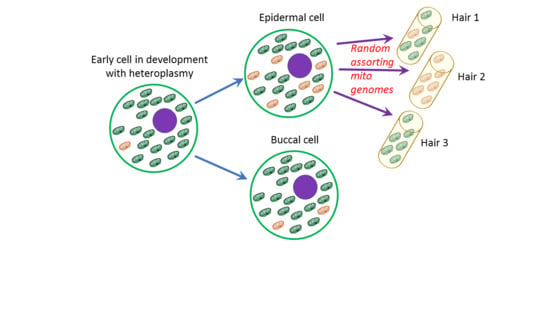
The possibility to type HP at a much lower level by MPS impacts the forensic comparison of evidentiary samples and references (as illustrated in
Supplementary Figure S1). The highly sensitive and truly quantitative information that is obtained by MPS provides detailed insight into HP variation within a person arising from the mtDNA bottleneck phenomenon [
5]. This improved sensitivity can be specifically informative for forensic comparisons of reference and hair specimens.
In the discussion, we have excluded LHP in HV2 and HV3, and any discussed LHP in this study is therefore limited to the HV1 C-stretch. Sites located in the middle of a potential C-stretch (T16189, T310 and T318–T319, surrounded by Cs at both sides) are regarded separately in the discussion as these sites show divergent patterns of variation even when the C-stretch is interrupted (further referred to as “PHP C-stretch-related sites”). Interpretation of these “PHP C-stretch-related sites” is different between MPS and Sanger as every MPS read represents a single molecule rather than the consensus signal of Sanger where LHP variation adjacent to the sites overlaps with the signal of the site itself.
3.1. HP Occurrence in Buccal Reference Samples Analysed with MPS
All 26 buccal reference samples were analysed through MPS. All mixed positions were identified, and the HP levels were categorised (percentage contribution of the minor variant), as shown in
Table 1. At lower levels, HP occurs more frequently. On average, a total of 1.8 PHP events per sample were observed by MPS analysis and 0.15 PHP events per sample by Sanger (haplotypes typed by Sanger and MPS are displayed in
Supplementary Table S1). Six buccal samples were only typed for HV1 and HV2, but none of the observed HP sites in these samples by MPS were located outside of the HV1 and HV2 region. In addition, LHP in HV1 was observed for half of the buccal samples showing 16189C when the transition resulted in an uninterrupted homopolymer of nine or more C residues. Many low-level PHP events (48% of the total number of HP events) were located in or around the HV2 C-stretch (position 310, 316, 318 and 319) or at HV1 position 16,189.
From the MPS-typed PHP events, most events with levels > 20% were originally typed by Sanger as expected (scenario 1 of
Supplementary Figure S1), except for 16320Y in X1. While 16320Y reached a level of 46% in the MPS analysis of this sample, a new inspection of the Sanger profile revealed only a minimal signal of the T-variant in the Sanger sequence (
Supplementary Figure S2). From the sites in the 10–20% MPS category, one out of four events was also typed by Sanger (the remaining MPS-typed HP sites follow scenario 2 of
Supplementary Figure S2).
It should be noted that new buccal swabs were used for the MPS analysis with sampling moments of up to several years between the Sanger and MPS samples which could explain some variation between the Sanger and MPS results. HP has been shown to accumulate with age [
16]. However, since most low-level buccal HP sites are also observed in at least part of the corresponding hairs (taken around the time of the Sanger buccal sample), the observed difference is more likely to be caused by the difference in the detection level than by the relatively small age difference of the individuals at the sampling moments of the buccal references.
3.2. MPS Analysis of Buccal References Resolves Mismatches between Buccals and Corresponding Hairs Seen with Sanger Sequencing
The increased sensitivity of mixed position detection by MPS revealed more HP events in the buccal reference samples. This may reduce the number of apparent homoplasmic mismatches when buccal references are compared to individual hairs. To focus the analyses, we specifically regard the two variable locations for which the samples had been selected and studied by Sanger sequencing (including 11 of the 18 “PHP events at other sites” from
Table 1): position 16,093 (14 samples), and positions 16,182 and 16,183 as a group (12 samples) [
7]. Sanger-sequenced data of the hairs [
4,
7] were compared to both Sanger- and MPS-analysed data of the corresponding buccal references (
Figure 2).
When regarding position 16,093, only one of the 14 buccal references showed a C/T (Y) PHP upon Sanger sequencing (X4,
Figure 2A), while for four references, only a C was detected, and for nine references, only a T was detected. Notwithstanding, not only the Y-typed reference, but also all four C-typed references showed different genotypes for the Sanger-sequenced hairs: X1, X2, X3 and X4 showed T, Y and C hairs and P11 showed Y and C hairs. The Sanger T-typing for X1, X2 and X3 (in total 16/54 hairs) is a mismatch with the C-typed reference (
Figure 2A, patterned bars). When the MPS data for the buccal references are regarded (with an analysis threshold of 3%), not only X4 but also X1, X2, X3 and P11 are typed as Y (black dots
Figure 2A, following scenario B of
Supplementary Figure S1) and these mismatches become matches. When regarding location 16,182–16,183, mismatches between Sanger-sequenced hairs and buccals are seen for P1_AA (P5 in [
4,
7]), P2_AA, P3_AA, P5_AA and P6_AA (in total 9/164 hairs) at position 16,183 (A-typed in buccals, C-typed in hairs). Again, these mismatches are resolved when the MPS buccal results are used as low-level HP is detected (patterned bars
Figure 2B, following scenario B of
Supplementary Figure S1).
Overall, due to its increased sensitivity, MPS indicated HP in the buccal references of 12 of the 26 individuals (P4_AA, P2_CC and P3_CC also show a minimal mixed contribution on position 16,183, but not within the MPS detection limit (scenario C of
Supplementary Figure S1)), while Sanger indicated HP for only one individual. When the Sanger reference data are used, for eight individuals, mismatches appear with homoplasmic hairs (25 events in total), but these mismatches are resolved when the MPS reference data are used as the homoplasmic hair variant corresponds to the minor heteroplasmic variant in the buccal reference. Thus, it is unlikely that these apparent homoplastic hair variants represent de novo mutations.
3.3. Minor Buccal HP Variants Observed as Apparent Homoplasmy or High-Level HP in Hairs by MPS
In the examination of three PHP sites in the previous section, we observed that the minor variant of a PHP site in the buccal can occur as an apparent homoplasmy in a portion of the hairs by Sanger (
Figure 2A: T-typed hairs for X1, X2, X3 and X4;
Figure 2B: AC-typed hairs for P1_AA (P5 [
4,
7]), P2_AA, P3_AA, P5_AA and P6_AA). Next, the overall occurrence of the phenomenon was examined by considering all HP events in the Control Region for all 26 individuals. Since MPS is more sensitive in detecting HP than Sanger sequencing, MPS analysis was used for both the 26 buccal references and the 475 corresponding hairs. We consider MPS homoplasmy when no minor variant exceeding the 3% allele calling threshold is observed (which does not exclude low-level heteroplasmy below 3%).
Figure 3A and
Supplementary Table S2 display the proportion of hairs with MPS homoplasmy of the minor buccal HP variant categorised by the corresponding level of HP in the buccal. Narrow ranges were used for low-level buccal HP categories and broader ranges for higher-level categories. As the lower categories are closest to the detection limit, they are important to gain insight down to which buccal HP level the minor HP variant can reach homoplasmy in corresponding hairs.
There are 11 apparent MPS homoplasmic occurrences of the minor buccal HP variant in the hairs (
Figure 3A), all involving PHP; five different HP sites are involved (195, 16,093, 16,183, 16,256, 16,320) and the frequency of the minor variant in the buccal reference is >4.5%. We also regarded the proportion of hairs approaching (>75% contribution of the buccal minor) homoplasmy, which roughly resembles hairs that could appear homoplasmic upon Sanger sequencing (
Figure 3B). Now, 13 incidents are observed including one additional LHP position (16193del), all with an HP level in the buccal above 4%. Sites 16,093 and 16,183 are most frequently involved since most individuals were selected for PHP at these positions [
4,
7]. This finding confirms the mismatch results found with Sanger sequencing of hairs as shown in
Figure 2 for sites 16,093 and 16,183. For site 16,183, it is noticeable that the buccal PHP levels are relatively low compared to other sites in the buccal references (between 4 and 10% only) even though occasionally high levels in hairs are observed. For all individuals involved, 16183C is directly adjacent to a long C-stretch (≥10 Cs) that occurs from a predominant C polymorphism at 16,189. Since, also for MPS, C-stretches and adjacent positions tend to exhibit increased error rates [
9], it could be that the obtained levels of the C-variant for 16,183 in the buccal samples are somewhat biased (here causing either an underrepresentation of the 16183C level in buccals or an overrepresentation of 16183C homoplasmy in hairs).
It is important to note that no MPS homoplasmic hairs were seen for which the variant was not detected as a (low-level) HP in the buccal reference. Interestingly, also the 16320Y HP that was hardly visible in the Sanger reference sample (
Supplementary Figure S2) was observed homoplasmic in one of the corresponding hairs and with >75% contribution in 32% of the hairs. Except for the previously discussed cases, no other Sanger buccal-to-hair mismatches were observed. In this study, we analysed 26 individuals and 475 corresponding hairs. While the numbers of hairs are substantial, the number of individuals is not sufficient to conclude that a complete mismatch between a buccal reference and a hair of the same individual cannot occur with MPS at all. For non-C-stretch-related positions, the buccal reference HP levels were at least 7.5%. Since this exceeds the 3% MPS analysis threshold by more than 2-fold, it is not very likely to observe an apparent MPS homoplasmic mismatch for a buccal and hair within the same individual. This is further supported by the fact that none of the HP minor variants with levels <4% were observed at an HP level of >75% in any of the tested hairs. However, since this study includes a total of 26 different haplotypes, it cannot be excluded that specific haplotypes or variants exist with a different pattern of HP variation.
Interestingly, in the buccal reference samples, all individuals carrying a C for 16,093 seem to have some level of PHP, while all individuals carrying a T appear to be homoplasmic, also at MPS resolution. This confirms the observations obtained with Sanger sequencing and indicates that 16093T is less prone to mutation than 16093C, as suggested before in several studies [
3,
7,
17].
3.4. Variation between the Observed HP Frequencies in Buccals and Corresponding Hairs by MPS
When a PHP site is detected in both a trace and a reference sample, it can provide additional confirmation that both samples may derive from the same individual. It is informative to assess how often PHP variants with a specific level in reference buccals are reproduced in hairs and vice versa. To examine the overall relation between the PHP levels in buccal references and corresponding hairs,
Figure 4 was generated. As PHP sites located within potential C-stretches exhibited a different pattern of PHP variation, we analysed these as separate groups (
Figure 4A,B). On the
y-axis, the proportion of hairs is presented categorised by the hair PHP level, while on the
x-axis, categories of PHP levels for the buccal samples are displayed. In the 26 buccal samples, HP was observed at 16 different positions: 1 LHP (position 16,193), 5 C-stretch-related and 10 non-C-stretch-related sites. The total number of buccal HP occurrences in the 26 individuals was 60 for buccals; the total number of investigated occurrences for these positions in hairs was 1099 (236 LHP, 552 C-stretch-related PHP and 311 non C-stretch-related PHP).
In general,
Figure 4 shows the trend that with higher PHP levels in buccals, high PHP levels in hairs are seen more frequently. However, the distribution of PHP levels in hairs is broad, as is expected from mtDNA bottleneck assorting during hair development [
5]. For instance, for buccal sites with a PHP level above 15%, >90% of the hairs show the PHP as well (categories 3–97%,
Figure 4A), at levels in a very wide range. For buccal sites with PHP levels of 7–10%, PHP is absent in a much larger percentage of the hairs (47% of the hairs in category 0–3%,
Figure 4A). Buccal PHP sites with levels below 10% are generally absent in the majority of hairs. Therefore, it could be considered to record PHP variants in databases only if levels exceed 10% and compare the lower-level PHP variants only in the case of a single mismatch. As this percentage approaches the PHP levels observed by Sanger sequencing, this strategy would also avoid large differences between database entries generated by Sanger or MPS.
Interestingly, mixed sites located on positions that lead to C-stretches tend to concur with lower mixed levels in hairs compared to other PHP sites while the extracts and libraries were processed in the same way. Since the C-stretch-related PHP sites were generally low-level, this might partly be the result of bias introduced by C-stretch-related sequencing errors seemingly causing an underestimation of HP levels of the C-stretch variant. For example, if we focus on position 310 (where a T > C SNP leads to a long C-stretch) as shown in
Supplementary Figure S3, PHP is frequently observed at low levels. A trend is observed of rising frequencies in the hairs for individuals with a higher frequency in the buccal, but the frequencies in the hairs are much lower than those observed for other PHP sites (
Figure 4A). While a portion of the seemingly PHP in the buccals could originate from PCR or sequence artefacts, reads of the molecules containing a long C-stretch are more likely to fail quality criteria during the basecalling process, resulting in biased lower levels of the C-stretch variant. Thereby, they will more often fall in the category below the allele calling threshold. However, the same errors would be expected in hairs and buccals, so the exact cause of the difference between these two remains unclear. For LHP sites, no clear trend was observed in the levels between buccals and hairs (
Supplementary Figure S3C).
3.5. Mixed Sites Observed in Hairs, but Not in Corresponding Buccals
Up to now, we examined the concurrence of buccal HP sites in hairs. Besides, 36% of the hairs were found to carry mixed positions (above the 3% calling threshold) on sites where no HP was observed in the corresponding buccal reference sample. A total of 579 mixed occurrences were observed in the 475 analysed hairs. These were dispersed over 162 different positions, and for 70 positions, the same mixed position was seen in more than one hair. A total of 306 instances (44 positions) involved at least two hairs of the same individual, suggesting that this HP might exist throughout the cells of these individuals but below the MPS detection level in buccals. The mixture levels were mostly very low (273 of the 579 occasions had a mixture level between 3 and 5%; 93 resided above 10%). Further, multiple “non-buccal” mixed sites could be found in the same hair, but none of the hairs showed more than one mixed site exceeding 10%.
While most hairs exhibit a maximum of three mixed HP sites, 12 hairs stood out since they contained 4–8 mixed sites at low levels. Interestingly, 11 of these 12 hairs belong to the same individual and the mixed sites were all in the same fragment that contains a C-stretch of 10 Cs (due to an insertion of four Cs after position 573). Since all the mixed sites in this fragment represent additional Cs (although mostly not adjacent to the C-stretch itself,
Supplementary Figure S4) and they are observed at similar levels in the buccal (although some just below the detection threshold), it suggests that this specific sequence results is an accumulation of errors rather than being actual mixed sites (the sites were excluded for other calculations in this paper). The suspected errors are present in the raw sequencing data from the instrument, so they could reside from either the PCR, the sequencing process or from the basecalling process in the sequencer.
Since several samples were specifically selected for containing a PHP at mtDNA positions 16,093, 16,182 or 16,183 [
4,
7], the tested samples do not represent a random population and seemingly “de novo” HP events in hairs could only be studied for positions that were not already HP in the buccal of the sample. From these sites, 12 HP sites stand out since they are observed in >10 hairs; all were observed in at least two hairs of one individual (
Figure 5A). Some positions are limited to two or three individuals (A16183M, and C16278Y); others are common and seen in at least eight individuals (T16224Y, C16290Y, G16390R, A73R, T152Y, G316R and A561M). Although these 12 HP sites shown in
Figure 5 stand out for the frequency at which HP is seen among hairs, the level of HP does not specifically stand out (
Figure 5B); of the 93 positions that have a mixture rate of >10%, only 26 are at these 12 HP sites. Although these sites stand out for the frequency at which HP is seen among hairs, the level of HP does not specifically stand out (
Figure 5B); of the 93 events with a mixture rate of >10%, only 26 are at these 12 HP sites.
In general, heteroplasmy detection is more sensitive with MPS than with Sanger sequencing, but its interpretation depends on several factors, such as background noise, coverage and strand bias. The authenticity of heteroplasmy also depends on the contamination rate; several examples can be found in the literature [
18,
19] where some of the reported heteroplasmies are most likely the result of contamination [
8].
Although contamination cannot be totally ruled out, three of these variants (C16290Y, G16390R and A561M) were not present in any of the samples that were prepared together with the hairs, nor in any of the positive and negative controls or the haplotypes of the analysts. Since most of the haplotypes differ for two or more positions from each other, contamination would mostly result in multiple (linked) mixed sites, which was not the case. While the number of individuals is too low to look at potential haplogroup-specific patterns of HP variation, data from the hairs suggest that these seven common “de novo HP sites” might be more prone to HP formation than other positions. Interestingly, position 152 overlaps with the five most frequent PHP sites in buccal cells and blood samples, as reported by Irwin et al. [
3] with Sanger sequencing. PHP at positions 16,183, 16,224, 16,278, 16,290, 16,311, 16,362, 16,390, 73 and 152 were previously found in [
20] or in [
3] or in both, while 316, 344 and 561 were not. From these three positions for which PHP has not been observed previously, 316 and 561 are adjacent to repeated Cs, suggesting that sequencing errors might be a factor here which would explain why they were observed mixed more frequently than other positions.