1. Introduction
The MAPK organizer-1 (MORG1) protein belongs to the family of the WD40 repeat domain-containing proteins [
1]. The WD40 domain-containing proteins are devoid of enzymatic activities, but due to their multiple WD-repeat structure, play an important role in assembling functional protein complexes [
2] and are involved in different cellular processes, such as transcriptional regulation [
3], ubiquitin-dependent protein degradation [
4] and chromatin modification [
5]. MORG1 was first discovered as a binding partner of the MP1-protein, that is, a part of the extracellular signal regulated kinases (ERKs) module, and facilitates ERK activation [
1]. Moreover, it was shown that MP1 is not the solely protein that interacts with MORG1 from the ERK-signaling path. In addition, it was shown that MEK-1, RAF-1, and ERK1,2 are binding partners of MORG1 [
1]. Simultaneously, MORG1 was identified by us in a yeast two hybrid system as a protein that associates with the HIF-prolyl hydroxylase domain containing protein 3 (PHD3) [
6]. PHDs are pivotal cellular regulators of the HIF transcription factors [
7,
8]. Those findings suggest an important function of MORG1 as a scaffold protein for two signal-transduction pathways regulating proliferation via a MAPK-signaling module and hypoxia/normoxia regulation and adaptation to hypoxia via its complex with PHD3.
Hypoxia often occurs because of a microcirculation injury and hypoperfusion in tissues and organs, including the kidney [
9]. Hypoxia contributes to renal injury in acute and chronic kidney diseases [
10] and as well as in CKD-related pathological processes as anaemia or inflammation [
11]. There is increasing evidence that HIF activation may protect against renal damage [
12,
13,
14]. Hypoxia Inducible Factors (HIFs) are the major transcription factors that are activated under hypoxic conditions and have an evident role in kidney injury and repair by inducing the expression of numerous HIF target genes involved in the cellular adaptation to hypoxia [
15]. HIFs are heterodimers, which are degraded under normoxic conditions due to prolyl-hydroxylation-dependent ubiquitination and subsequent proteasomal degradation [
16]. Our recent data revealed that the reduced expression of MORG1 in
MORG1+/− mice improved renal damage in animal models of type 1 and type 2 diabetes mellitus [
17,
18] and ameliorated renal injury during systemic hypoxia [
19]. Thus, our findings depicted that a better understanding of the potential mechanism of MORG1 regulation may eventually lead to the generation of novel therapeutic tools to treat hypoxic renal damage.
2. Materials and Methods
2.1. Cells and Treatments
The regulation of the human MORG1 expression was studied in parental human HEK 293 cells (kindly provided by J. Mueller, Center of Molecular Biomedicine, Jena, Germany). HEK 293 cells were cultured in DMEM (Invitrogen, Schwerte, Germany) containing 4.5 g/L glucose, supplemented with 10% heat inactivated fetal bovine serum (Invitrogen, Germany), 100 U/mL penicillin and 0.1 mg/mL streptomycin (Sigma-Aldrich, Darmstadt Taufkirchen, Germany). For analyses, 3 × 105 cells per well were seeded into 12 well plates. On the next day, the medium was replaced with DMEM, containing 1 g/L glucose, supplemented with 5% heat inactivated fetal bovine serum, (Invitrogen, Germany) and penicillin/ streptomycin. To study the effect of hypoxia on the MORG1 expression in HEK 293 cells, they were exposed for 6, 12 or 24 h to 10% O2 in hypoxia cell culture incubator from TermoScientific (Schwerte, Germany), containing 5% CO2, 10% oxygen and 90% nitrogen gas. In addition to hypoxia, we studied the effect of two HIF-prolyl-hydroxylase (PHDs) inhibitors: 500 µM 3,4-dihydroxybenzoic acid (3,4-DHB) and 500 µM L-mimosine, both purchased from Sigma-Aldrich (Taufenkirchen, Germany).
2.2. Reverse-Transcription and Quantitative Real-Time PCR Analyses
Total RNA was isolated from HEK 293 cells using the RNeasy kit (Qiagen GmbH, Hilden, Germany), according to the manufacturers’ recommendation. To assess the quality of the RNA, gel electrophoresis with RNA stained with SYBR Green dye (Molecular Probes) was performed on samples. RNA concentrations were determined, and one µg total RNA was reverse transcribed using an M-MLV-RT kit (Promega, Mannheim, Germany) for 1 h at 37 °C. The gene expression was estimated by real-time PCR (RT-PCR) in a multiplex assay as previously described [
20] using 0.5 µM of forward reverse gene specific primers and a LightCycler
® 480 SYBR Green I Master Kit (Roche Diagnostics, Mannheim, Germany). The amplifications were performed using a Q-Tower thermocycler (Analytik Jena Bio Solutions, Jena, Germany). The sequences of the primers used for analyses were as follows: human
gapdh forward 5′-GGAGTCAACGGATTTGGTCG-3′, reverse: 5′-CAGTGGACTCCACGACGTAC-3′, human
MORG1 forward 5′-AGGAATACAAGCTGGACTG-3′, reverse: 5′-AGCTACGTCTCTAGAATAAGATGG-3′, human
VEGFA forward 5′-TTGCCTTGCTGCTCTACCTCCA-3′, reverse 5′-GATGGCAGTAGCTGCGCTGATA. The relative gene expression was quantified by ΔΔCT method, and the relative expression ratio was calculated according to Livak et al. [
21], where
Ratio = 2
−ΔΔCT [
21]. The primers sequences used for the analyses of the gene expression in HEK 293 cells were purchased from Invitrogen (ThermoFisher, Invitrogen, Life technologies, Darmstadt, Germany). Annealing temperatures and amplification conditions were as previously described [
17,
18]
2.3. Cloning of the Murine MORG1 Promoter and Generation of the Luciferase Reporter Constructs
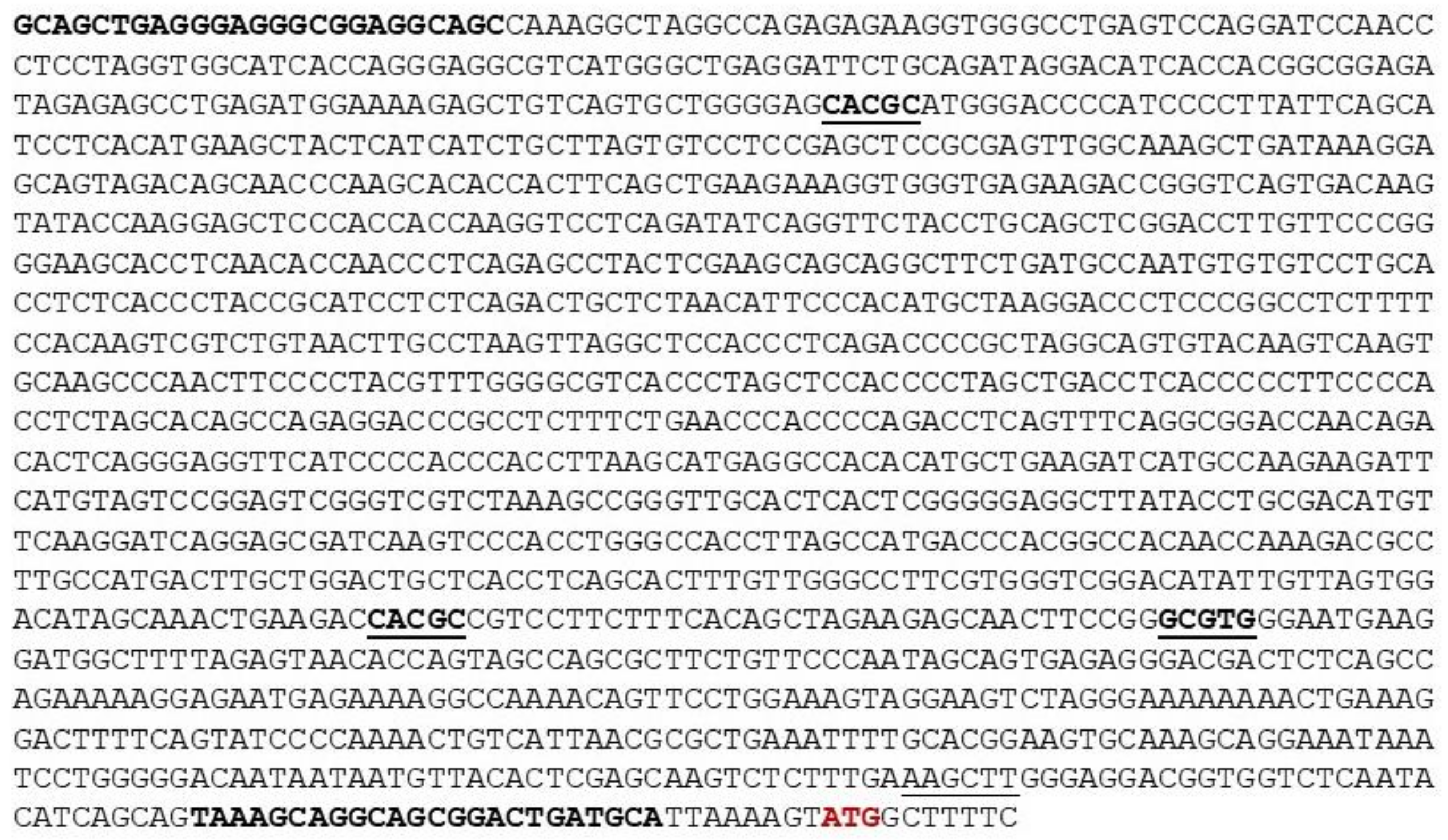
A fragment corresponding to 1473 bp of the 5′ untranslated region of the murine MORG1 genomic sequence upstream of the ATG codon was amplified from a DNA isolated from a BAC clone pMQ155E2 purchased from Source Bioscience, Nottingham, U.K., containing the murine genomic MORG1 clone with the following primers: forward primer sequence 5′ GCAGCTGAGGGAGGGCGGA3′ and reverse primer sequence 5′ TGCATCAGTCCGCTGCCT 3′. The fragment was amplified with PhusionTM High-Fidelity DNA polymerase (TermoFisher, Schwerte, Germany). After that, the PCR product was digested with HindIII endonuclease enzyme; the location of the internal HindIII restriction side is underlined on the murine promoter sequence. Next, the fragment was gel purified and cloned into the digested with SmaI and HindIII promoter-less pGL2-basic vector (Promega), containing the firefly luciferase gene as a reporter gene. The corresponding sequence of the murine MORG1 promoter was confirmed by sequencing.
Subsequently, we performed a specific analysis of the hypoxia-inducible factor binding sites on the h
MORG1 promoter by the generation of point mutations in the putative HRE binding sites, where the A/GCGTG was mutated to A/GAAAG in the corresponding constructs. The point mutations were generated using 100 ng of the WT
MORG1 promoter reporter construct as a template, 5 U/reaction of Pfu-Turbo DNA polymerase (Agilent Technologies, Waldbronn, Germany) and 125 ng of each forward and reverse primer, caring the mutated site of the putative human
MORG1 hypoxia-responsible element binding site (HBS). The reactions were performed in a final volume of 50 µL. The primer sequences used for the mutagenesis are presented in
Table S1. The cycling conditions were as follows: 95 °C denaturation for 5 min, followed by 18 cycles of 95 °C for 30 s, 60 °C annealing for 45 s and 16 min elongation step. Next, the PCR reactions were digested with 5 U
DpnI restriction endonuclease (TermoFisher, Schwerte, Germany) for 3 h at 37 °C and the enzyme activity was inactivated for 15 min at 75 °C. 5 µls of the
DpnI treated reactions were transformed into
E. coli XL-1 Blue competent cells. Plasmid DNA was isolated from several colonies and subjected to sequencing using the sequencing primers pGL2 forward: 5′ CAATGTATCTTATGGTACTG 3′ and pGL2 reverse: 5′ CCTTATGCAGTTGCTCTCCA 3′. For simplicity, the constructs of the human
MORG1 promoter caring the corresponding single mutations were termed HBS A, HBS B, HBS C and HBS D, respectively. For the generation of the double HBS-binding site mutations of the h
MORG1 promoter, the plasmids caring the single HBS A, HBS B and HBS C mutants were digested with
XhoI and
HindIII endonucleases (Analytic Jena, Jena, Germany). The corresponding 496 bp fragment, containing the mutated sites, was gel purified and ligated into
XhoI and
HindIII digested HBS D vector plasmid containing the HBS D single mutation. Thus, were generated the
MORG1 promoter constructs caring double mutations as follows: HBS A + D, HBS B + D and HBS C + D; the corresponding reporter constructs of the human
MORG1 promoter were named thereafter as the double sites HBS A + B, HBS B + D and HBS C + D. All mutations were confirmed by sequencing analyses.
2.4. Transfections and Reporter Analysis
To analyse the transcriptional activation of the h
MORG1 promoter 3 × 10
5 HEK 293 cells were seeded into 12 well plates as described above and transiently transfected on the next day in DMEM medium, containing 1 g/L glucose, 5% heat inactivated serum and antibiotics. For transient transfections, 2 µg of the corresponding human WT
MORG1 promoter reporter constructs or mutants of the h
MORG1 promoter reporter plasmid were co-transfected with 0.5 µg of β-galactosidase control plasmid (TAKARA Bio, San Jose, CA, USA), using TurboFect transfection reagent (ThermoScientific, Schwerte, Germany) for 12 h. Thereafter, a fresh DMEM medium, supplemented with 1 g/L glucose, 5% heat inactivated serum, antibiotics and the corresponding stimuli, was added and the cells were incubated for different time periods at the absence (control, respectively normoxia samples) or at the presents of stimuli as shown under the figure legends. Finally, the cells were washed with sterile PBS and lysed in 100 µL of cell lysis reagent (Promega). The cell lysates were vortexed and centrifuged at 13,000 rpm for 15 min. The luciferase activity was determined on a Tecan Infinite M200 (Tecan, Austria GmbH, Grödig, Austria) with a luminometer mode using 20 µL of the cells lysates and 100 µL firefly luciferase substrate (Promega). The ability of 3,4-DHB, L-mimosine and hypoxic conditions to induce HIFs activation was assessed by transient transfection of the
pHIF-Luc reporter plasmid (Signosis, Sunnyvale, CA, USA) [
20] and 0.5 µg of a β-galactosidase control plasmid (TAKARA Bio). The corresponding
pHIF-Luc reporter activity was assayed as described above.
2.5. Western Blot Analyses
The protein expression was analyzed using Western blots after separation of proteins on 10 or 12% SDS-PAGE gels. After stimulations as appropriate, the cells were washed with PBS and lysed in complete lysis M buffer, supplemented with protease inhibitors (Roche, Mannheim, Germany) and 100 mM of Na3VaO4 (Sigma-Aldrich). The protein concentration was detected in DeNovix, DS-11FX Spectrophotometer/Fluorometer (DeNovix Inc., Wilmington, DE, USA) under UV 280 mode. Protein expression was detected with ECL reagent (PerkinElmer, Waltham, MA, USA) and documented with a Fusion FX7 EDGE imaging system (Vilber Lourmat GmbH, Eberhardzell, Germany). Protein expressions were measured via ImageJ software and normalized to the corresponding loading control protein. For the detection of MORG1 protein, a rabbit polyclonal antibody generated in Biotrend (Berlin, Germany) was used in dilution 1: 1000. The equal loading of total protein or cytosolic fraction protein was monitored by the detection of vinculin expression using a mouse antibody to vinculin 1: 1000 dilution (Santa Cruz Biotechnology Inc., Heidelberg, Germany). For detection of HIF-1α protein (Abcam, Cambridge, UK, 1:1000) and for HIF-2α, an antibody from Novus Biologicals (Wiesbaden, Germany, 1:500) was used. The PCNA (Proliferating cell nuclear antigen) antibody was purchased from Santa Cruz Biotechnology Inc. and it was used in dilution 1:500. The corresponding secondary antibodies were used in dilution 1:2000 purchased from KPL (Sera Care, Gaithersburg, MD, USA).
2.6. Biotin-11 UTP Labelling of ssDNA Oligonucleotides
The primers for electrophoretic mobility assay were labelled using BIO 3′-END oligonucleotide labelling kit with Biotin 11-UTP (Jena Bioscience, Jena, Germany), according to the manufacturer’s instructions. In brief, 5 pmol ssDNA oligonucleotides were labelled with 50 pmol Biotin 11-UTP by terminal deoxynucleotidyl transferase (TdT) for 30 min at 37 °C. We evaluated the Biotin 11-UTP labelling by dot blot analyses. Afterwards, two complementary ssDNA oligonucleotides (sense and anti-sense) were annealed in annealing buffer containing: 10 mM TRIS-HCl pH 7.5; 1 mM EDTA and 50 mM KCl in a heating block at 95 °C for 5 min, then the mixture was allowed to cool down to 25 °C.
2.7. Electrophoretic Mobility Shift Assay
We further analyzed the
MORG1 putative HIF binding sites by electrophoretic mobility shift assay (EMSA). Nuclear protein extracts for EMSA were prepared using a mammalian nuclear and cytoplasmic protein extraction kit (Serva Electrophoresis GmbH, Heidelberg, Germany), as recommended by the manufacturer. The protein concentration was measured in DeNovixDS-11FX Spectrophotometer (DeNovix Inc., Wilmington, DE, U.S.A) and the proteins were kept frozen at −20 °C until the analyses. The EMSA binning reactions were performed for all four putative HBS of the h
MORG1 promoter region by the aid of an active motif non-radioactive assay kit (BIOZOL Diagnostica, Eching, Germany). The reactions were performed in final volume of 20 µL and usually contained 10 µg of nuclear extracts, 1 µL of 10× reaction binding buffer, 1 µL of 2.5 mg/mL BSA, 1 µL of 50% glycerol, 2 µL of 0.1 mg/mL dI-dC, and 2 µL of 3′-end Biotin-11 UTP labelled dsDNA oligonucleotides. After that, the reactions were incubated for 30 min at room temperature and loaded on a 6% non-denaturing PAGE. The gel was transferred via semi-dry transfer system (Bio-Rad Laboratories Inc., Feldkirchen, Germany) on a positively charged membrane for 1 h. Next, the membrane was removed and placed side down on a UV-transiluminator Vilber (Vilber Lourmat GmbH, Eberhardzell, Germany) equipped with 4 × 8 W 365 nm UV-lamp and cross-linked for 15 min. The membrane was blocked for 1 h in blocking buffer, followed by 1 h incubation with a stabilized Streptavidin–HRP conjugate in 1:330 dilution in blocking buffer. The membrane was washed 4 times each 5 min with washing buffer. Finally, the membrane was equilibrated for 10 min in equilibration buffer and incubated with an EMSA-ECL solution for 10 min. The signals were visualized on a Fusion FX7 EDGE imaging system (Vilber Lourmat GmbH, Eberhardzell, Germany). All buffers, the streptavidin–HRP conjugate and ECL reagents were provided with the active motif assay kit (BIOZOL Diagnostica, Eching, Germany). For competitor studies to the binding reactions, either 100-fold excess of the corresponding unlabeled dsDNA oligonucleotides or 10-fold excess of unlabeled mutated dsDNA oligonucleotides were added. For super shift analyses, 1 µL of HIF-1α of HIF-2α antibodies from R&D System (Wiesbaden, Germany) were added for 15 min, prior to addition of the 3′-end Biotin-11 UTP labelled dsDNA oligonucleotides, followed by incubation for an additional 30 min at RT. The sequences of the oligonucleotides used in the EMSA analyses are shown in
Supplementary Table S1.
2.8. Statistics
Statistical analyses were performed with the statistical package SigmaPlot 14.5 software (SYSTAT Software, San Jose, CA, USA). The results were evaluated by Analysis of Variance (ANOVA) followed by a t-test to analyze the differences between the two groups. The data are graphically presented as a box plot, where the values are shown as the median and percentiles, and to the box plot was added additional a vertical point plot of all sample’s value. In the bar graphs, the values are presented as mean ± standard error of mean (SEM). Differences were considered significant when p < 0.05. * p < 0.05; ** p < 0.01. *** p < 0.001.
4. Discussion
The identification of the MAPK-organizer protein 1 (MORG1) as a binding partner of several binding modules provoked an interest in the function and regulation of the MORG1 gene and protein expression in health and diseased conditions. Although studies from MORG1
+/− mice revealed that reduced MORG1 expression is renoprotective and diminished the development of diabetic nephropathy in a mouse model of type 2 diabetes mellitus [
18], it reduced renal injury in systemic hypoxia [
19] and shows an anti-inflammatory function in septic AKI models [
27]; the mechanisms regulating the MORG1 expression are not investigated. With this study, we cloned murine and human MORG1 promoter regions. Exploring the ALGGEN-PROMO link [
23], we identified putative regulatory elements in both promoter sequences. The numerous predicted transcription factor binding sites in the murine and human MORG1 promoter regions need to be further investigated in more detail. Surprisingly, among other depicted putative TF binding site, several predicted hypoxia-responsible elements (HRE) binding motives were identified in both murine and human MORG1 promoter sequences corresponding to the canonical HRE binding sequence 5′-A/GCGTG-3. HIF binding motifs are located similarly in the murine and human MORG1 promoter sequences, except for the most proximal putative HRE binding site on the human promoter, the HBS A, which is situated just 67 bp upstream of the ATG codon. To this sequence has been also predicted the binding of ARNT, aryl hydrocarbon receptor nuclear translocator, also known as HIF-1β and the AHR (aryl hydrocarbon receptor). The heterodimer ARNT: AHR binds to the core DNA sequence 5′-TGCGTG-3′ within the dioxin response element (DRE) of the promoter sequences and activates their transcription [
28]. Furthermore, environmental factors act through AHR receptor activation and regulate energy metabolism, and the disruption of energy metabolism [
29] can result in metabolic diseases such as obesity [
30], diabetes and hypertension [
31]. This complex is closely related to hypoxia response elements as ARNT interacts with HIF-1α, HIF-2α in hypoxic conditions [
32,
33,
34]. The interrelated roles of AHR and HIF-1α signaling play an important role in the coordination of such processes as infection and inflammatory diseases [
33]. Thus, some of the predicted transcription factors binding sites correlate with our current knowledge well for the role of MORG1 in physiological and diseases conditions. Although we have more data supporting the involvement of MORG1 in the HIFs activation from animal studies, we selected to focus initially on the h
MORG1 promoter for our analyses, because understanding the mechanisms of
MORG1 regulation could be of potential therapeutic interest and, second, there are no data about the regulation and function of human MORG1. We explored the human HEK 293 cells for our studies as it was already shown that MORG1 is expressed in those cells [
6]. It is well documented that murine MORG1 protein plays an important role in the stabilization and accumulation of HIFs via MORG1/PHD3 complex [
6,
17,
20], but is unknown whether h
MORG1 expression is affected by hypoxic conditions. The finding that four putative hypoxia-inducible binding sites are present in the h
MORG1 promoter sequence raised our interest to study in more detail the ability of hypoxic conditions to affect the transcriptional regulation of the h
MORG1 promoter. Recent studies in HKC-8 cells have reported that HIF-1α and HIF-2α binding sites are independent of the duration and severity of hypoxia [
24]. Our data revealed that mild hypoxic conditions, as 10% hypoxia in a time-dependent manner, affected the hMORG1 reporter activity. While early time points were associated with a significant inhibition of the wild-type hMORG1 promoter activity, 24 h exposure to mild hypoxia showed low, but significant, enchantment of the reporter gene activation. Therefore, we observed a biphasic effect of hypoxic conditions on the regulation of the hMORG1 promoter. We have to point at this point that we were able to confirm that HIFs were activated at 12 h of 10% hypoxia as we observed a significantly increased expression of the hVEGFA mRNA under above conditions. Furthermore, exploring HIF-reporter assays, we measured an increased luciferase activation after 6 h and 12 h in HEK 293 cells exposed to 10% hypoxia or treated with 3,4-DHB or L-mimosine.
Although, in general, it is expected that the genes that are HIF targets in hypoxic conditions are positively regulated, characterized with an increased transcriptional activation [
35], it was also reported that hypoxia can inhibit TNF-dependent VCAM1 induction in endothelial cells [
36]. One explanation for the inhibitory effect of hypoxic conditions could be that at the first 6-to-12 h mild hypoxic conditions could induce the hypoxia-dependent activation of other transcription factor(s) regulated by HIFs, which in turn negatively regulate h
MORG1 transcriptional activity. Indeed, 10% hypoxia is approximately a 50% reduction of the normal oxygen supply. A very recent report demonstrated that even mild hypoxia can enhance HIF stabilization and the expression of VEGF and may induce destructive changes in kidney tissues [
37]. On the other hand, hypoxia could lead to the reduced expression of several genes, regulated by transcription factors activated by hypoxic stress (for example HIF-1α activation is shown to induce an NF-κB activation [
38]). It is also possible that the elevated
MORG1 reporter activation after 24 h exposure to mild hypoxia could be a result of the direct binding of HIFs to the putative HREs on the
MORG1 promoter. This binding may enhance the promoter reporter activity at the latter time point because the association is likely time dependent, or it is a result of competitive binding.
In contrast to hypoxia-dependent HIF activation, the suppression of HIF hydroxylation by PHD enzymes under normoxic conditions, by the application of pharmacological inhibitors of PHDs, was associated with an enhanced hMORG
1 promoter activity starting at 6 h for the 3,4-DHB inhibitor. We detected a sustained h
MORG1 promoter reporter activation at 24 h and 48 h with a single treatment of the reagent. Furthermore, the application of L-mimosine, another PHD unspecific inhibitor, was less effective at the early time point, but its effect was undistinguishable from the 3,4-DHB action at 12 h, 24 h or 48 h treatments. Interestingly, these data suggest that small PHD inhibitor molecules probably need more initial time until HIF transcription factors accumulate and promote the h
MORG1 transcriptional activity. Therefore, HIFs could have a slower kinetic of accumulation at the presence of L-mimosine compared with 3,4-DHB treatment, which could be due to the slower effect of L-mimonsine on the suppression of the PHD enzymatic activity. Such a stronger, but slower, inhibitory action of L-mimosine on tyrosinase inhibition has already been reported [
39]. Although both PHD inhibitors have common effects on the suppression of PHDs as well as the stabilization and activation of HIFs in normal conditions, it was shown that 3,4-DHB could suppress trophic factor deprivation-induced apoptosis partially in a HIF-independent manner in neuronal cells [
40]. Consequently, the mechanisms used from both pharmacological PHD inhibitors to induce HIF activation could be slightly different and cell-type specific. Several studies reported that hypoxia mimetics induced preferentially HIF-2α activation, thus acting less on HIF-1α or even suppressing HIF-1α activation showing a mutual antagonism of HIF isoforms in cardiac, vascular and renal disorders [
41]. Our data demonstrated that hypoxia and hypoxia mimetics induce HIF activation via different mechanisms, which may explain the differential effects observed on the regulation of h
MORG1 transcriptional regulation. This hypothesis is also supported from our experiments to reverse the hypoxic inhibition of h
MORG1 promoter by simultaneous exposure of the HEK 293 cells to 10% hypoxia and 3,4 –DHB. Although 3,4-DHB very potently reversed the h
MORG1 promoter reporter activation after 6 h, it was not able to overcome the hypoxic inhibition of the h
MORG1 promoter after 12 h co-treatment. This finding suggests that, at 12 h of mild hypoxia, different transcription factor(s) could be activated by HIFs indirectly, causing an inhibitory effect on
MORG1 transcriptional activity. Both HIF-1α and HIF-2α are expressed in HEK 293 cell, but the kinetic how they associate with the putative MORG1 HRE and how are activated from hypoxia or 3,4-DHB alone or in simultaneous treatment is unknown at present.
Furthermore, mutational analyses of the putative HIF binding sites on the hMORG1 promoter revealed that single HBS A or HBS C mutants were able to reverse the inhibitory effect of 10% hypoxia compared with the wild-type hMORG1 promoter reporter activity, suggesting that these sites have an inhibitory function on the hMORG1 promoter at 12 h treatments. We detected opposite results when hMORG1 reporter constructs carrying single mutation at the HBS B or HBS D were used. Interestingly, hMORG1 reporter constructs carrying mutation in the HBS A and HBS B further increased the activity of the mutated promoter’s reporter constructs, compared with the wild-type hMORG1 reporter activity, while HBS C and HBS D mutations were able to suppress the 3,4-DHB- or L-mimosine-dependent hMORG1 promoter activation at 12 h treatments. Therefore, these HIFs putative binding sites in the hMORG1 promoter could be more sensitive to the pharmacological activation of HIFs. The analyses of the hMORG1 promoter reporter constructs carrying two mutations revealed that the presence of the mutated HBS D binding site was sufficient to reduce the reverse the effects observed in the single mutants’ analyses. These data suggest that the distal putative HIF binding site in the hMORG1 promoter sequence HBS D could play an essential role for the HIF-dependent regulation of the hMORG1 promoter.
Moreover, HIFα subunits can also be hydroxylated at asparagine 851 blocking interactions with coactivators, suggesting an additional mechanism of regulation [
42].
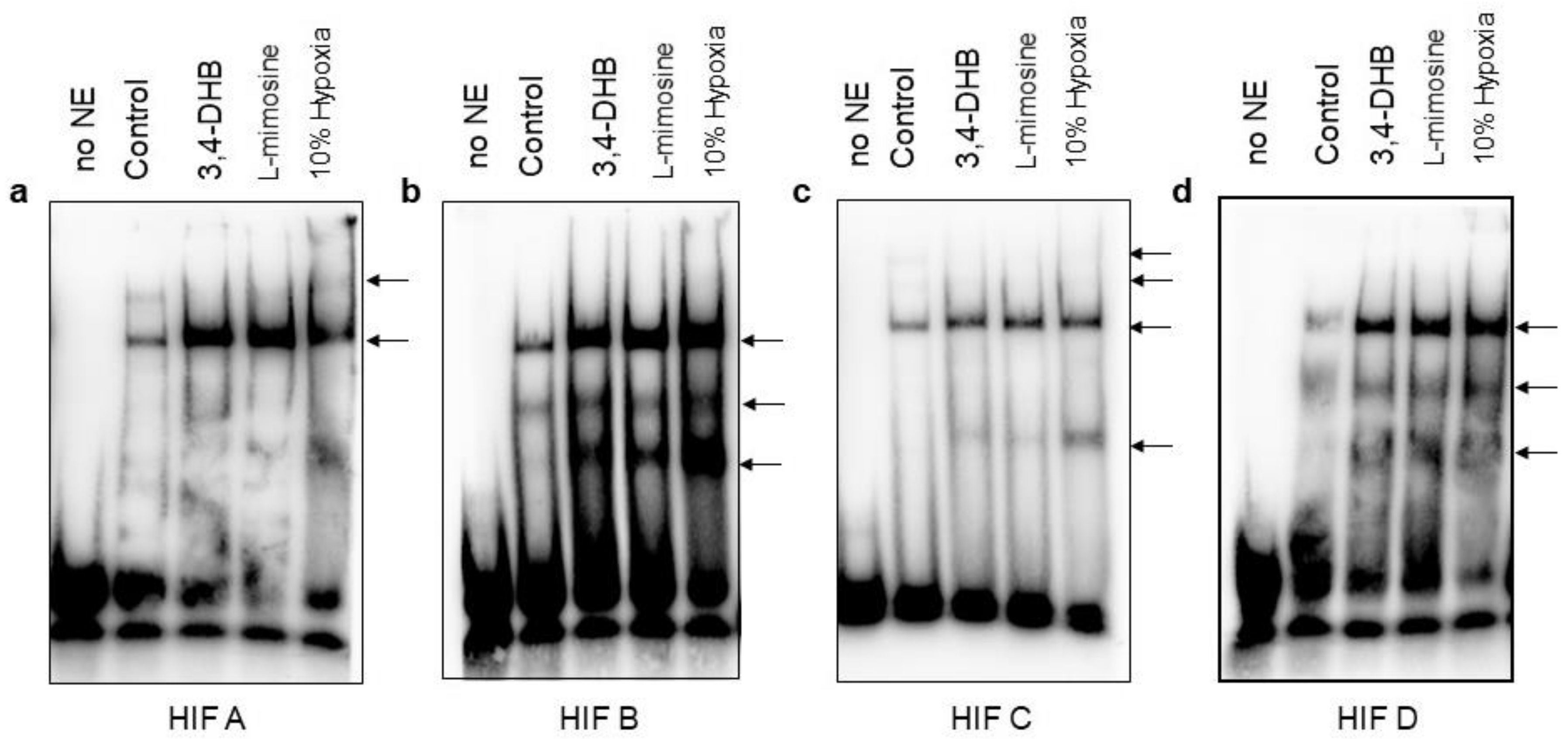
EMSA assays confirmed the complex specificity between the hMORG1 promoter putative HIF binding sites and HIF-1α and/or HIF-2α TFs, but is not yet clear how the exact complex formation occurs in hypoxic conditions and the pharmacological activation of HIFs via PHD suppression under normoxic conditions. Because the h
MORG1 promoter region contains four HIF binding sites, it is difficult to exactly determine whether there are preferential binding sites. It is expected that the proximal TF binding sites are probably positive regulators of the promoter activity, although we observed here that the HBS A presumable negatively regulate the h
MORG1 promoter activity in hypoxic conditions and by the pharmacological activation of HIFs in normoxic conditions through the suppression of PHD enzymes. In addition,
cis-active elements may build complexes with other
trans-factor and/or coactivators, further adding to the complexity of the regulation [
43]. Further approaches, as a CHIP assay, bioinformatic approaches as well as functional motif analysis of the promotor, are necessary to be performed to learn more regarding the complex regulation of the MORG1 promoter under hypoxic conditions. In addition, experiments with other cell types will be of advantage to find out whether this complex regulation is cell-type specific. Nevertheless, our data provide for the first time evidence that not only
MORG1 regulates HIF stabilization through a PHD complex, but also, vice versa, HIFs control MORG1 expression directly or indirectly by a complicated regulatory mechanism, which needs to be further elucidated.