Complete Mitochondrial Genome of Tanypus chinensis and Tanypus kraatzi (Diptera: Chironomidae): Characterization and Phylogenetic Implications
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling and Sequencing
2.2. Assembly, Annotation, and Composition Analyses
| Family | Species | GenBank Accession Number | Reference |
|---|---|---|---|
| Chironomidae | Clinotanypus yani | MW373524 | [2] |
| Tanypus chinensis | PQ014462 | This study | |
| Tanypus punctipennis | MZ475054 | [14] | |
| Tanypus kraatzi | PQ014453 | This study | |
| Parochlus steinenii | NC027591 | NCBI | |
| Diamesa tonsa | MZ158292 | [11] | |
| Diamesa loefferi | MZ127838 | [11] | |
| Sympotthastia takatensis | MZ231026 | [11] | |
| Boreoheptagyia kurobebrevis | MZ043576 | [11] | |
| Boreoheptagyia zhengi | OM302508 | [11] | |
| Prodiamesa olivacea | MW373525 | [26] | |
| Monodiamesa sp. | MW837769 | [26] | |
| Monodiamesa bonalpicola | MW837770 | [26] | |
| Propsilocerus akamusi | MW846253 | [26] | |
| Propsilocerus taihuensis | MW837766 | [26] | |
| Chironomus kiiensis | ON838253 | [27] | |
| Chironomus plumosus | ON838252 | [27] | |
| Stenochironomus okialbus | OL753645 | [12] | |
| Stenochironomus gibbus | OL742440 | [12] | |
| Polypedilum heberti | OP950225 | [12] | |
| Polypedilum nubifer | MZ747090 | [27] | |
| Cricotopus trifasciatus | OP006250 | [28] | |
| Cricotopus triannulatus | OP006254 | [28] | |
| Thienemanniella tusimufegea | OR333983 | [29] | |
| Thienemanniella triangula | OR333981 | [29] | |
| Ceratopogonidae | Forcipomvia pulchrithorax | NC084322 | [30] |
| Forcipomyia makanensis | MK000395 | [31] |
2.3. Phylogenetic Analyses
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Armitage, P.; Cranston, P.; Pinder, C. The Chironomidae: Biology and Ecology of Non-Biting Midges; Chapman & Hall: London, UK, 1995; 572p. [Google Scholar]
- Karima, Z. Chironomidae: Biology, Ecology and Systematics. In The Wonders of Diptera—Characteristics, Diversity, and Significance for the World’s Ecosystems; IntechOpen: London, UK, 2021; 188p. [Google Scholar] [CrossRef]
- Liu, W.; Chang, T.; Zhao, K.; Sun, X.; Qiao, H.; Yan, C.; Wang, Y. Genome-wide annotation of cuticular protein genes in non-biting midge Propsilocerus akamusi and transcriptome analysis of their response to heavy metal pollution. Int. J. Biol. Macromol. 2022, 223, 555–566. [Google Scholar] [CrossRef] [PubMed]
- Cranston, P.; Hardy, N.; Morse, G. A dated molecular phylogeny for the Chironomidae (Diptera). Syst. Entomol. 2011, 37, 172–188. [Google Scholar] [CrossRef]
- Ashe, P.; O’Connor, J.P. Part 1. Buchonomyiinae, Chilenomyiinae, Podonominae, Aphroteniinae, Tanypodinae, Usambaromyiinae, Diamesinae, Prodiamesinae and Telmatogetoninae. In A World Catalogue of Chironomidae (Diptera); Irish Biogeographical Society & National Museum of Ireland: Dublin, Ireland, 2009; 445p. [Google Scholar]
- Andersen, T.; Ekrem, T.; Cranston, P.S. The larvae of the Holarctic Chironomidae (Diptera)—Introduction. Insect Syst. Evol. Suppl. 2013, 66, 7–12. [Google Scholar]
- Silva, F.L.; Oliveira, C.S. Tanypus urszulae, a new Tanypodinae (Diptera: Chironomidae) from the Neotropical Region. Zootaxa 2016, 4178, 593–600. [Google Scholar] [CrossRef]
- Wang, X. A revised checklist of chironomids from China (Diptera). In Late 20th Century Research on Chironomidae: An Anthology from the 13th International Symposium on Chironomidae; Hoffrichter, O., Ed.; Shaker Verl: Aachen, Germany, 2000; pp. 629–652. [Google Scholar]
- Brown, W.M.; George, M.; Wilson, A.C. Rapid evolution of animal mitochondrial DNA. Proc. Natl. Acad. Sci. USA 1979, 76, 1967–1971. [Google Scholar] [CrossRef] [PubMed]
- Cameron, S. Insect Mitochondrial Genomics: Implications for Evolution and Phylogeny. Annu. Rev. Entomol. 2014, 59, 95–117. [Google Scholar] [CrossRef]
- Lin, X.; Liu, Z.; Yan, L.; Duan, X.; Bu, W.; Wang, X.; Zheng, C. Mitogenomes provide new insights of evolutionary history of Boreheptagyiini and Diamesini (Diptera: Chironomidae: Diamesinae). Ecol. Evol. 2022, 12, e8957. [Google Scholar] [CrossRef]
- Zhang, D.; He, F.; Li, X.; Aishan, Z.; Lin, X. New Mitogenomes of the Polypedilum Generic Complex (Diptera: Chironomidae): Characterization and Phylogenetic Implications. Insects 2023, 14, 238. [Google Scholar] [CrossRef]
- Zheng, C.; Ye, Z.; Zhu, X.; Zhang, H.; Dong, X.; Chen, P.; Bu, W. Integrative taxonomy uncovers hidden species diversity in the rheophilic genus Potamometra (Hemiptera: Gerridae). Zool. Scr. 2019, 49, 174–186. [Google Scholar] [CrossRef]
- Jiang, Y.; Zhao, Y.; Lin, X. First report of the complete mitogenome of Tanypus punctipennis Meigen, 1818 (Diptera, Chironomidae) from Hebei Province, China. Mitochondrial DNA Part B 2022, 7, 215–216. [Google Scholar] [CrossRef]
- Kieffer, J.J. Quelques nouveaux Tendipédides [Dipt.] obtenus d’éclosion (2e note). Bull. De La Soc. Entomol. De Fr. 1912, 4, 101–103. [Google Scholar] [CrossRef]
- Wang, S. New and little-known Chironomidae (Diptera) from the southern provinces of China. Entomotaxonomia 1994, 16, 135–148. [Google Scholar]
- Ge, X.; Zang, H.; Ye, X.; Peng, L.; Wang, B.; Lian, G.; Sun, C. Comparative Mitogenomic Analyses of Hydropsychidae Revealing the Novel Rearrangement of Protein-Coding Gene and tRNA (Trichoptera: Annulipalpia). Insects 2022, 13, 759. [Google Scholar] [CrossRef]
- Ge, X.Y.; Peng, L.; Vogler, P.A.; Morse, J.C.; Yang, L.F.; Sun, C.H.; Wang, B.X. Massive gene rearrangements of mitochondrial genomes and implications for the phylogeny of Trichoptera (Insecta), Trichoptera (Insecta). Syst. Entomol. 2023, 48, 278–295. [Google Scholar] [CrossRef]
- Bolger, A.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Dierckxsens, N.; Mardulyn, P.; Smits, G. NOVOPlasty: De novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 2016, 45, e18. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Le, S.; Li, Y.; Hu, F. SeqKit: A Cross-Platform and Ultrafast Toolkit for FASTA/Q File Manipulation. PLoS ONE 2016, 11, e0163962. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; RamosOnsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA Sequence Polymorphism Analysis of Large Data Sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Zhao, Y.; Yan, L.; Liu, W.; Bu, W.; Wang, X.; Zheng, C. Mitogenomes provide new insights into the evolutionary history of Prodiamesinae (Diptera: Chironomidae). Zool. Scr. 2021, 51, 119–132. [Google Scholar] [CrossRef]
- Fang, X.; Wang, X.; Mao, B.; Xiao, Y.; Shen, M.; Fu, Y. Comparative mitogenome analyses of twelve non-biting flies and provide insights into the phylogeny of Chironomidae (Diptera: Culicomorpha). Sci. Rep. 2023, 13, 9200. [Google Scholar] [CrossRef]
- Li, S.; Chen, M.; Sun, L.; Wang, R.; Li, C.; Gresens, S.; Li, Z.; Lin, X. New mitogenomes from the genus Cricotopus (Diptera: Chironomidae, Orthocladiinae): Characterization and phylogenetic implications. Arch. Insect Biochem. Physiol. 2023, 115, e22067. [Google Scholar] [CrossRef]
- Qi, Y.; Bu, W.; Zheng, C.; Lin, X.; Jiao, K. New data on mitogenomes of Thienemanniella Kieffer, 1911 (Diptera: Chironomidae, Orthocladiinae). Arch. Insect Biochem. Physiol. 2023, 114, 1–9. [Google Scholar] [CrossRef]
- Karademir, G.K.; Teber, S.; Caner Kulig, C.; Toroslu, A.M.; Ibis, O.; YildirimI, A. Complete Mitochondrial Genome Analyses of Forcipomyia pulchrithorax (Diptera: Ceratopogonidae): Genome Orientation and Phylogenetic Implications. Kafkas Univ. Vet. Fak. Derg. 2023, 30, 223–231. [Google Scholar] [CrossRef]
- Jiang, X.; Han, X.; Liu, Q.; Hou, X. The mitochondrial genome of Forcipomyia makanensi (Insecta: Diptera: Ceratopogonidae). Mitochondrial DNA Part B 2019, 4, 344–345. [Google Scholar] [CrossRef]
- Capella-Gutiérrez, S.; Silla-Martínez, J.; Gabaldón, T. trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Kück, P.; Meid, S.A.; Groß, C.; Wägele, J.W.; Misof, B. AliGROOVE—Visualization of heterogeneous sequence divergence within multiple sequence alignments and 6 detection of inflated branch support. BMC Bioinform. 2014, 15, e294. [Google Scholar] [CrossRef]
- Brundin, L. Transantarctic relationships and their significance, as evidenced by chironomid midges with a monograph of the subfamilies Podonominae and Aphroteniinae and the austral Heptagyiae. K. Sven. Vetenskapsakademiens Handl. 1966, 11, 1–472. [Google Scholar]
- Brundin, K.; Sæther, O.A. Buchonomyia burmanica sp.n. and Buchonomyiinae, a new subfamily among the Chironomidae (Diptera). Zool. Scr. 1978, 7, 269–275. [Google Scholar] [CrossRef]
- Murray, D.A.; Ashe, P. A description of the adult female of Buchonomyia thienemanni Fittkau and a reassessment of the phylogenetic position of the subfamily Buchonomyiinae. Spix. Suppl. 1985, 11, 149–160. [Google Scholar]
- Sæther, O.A. Female genitalia in Chironomidae and other Nematocera: Morphology, phylogenies, keys. Bull. Fish. Res. Bd. Can. 1977, 1–209. [Google Scholar]
- Sæther, O.A. Phylogeny of the subfamilies of Chironomidae (Diptera). Syst. Entomol. 2000, 25, 393–403. [Google Scholar] [CrossRef]
- Cranston, P.S.; Hardy, N.B.; Morse, G.E.; Puslednik, L.; McCluen, S.R. When morphology and molecules concur: The ‘Gondwanan’ midges (Diptera: Chironomidae). Syst. Entomol. 2010, 35, 636–648. [Google Scholar] [CrossRef]
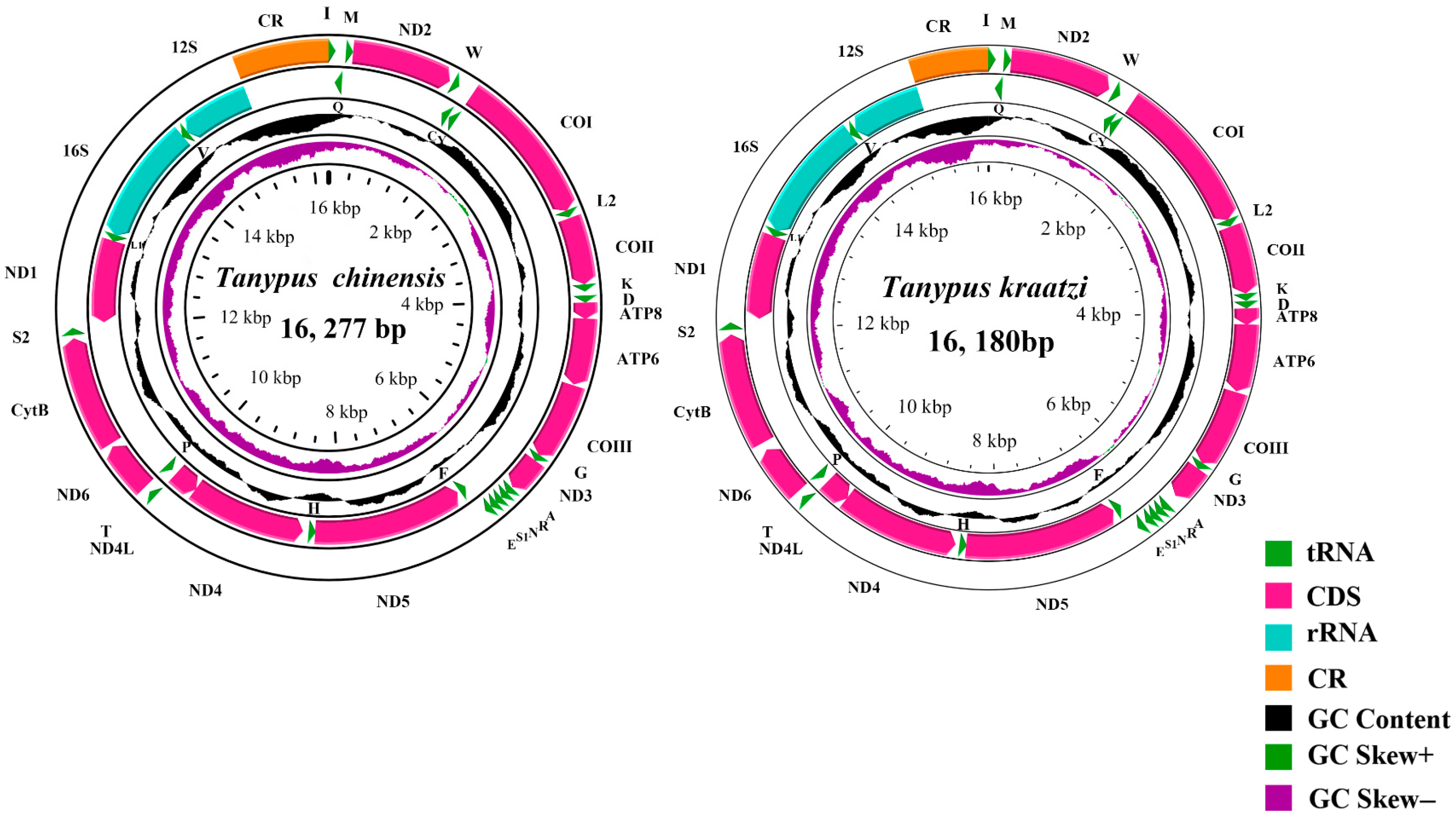
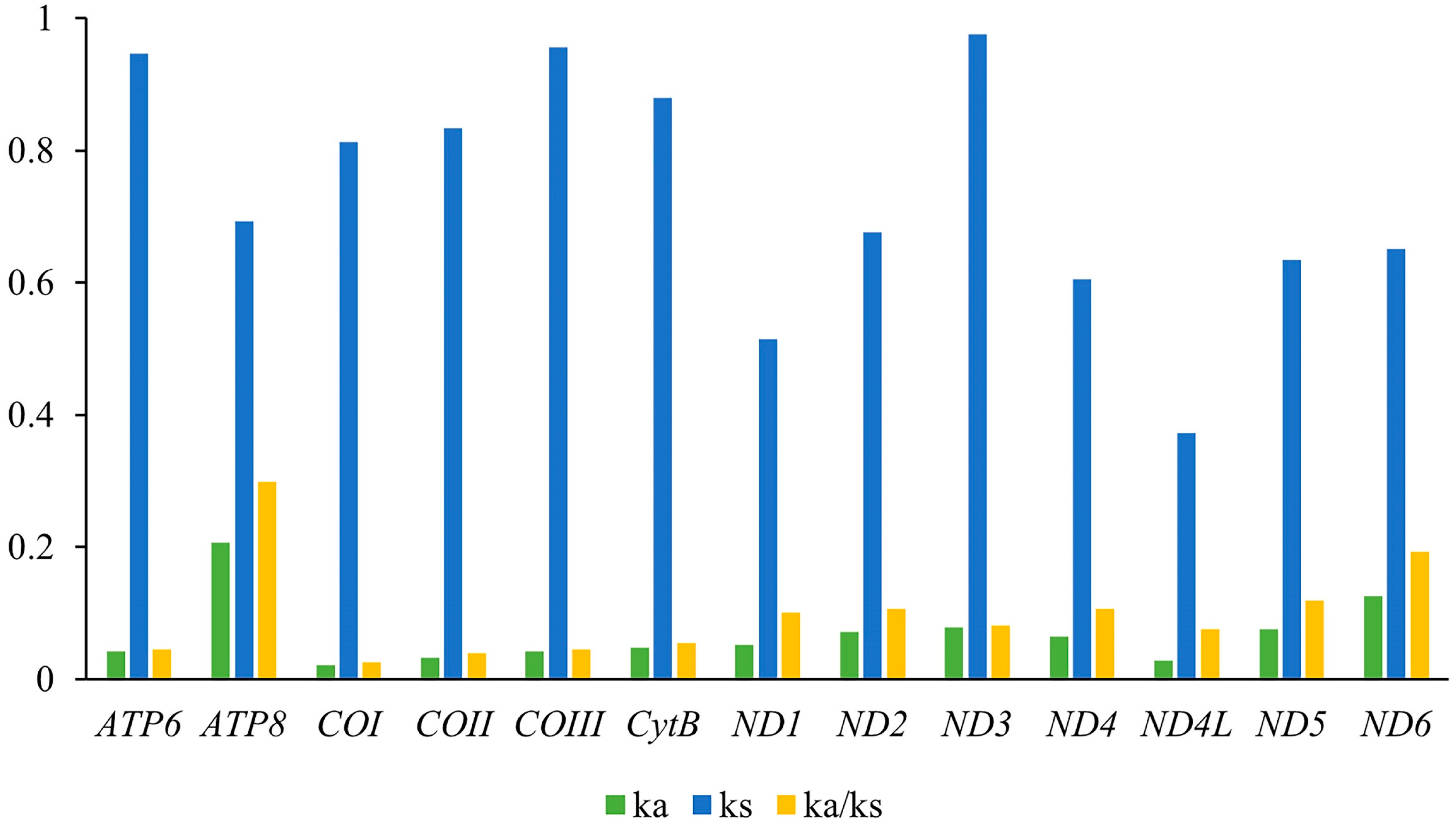

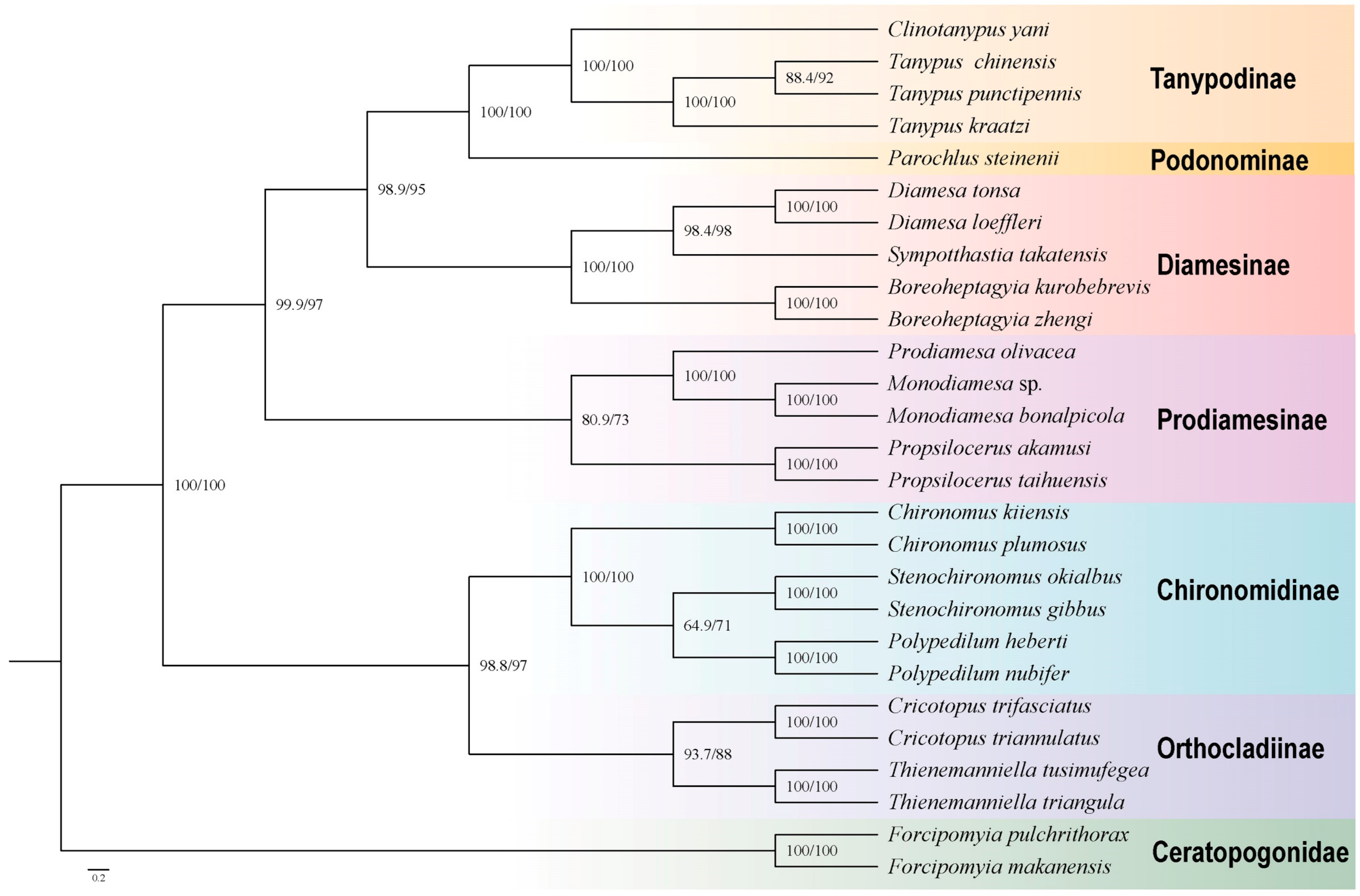
| Gene Type | Length (bp) | Base Composition (%) | Skew | ||||||
|---|---|---|---|---|---|---|---|---|---|
| A | T | C | G | A + T | G + C | AT-Skew | GC-Skew | ||
| Whole genome | 16,266 | 39.61 | 36.27 | 14.55 | 9.57 | 75.88 | 24.12 | 0.044 | −0.206 |
| PCG | 11,216 | 31.28 | 42.73 | 13.45 | 12.54 | 74.00 | 26.00 | −0.155 | −0.035 |
| PCG 1st codon position | 3740 | 31.96 | 36.47 | 12.22 | 19.35 | 68.43 | 31.57 | −0.066 | 0.226 |
| PCG 2nd codon position | 3738 | 20.99 | 45.57 | 19.93 | 13.51 | 66.56 | 33.44 | −0.369 | −0.192 |
| PCG 3rd codon position | 3738 | 40.87 | 46.15 | 8.21 | 4.77 | 87.03 | 12.98 | −0.061 | −0.265 |
| ATP6 | 678 | 32.45 | 41.00 | 15.63 | 10.91 | 73.45 | 26.54 | −0.116 | −0.178 |
| ATP8 | 168 | 42.86 | 39.88 | 12.50 | 4.76 | 82.74 | 17.26 | 0.036 | −0.448 |
| COX1 | 1534 | 28.68 | 37.87 | 17.67 | 15.78 | 66.55 | 33.45 | −0.138 | −0.057 |
| COX2 | 688 | 35.03 | 37.79 | 15.12 | 12.06 | 72.82 | 27.18 | −0.038 | −0.113 |
| COX3 | 789 | 30.54 | 36.88 | 17.74 | 14.83 | 67.42 | 32.57 | −0.094 | −0.089 |
| CYTB | 1137 | 32.63 | 37.03 | 17.77 | 12.58 | 69.66 | 30.35 | −0.063 | −0.171 |
| ND1 | 948 | 24.58 | 49.05 | 9.07 | 17.30 | 73.63 | 26.37 | −0.332 | 0.312 |
| ND2 | 1026 | 32.46 | 45.42 | 12.87 | 9.26 | 77.88 | 22.13 | −0.166 | −0.163 |
| ND3 | 354 | 31.07 | 41.81 | 16.38 | 10.73 | 72.88 | 27.11 | −0.147 | −0.208 |
| ND4 | 1341 | 28.34 | 47.35 | 8.58 | 15.73 | 75.69 | 24.31 | −0.251 | 0.294 |
| ND4L | 294 | 27.55 | 52.04 | 6.80 | 13.61 | 79.59 | 20.41 | −0.308 | 0.334 |
| ND5 | 1734 | 28.43 | 45.50 | 9.69 | 16.38 | 73.93 | 26.07 | −0.231 | 0.257 |
| ND6 | 525 | 34.48 | 46.48 | 12.19 | 6.86 | 80.96 | 19.05 | −0.148 | −0.280 |
| All rRNA | 2202 | 37.30 | 42.91 | 6.73 | 13.07 | 80.21 | 19.79 | −0.070 | 0.320 |
| 12S | 807 | 36.68 | 42.38 | 7.43 | 13.51 | 79.06 | 20.94 | −0.072 | 0.290 |
| 16S | 1395 | 37.92 | 43.44 | 6.02 | 12.62 | 81.36 | 18.64 | −0.068 | 0.354 |
| CR | 952 | 47.16 | 43.59 | 7.14 | 2.10 | 90.75 | 9.24 | 0.039 | −0.545 |
| tRNA | 1503 | 38.59 | 37.59 | 9.98 | 13.84 | 76.18 | 23.82 | 0.013 | 0.162 |
| Gene Type | Length (bp) | Base Composition (%) | Skew | ||||||
|---|---|---|---|---|---|---|---|---|---|
| A | T | C | G | A + T | G + C | AT-Skew | GC-Skew | ||
| Whole genome | 16,180 | 39.10 | 36.51 | 14.33 | 10.06 | 75.61 | 24.39 | 0.034 | −0.175 |
| PCG | 11,216 | 31.13 | 43.19 | 12.98 | 12.70 | 74.32 | 25.68 | −0.162 | −0.011 |
| PCG 1st codon position | 3740 | 32.55 | 37.08 | 11.56 | 18.82 | 69.62 | 30.38 | −0.065 | 0.239 |
| PCG 2nd codon position | 3738 | 20.63 | 45.64 | 19.78 | 13.95 | 66.27 | 33.73 | −0.377 | −0.173 |
| PCG 3rd codon position | 3738 | 40.20 | 46.87 | 7.60 | 5.33 | 87.07 | 12.93 | −0.077 | −0.176 |
| ATP6 | 678 | 29.94 | 40.27 | 16.67 | 13.13 | 70.21 | 29.80 | −0.147 | −0.119 |
| ATP8 | 168 | 41.67 | 38.69 | 12.50 | 7.14 | 80.36 | 19.64 | 0.037 | −0.273 |
| COX1 | 1534 | 29.01 | 37.09 | 17.99 | 15.91 | 66.10 | 33.90 | −0.122 | −0.061 |
| COX2 | 688 | 34.74 | 36.63 | 15.84 | 12.79 | 71.37 | 28.63 | −0.026 | −0.107 |
| COX3 | 789 | 29.66 | 37.90 | 16.86 | 15.59 | 67.56 | 32.45 | −0.122 | −0.039 |
| CYTB | 1137 | 31.75 | 37.99 | 17.24 | 13.02 | 69.74 | 30.26 | −0.089 | −0.139 |
| ND1 | 948 | 24.58 | 49.05 | 8.97 | 17.41 | 73.63 | 26.38 | −0.332 | 0.320 |
| ND2 | 1026 | 32.46 | 45.22 | 12.48 | 9.84 | 77.68 | 22.32 | −0.164 | −0.118 |
| ND3 | 354 | 31.64 | 41.81 | 15.25 | 11.30 | 73.45 | 26.55 | −0.138 | −0.149 |
| ND4 | 1341 | 28.11 | 47.20 | 9.40 | 15.29 | 75.31 | 24.69 | −0.253 | 0.239 |
| ND4L | 294 | 25.85 | 52.38 | 8.16 | 13.61 | 78.23 | 21.77 | −0.339 | 0.250 |
| ND5 | 1734 | 29.01 | 45.67 | 9.57 | 15.74 | 74.68 | 25.31 | −0.223 | 0.244 |
| ND6 | 525 | 33.71 | 48.95 | 10.67 | 6.67 | 82.66 | 17.34 | −0.184 | −0.231 |
| All rRNA | 2193 | 38.06 | 42.08 | 7.17 | 12.70 | 80.13 | 19.87 | −0.050 | 0.279 |
| 12S | 811 | 37.98 | 40.81 | 7.89 | 13.32 | 78.79 | 21.21 | −0.036 | 0.256 |
| 16S | 1382 | 38.13 | 43.34 | 6.44 | 12.08 | 81.47 | 18.52 | −0.064 | 0.305 |
| CR | 781 | 44.81 | 46.73 | 6.40 | 2.05 | 91.54 | 8.45 | −0.021 | −0.515 |
| tRNA | 1508 | 37.86 | 38.46 | 10.21 | 13.46 | 76.32 | 23.67 | −0.008 | 0.137 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gao, S.; Wang, C.; Tang, Y.; Zhang, Y.; Ge, X.; Zhang, J.; Liu, W. Complete Mitochondrial Genome of Tanypus chinensis and Tanypus kraatzi (Diptera: Chironomidae): Characterization and Phylogenetic Implications. Genes 2024, 15, 1281. https://doi.org/10.3390/genes15101281
Gao S, Wang C, Tang Y, Zhang Y, Ge X, Zhang J, Liu W. Complete Mitochondrial Genome of Tanypus chinensis and Tanypus kraatzi (Diptera: Chironomidae): Characterization and Phylogenetic Implications. Genes. 2024; 15(10):1281. https://doi.org/10.3390/genes15101281
Chicago/Turabian StyleGao, Shaobo, Chengyan Wang, Yaning Tang, Yuzhen Zhang, Xinyu Ge, Jiwei Zhang, and Wenbin Liu. 2024. "Complete Mitochondrial Genome of Tanypus chinensis and Tanypus kraatzi (Diptera: Chironomidae): Characterization and Phylogenetic Implications" Genes 15, no. 10: 1281. https://doi.org/10.3390/genes15101281
APA StyleGao, S., Wang, C., Tang, Y., Zhang, Y., Ge, X., Zhang, J., & Liu, W. (2024). Complete Mitochondrial Genome of Tanypus chinensis and Tanypus kraatzi (Diptera: Chironomidae): Characterization and Phylogenetic Implications. Genes, 15(10), 1281. https://doi.org/10.3390/genes15101281







