Expression and Characterization of an Efficient Alginate Lyase from Psychromonas sp. SP041 through Metagenomics Analysis of Rotten Kelp
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Metagenomic Screening and Cloning of Alginate Lyase Gene
2.3. Structure Prediction of Alg169
2.4. Expression and Purification of Alg169
2.5. Enzyme Activity Assay
2.6. Characterization of Recombinant Alg169
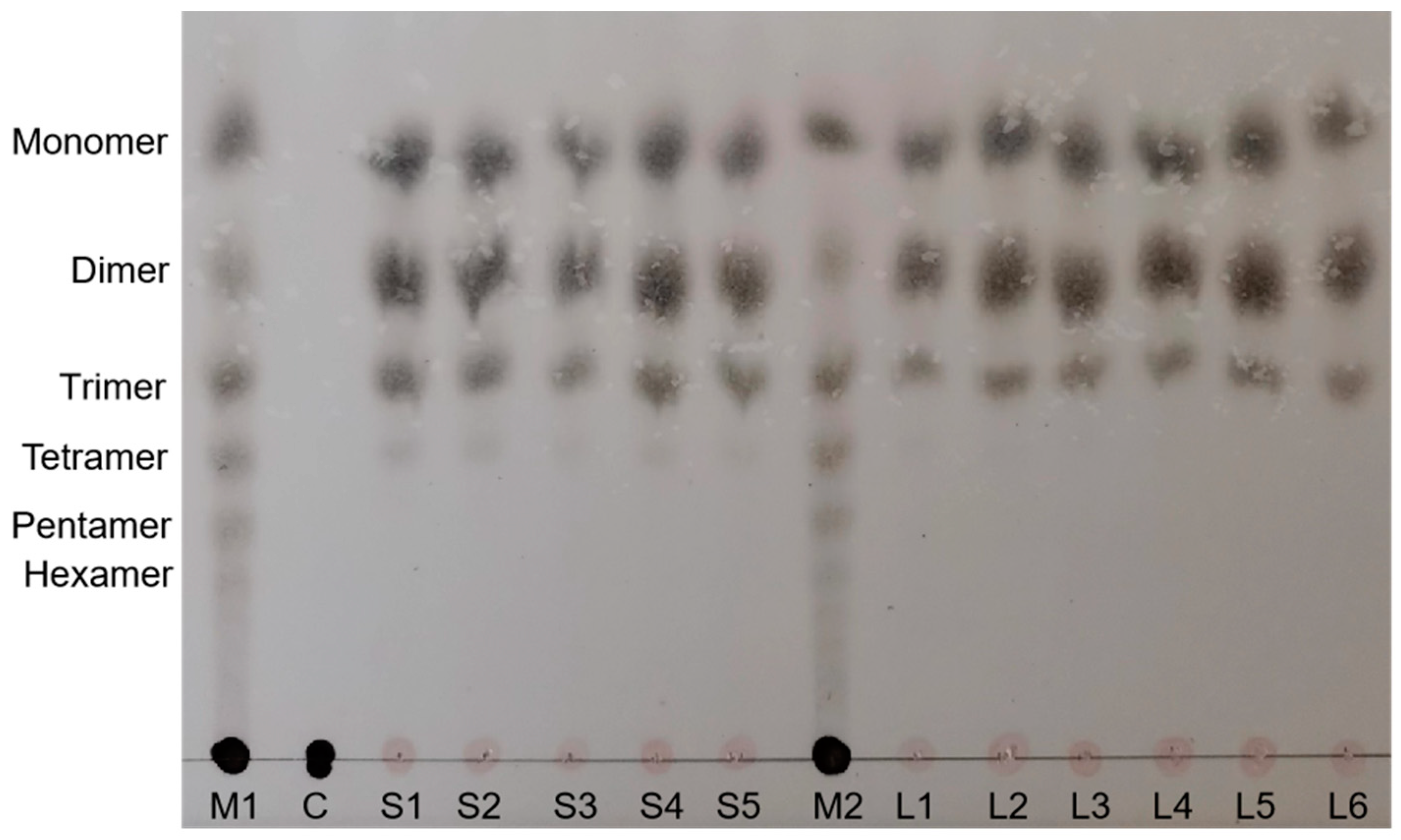
2.7. Analysis of the Enzymatic Hydrolysis Products of Alginate
3. Results
3.1. Metagenomic Data Analysis
3.2. Sequence Analysis and Molecular Docking of Alg169
3.3. Analysis of the Three-Dimensional Structure of Alg169
3.4. Expression and Purification of Alg169
3.5. The Optimum Temperature and Temperature Stability of Alg169
3.6. The Optimum pH and pH Stability of Alg169
3.7. The Effect of Metal Ions on Alg169 Enzyme Activity
3.8. The Effect of NaCl Concentration on the Activity of Alg169
3.9. Enzyme Kinetic Parameters and Products of Alg169
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Wu, Y.; Jin, X.; Zhang, Y.; Liu, J.; Wu, M.; Tong, H. Bioactive Compounds from Brown Algae Alleviate Nonalcoholic Fatty Liver Disease: An Extensive Review. J. Agric. Food Chem. 2023, 71, 1771–1787. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Wang, Y.; Qin, Y.M.; Shen, P.L.; Peng, Q. Structures, properties and application of alginic acid: A review. Int. J. Biol. Macromol. 2020, 162, 618–628. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Yin, H. Alginate lyase: Review of major sources and classification, properties, structure-function analysis and applications. Bioengineered 2015, 6, 125–131. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.X.; Yao, L.Y.; Nie, S.; Xu, Y. Low-viscosity sodium alginate combined with TiO(2) nanoparticles for improving neuroblastoma treatment. Int. J. Biol. Macromol. 2021, 167, 921–933. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.F.; Lai, S.S.; Chen, Y.H.; Liu, D.; Liu, B.; Ai, C.; Wan, X.Z.; Gao, L.Y.; Chen, X.H.; Zhao, C. Anti-diabetic effect of oligosaccharides from seaweed Sargassum confusum via JNK-IRS1/PI3K signalling pathways and regulation of gut microbiota. Food Chem. Toxicol. 2019, 131, 110562. [Google Scholar] [CrossRef]
- Zhang, P.F.; Liu, J.; Xiong, B.H.; Zhang, C.; Kang, B.N.; Gao, Y.S.; Li, Z.K.; Ge, W.; Cheng, S.F.; Hao, Y.N.; et al. Microbiota from alginate oligosaccharide-dosed mice successfully mitigated small intestinal mucositis. Microbiome 2020, 8, 112. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.Y.; Sun, G.Q.; Feng, T.; Zhang, J.; Huang, X.; Wang, T.; Xie, Z.Q.; Chu, X.K.; Yang, J.; Wang, H.; et al. Sodium oligomannate therapeutically remodels gut microbiota and suppresses gut bacterial amino acids-shaped neuroinflammation to inhibit Alzheimer’s disease progression. Cell Res. 2019, 29, 787–803. [Google Scholar] [CrossRef] [PubMed]
- Ueno, M.; Cho, K.; Nakazono, S.; Isaka, S.; Abu, R.; Takeshita, S.; Yamaguchi, K.; Kim, D.; Oda, T. Alginate oligomer induces nitric oxide (NO) production in RAW264.7 cells: Elucidation of the underlying intracellular signaling mechanism. Biosci. Biotechnol. Biochem. 2015, 79, 1787–1793. [Google Scholar] [CrossRef]
- Jiang, Z.D.; Zhang, X.W.; Wu, L.Y.; Li, H.B.; Chen, Y.H.; Li, L.J.; Ni, H.; Li, Q.B.; Zhu, Y.B. Exolytic products of alginate by the immobilized alginate lyase confer antioxidant and antiapoptotic bioactivities in human umbilical vein endothelial cells. Carbohydr. Polym. 2021, 251, 116976. [Google Scholar] [CrossRef]
- Wang, Y.T.; Li, L.L.; Ye, C.Q.; Yuan, J.Y.; Qin, S. Alginate oligosaccharide improves lipid metabolism and inflammation by modulating gut microbiota in high-fat diet fed mice. Appl. Microbiol. Biotechnol. 2020, 104, 3541–3554. [Google Scholar] [CrossRef]
- Yan, S.; Zhu, Y.; Li, L.; Qin, S.; Yuan, J.; Chang, X.; Hu, S. Alginate oligosaccharide ameliorates azithromycin-induced gut microbiota disorder via Bacteroides acidifaciens-FAHFAs and Bacteroides-TCA cycle axes. Food Funct. 2023, 14, 427–444. [Google Scholar] [CrossRef] [PubMed]
- Bi, D.C.; Yang, X.; Lu, J.; Xu, X. Preparation and potential applications of alginate oligosaccharides. Crit. Rev. Food Sci. Nutr. 2023, 63, 10130–10147. [Google Scholar] [CrossRef] [PubMed]
- Abd El-Mohdy, H.L. Radiation-induced degradation of sodium alginate and its plant growth promotion effect. Arab. J. Chem. 2017, 10, 431–438. [Google Scholar] [CrossRef]
- Aida, T.M.; Yamagata, T.; Watanabe, M.; Smith, R.L. Depolymerization of sodium alginate under hydrothermal conditions. Carbohyd. Polym. 2010, 80, 296–302. [Google Scholar] [CrossRef]
- Rodrigues, D.; Sousa, S.; Silva, A.; Amorim, M.; Pereira, L.; Rocha-Santos, T.A.P.; Gomes, A.M.P.; Duarte, A.C.; Freitas, A.C. Impact of enzyme- and ultrasound-assisted extraction methods on biological properties of red, brown, and green seaweeds from the central west coast of portugal. J. Agric. Food Chem. 2015, 63, 3177–3188. [Google Scholar] [CrossRef] [PubMed]
- Zimoch-Korzycka, A.; Kulig, D.; Król-Kilińska, Ż.; Żarowska, B.; Bobak, Ł.; Jarmoluk, A. Biophysico-chemical properties of alginate oligomers obtained by acid and oxidation depolymerization. Polymers 2021, 13, 2558. [Google Scholar] [CrossRef]
- Mao, S.R.; Zhang, T.T.; Sun, W.; Ren, X.H. The depolymerization of sodium alginate by oxidative degradation. Pharm. Dev. Technol. 2012, 17, 763–769. [Google Scholar] [CrossRef] [PubMed]
- Boucelkha, A.; Petit, E.; Elboutachfaiti, R.; Molinie, R.; Amari, S.; Yahaoui, R.Z. Production of guluronate oligosaccharide of alginate from brown algae Stypocaulon scoparium using an alginate lyase. J. Appl. Phycol. 2017, 29, 509–519. [Google Scholar] [CrossRef]
- Li, S.Y.; Wang, Z.P.; Wang, L.N.; Peng, J.X.; Wang, Y.N.; Han, Y.T.; Zhao, S.F. Combined enzymatic hydrolysis and selective fermentation for green production of alginate oligosaccharides from Laminaria japonica. Bioresour. Technol. 2019, 281, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Liu, S.; Huang, J.; Ren, K.; Zhu, Y.; Yang, S. Alginate: Microbial production, functionalization, and biomedical applications. Int. J. Biol. Macromol. 2023, 242, 125048. [Google Scholar] [CrossRef]
- Chen, K.; Pachter, L. Bioinformatics for whole-genome shotgun sequencing of microbial communities. PLoS Comput. Biol. 2005, 1, 106–112. [Google Scholar] [CrossRef] [PubMed]
- Ariaeenejad, S.; Gharechahi, J.; Foroozandeh Shahraki, M.; Fallah Atanaki, F.; Han, J.L.; Ding, X.Z.; Hildebrand, F.; Bahram, M.; Kavousi, K.; Hosseini Salekdeh, G. Precision enzyme discovery through targeted mining of metagenomic data. Nat. Prod. Bioprospect 2024, 14, 7. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.S.; Lee, C.G.; Lee, E.Y. Alginate lyase: Structure, property, and application. Biotechnol. Bioprocess Eng. 2011, 16, 843–851. [Google Scholar] [CrossRef]
- Sun, H.H.; Gao, L.; Xue, C.H.; Mao, X.Z. Marine-polysaccharide degrading enzymes: Status and prospects. Compr. Rev. Food Sci. Food Saf. 2020, 19, 2767–2796. [Google Scholar] [CrossRef]
- Helbert, W.; Poulet, L.; Drouillard, S.; Mathieu, S.; Loiodice, M.; Couturier, M.; Lombard, V.; Terrapon, N.; Turchetto, J.; Vincentelli, R.; et al. Discovery of novel carbohydrate-active enzymes through the rational exploration of the protein sequences space. Proc. Natl. Acad. Sci. USA 2019, 116, 6063–6068. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Zhu, Y.M.; Men, Y.; Zeng, Y.; Sun, Y.X. Purification and characterization of a novel alginate lyase from the marine bacterium Bacillus sp. Alg07. Mar. Drugs 2018, 16, 86. [Google Scholar] [CrossRef] [PubMed]
- De Santi, C.; Tedesco, P.; Ambrosino, L.; Altermark, B.; Willassen, N.P.; de Pascale, D. A new alkaliphilic cold-active esterase from the psychrophilic marine bacterium Rhodococcus sp.: Functional and structural studies and biotechnological potential. Appl. Biochem. Biotechnol. 2014, 172, 3054–3068. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.; Ni, F.; Ning, L.; Sun, Y.; Yao, Z. Cloning and characterization of a new pH-stable alginate lyase with high salt tolerance from marine Vibrio sp. NJ-04. Int. J. Biol. Macromol. 2018, 115, 1063–1070. [Google Scholar] [CrossRef]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar]
- Almagro Armenteros, J.J.; Tsirigos, K.D.; Sønderby, C.K.; Petersen, T.N.; Winther, O.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 2019, 37, 420–423. [Google Scholar] [CrossRef]
- Lu, S.; Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; Gwadz, M.; Hurwitz, D.I.; Marchler, G.H.; Song, J.S.; et al. CDD/SPARCLE: The conserved domain database in 2020. Nucleic Acids Res. 2020, 48, 265–268. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Zhang, X.; Tang, Y.; Gao, F.; Xu, X.; Chen, G.; Li, Y.; Wang, L. Low-cost and efficient strategy for brown algal hydrolysis: Combination of alginate lyase and cellulase. Bioresour. Technol. 2024, 397, 130481. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Wang, P.; Zhang, Y.Z.; Chen, X.L. Diversity of three-dimensional structures and catalytic mechanisms of alginate lyases. Appl. Environ. Microbiol. 2018, 84, e02040-17. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, M.; Ogura, K.; Hashimoto, W.; Mikami, B.; Murata, K. A structural basis for depolymerization of alginate by polysaccharide lyase family-7. J. Mol. Biol. 2005, 352, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Colovos, C.; Yeates, T.O. Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Sci. 1993, 2, 1511–1519. [Google Scholar] [CrossRef] [PubMed]
- Garron, M.L.; Cygler, M. Structural and mechanistic classification of uronic acid-containing polysaccharide lyases. Glycobiology 2010, 20, 1547–1573. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.H.; Sun, X.H.; Chen, X.L.; Li, P.Y.; Qin, Q.L.; Zhang, Y.Q.; Xu, F. Synergy of the two alginate lyase domains of a novel alginate lyase from Vibrio sp. NC2 in alginate degradation. Appl. Environ. Microbiol. 2022, 88, e0155922. [Google Scholar] [CrossRef]
- Sun, X.M.; Xue, Z.; Sun, M.L.; Zhang, Y.; Zhang, Y.Z.; Fu, H.H.; Zhang, Y.Q.; Wang, P. Characterization of a novel alginate lyase with two alginate lyase domains from the marine bacterium Vibrio sp. C42. Mar. Drugs 2022, 20, 746. [Google Scholar] [CrossRef]
- Gu, X.; Fu, L.; Wang, Z.; Cao, Z.; Zhao, L.; Seswita-Zilda, D.; Zhang, A.; Zhang, Q.; Li, J. Novel bifunctional alginate lyase and antioxidant activity of the enzymatic hydrolysates. J. Agric. Food Chem. 2024, 72, 4116–4126. [Google Scholar] [CrossRef]
- Li, L.; Cao, S.S.; Zhu, B.W.; Yao, Z.; Zhu, B.; Qin, Y.M.; Jiang, J.J. Efficient degradation of alginate and preparation of alginate oligosaccharides by a novel biofunctional alginate lyase with high activity and excellent thermophilic features. Mar. Drugs 2023, 21, 180. [Google Scholar] [CrossRef] [PubMed]
- Inoue, A.; Takadono, K.; Nishiyama, R.; Tajima, K.; Kobayashi, T.; Ojima, T. Characterization of an alginate lyase, FlAlyA, from Flavobacterium sp. strain UMI-01 and its expression in Escherichia coli. Mar. Drugs 2014, 12, 4693–4712. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Chen, X.L.; Sun, X.H.; Dong, F.; Li, C.Y.; Li, P.Y.; Ding, H.; Chen, Y.; Zhang, Y.Z.; Wang, P. Structural and molecular basis for the substrate positioning mechanism of a new PL7 subfamily alginate lyase from the arctic. J. Biol. Chem. 2020, 295, 16380–16392. [Google Scholar] [CrossRef] [PubMed]
- Peng, C.N.; Wang, Q.B.; Lu, D.R.; Han, W.J.; Li, F.C. A novel bifunctional endolytic alginate lyase with variable alginate-degrading modes and versatile monosaccharide-producing properties. Front. Microbiol. 2018, 9, 167. [Google Scholar] [CrossRef] [PubMed]
- Shu, Z.; Wang, G.; Liu, F.; Xu, Y.; Sun, J.; Hu, Y.; Dong, H.; Zhang, J. Genome sequencing-based mining and characterization of a novel alginate lyase from Vibrio alginolyticus S10 for specific production of disaccharides. Mar. Drugs 2023, 21, 564. [Google Scholar] [CrossRef] [PubMed]
- Zhu, B.W.; Tan, H.D.; Qin, Y.Q.; Xu, Q.S.; Du, Y.G.; Yin, H. Characterization of a new endo-type alginate lyase from Vibrio sp. W13. Int. J. Biol. Macromol. 2015, 75, 330–337. [Google Scholar] [CrossRef] [PubMed]
- Xing, M.; Cao, Q.; Wang, Y.; Xiao, H.; Zhao, J.; Zhang, Q.; Ji, A.; Song, S. Advances in research on the bioactivity of alginate oligosaccharides. Mar. Drugs 2020, 18, 144. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.X.; Xu, S.S.; Yin, X.J.; Wang, F.L.; Li, Y. Characterization of a new bifunctional and cold-adapted polysaccharide lyase (PL) family 7 alginate lyase from Flavobacterium sp. Mar. Drugs 2020, 18, 388. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Qin, S.; Zhai, S.; Gao, Y.; Li, L. Inulin with different degrees of polymerization modulates composition of intestinal microbiota in mice. FEMS Microbiol. Lett. 2017, 364, fnx075. [Google Scholar] [CrossRef]
- Li, L.; Zhu, B.; Yao, Z.; Jiang, J. Directed preparation, structure-activity relationship and applications of alginate oligosaccharides with specific structures: A systematic review. Food Res. Int. 2023, 170, 112990. [Google Scholar] [CrossRef]
- Robinson, S.L.; Piel, J.; Sunagawa, S. A roadmap for metagenomic enzyme discovery. Nat. Prod. Rep. 2021, 38, 1994–2023. [Google Scholar] [CrossRef] [PubMed]
- Nam, N.N.; Do, H.D.K.; Loan Trinh, K.T.; Lee, N.Y. Metagenomics: An effective approach for exploring microbial diversity and functions. Foods 2023, 12, 2140. [Google Scholar] [CrossRef] [PubMed]










| Substrate | Sodium Alginate | polyG | polyM |
|---|---|---|---|
| Km (mg/mL) | 7.23 | 0.53 | 1.29 |
| Vmax (μg/min) | 249.30 | 104.10 | 71.15 |
| kcat (s−1) | 2678.63 | 2082.00 | 1423.00 |
| kcat/Km (s−1/(mg/mL)) | 370.49 | 3928.30 | 1103.10 |
| Alginate Lyase | Specific Activity (U/mg) | Optimal Temperature (°C) | pH | Cation Activators | Source | Reference |
|---|---|---|---|---|---|---|
| Alg169 | 117,081 | 25 | 7.0 | Mn2+, Co2+, Ca2+, Mg2+, Ni2+, Ba2+, Cu2+, Zn2+, Na+, K+ | Psychromonas sp. SP041 | This study |
| AlyRm3 | 37,315 | 70 | 8.0 | Na+, K+, Ca2+, Mg2+, Co2+, Fe3+ | Rhodothermus marinus DSM 4252 | [41] |
| FlAlyA | 23,478 | 55 | 7.7 | Na+, K+, Mg2+, Ca2+, EGTA | Flavobacterium sp. strain UMI-01 | [42] |
| AlgA | 8306 | 40 | 7.5 | Na+, Mg2+, Ca2+ | Acillus weihaiensis Alg07 | [26] |
| AlyC3 | 6000 | 20 | 8.0 | Na+ | Psychromonas sp.C-3 | [43] |
| AlgNJ-04 | 2416 | 40 | 7.0 | Na+, K+, Ca2+ | Vibrio sp. NJ-04 | [28] |
| Aly2 | 2025 | 40 | 6.0 | EDTA, glycerol, 2-mercaptoethanol | Flammeovirga sp. MY04 | [44] |
| Alg4755 | 961 | 35 | 8.0 | Al3+, Co2+, Ba2+, Fe3+, Ni2+, Na+ | Vibrio alginolyticus S10 | [45] |
| Algb | 457 | 30 | 8.0 | Na+, Ca2+, Co2+, Fe2+ | Vibrio sp. W13 | [46] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, P.; Cai, Y.; Zhong, H.; Chen, R.; Yi, Y.; Ye, Y.; Li, L. Expression and Characterization of an Efficient Alginate Lyase from Psychromonas sp. SP041 through Metagenomics Analysis of Rotten Kelp. Genes 2024, 15, 598. https://doi.org/10.3390/genes15050598
Wang P, Cai Y, Zhong H, Chen R, Yi Y, Ye Y, Li L. Expression and Characterization of an Efficient Alginate Lyase from Psychromonas sp. SP041 through Metagenomics Analysis of Rotten Kelp. Genes. 2024; 15(5):598. https://doi.org/10.3390/genes15050598
Chicago/Turabian StyleWang, Ping, Yi Cai, Hua Zhong, Ruiting Chen, Yuetao Yi, Yanrui Ye, and Lili Li. 2024. "Expression and Characterization of an Efficient Alginate Lyase from Psychromonas sp. SP041 through Metagenomics Analysis of Rotten Kelp" Genes 15, no. 5: 598. https://doi.org/10.3390/genes15050598
APA StyleWang, P., Cai, Y., Zhong, H., Chen, R., Yi, Y., Ye, Y., & Li, L. (2024). Expression and Characterization of an Efficient Alginate Lyase from Psychromonas sp. SP041 through Metagenomics Analysis of Rotten Kelp. Genes, 15(5), 598. https://doi.org/10.3390/genes15050598








