Characterization and Phylogenetic Analysis of the Chloroplast Genomes of Stephania japonica var. timoriensis and Stephania japonica var. discolor
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection, DNA Extraction, and Sequencing
2.2. CpDNA Assembly and Annotation
2.3. Codon Preference Analysis and Repeat Sequence Analysis
2.4. Comparative Genomics Analysis of the Chloroplast Genome
2.5. Phylogenetic Analysis
3. Results
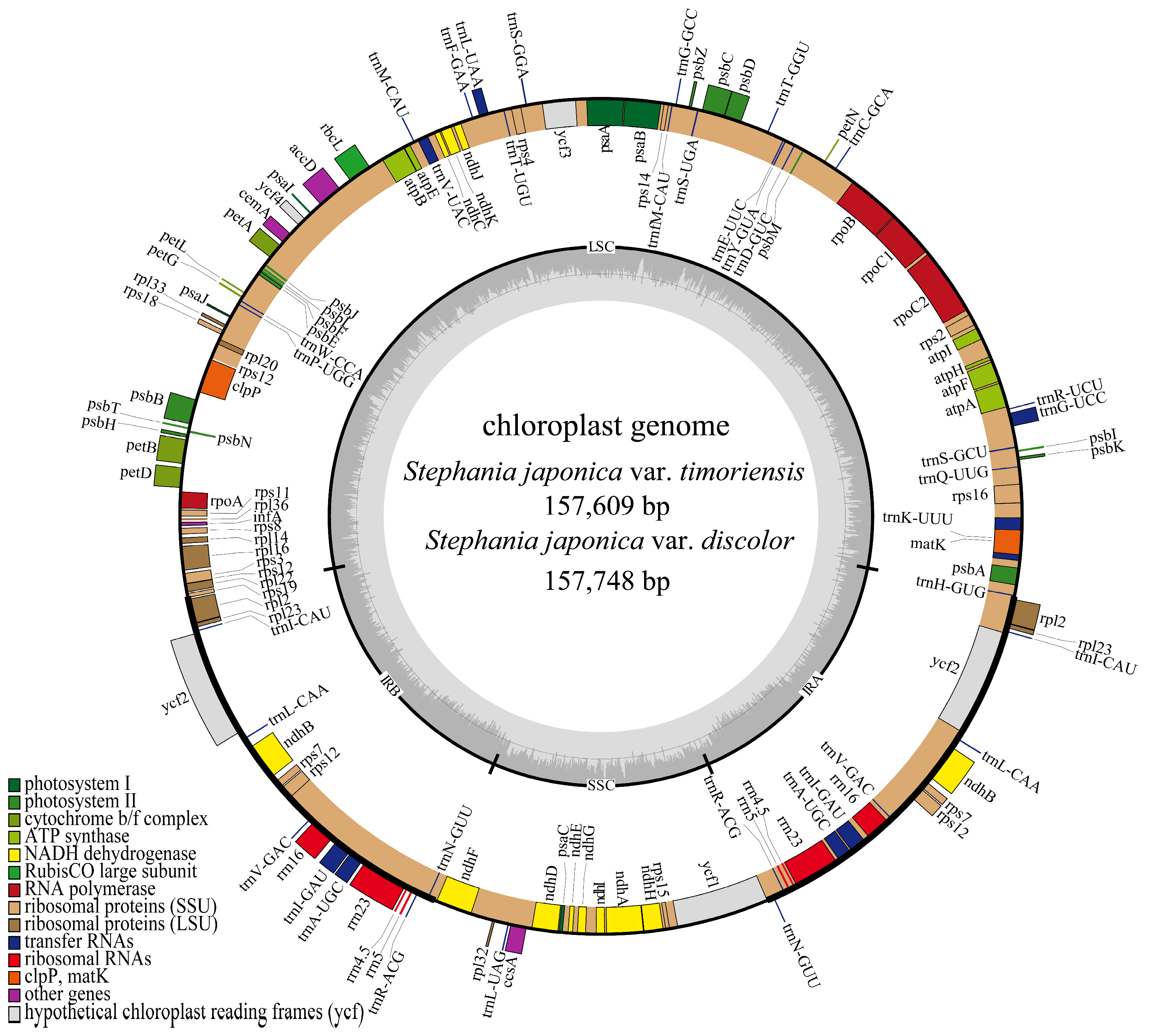
3.1. Analysis of Characteristics of Chloroplast Genomes
3.2. Codon Usage Bias
3.3. Repeat Sequences and SSR Analysis
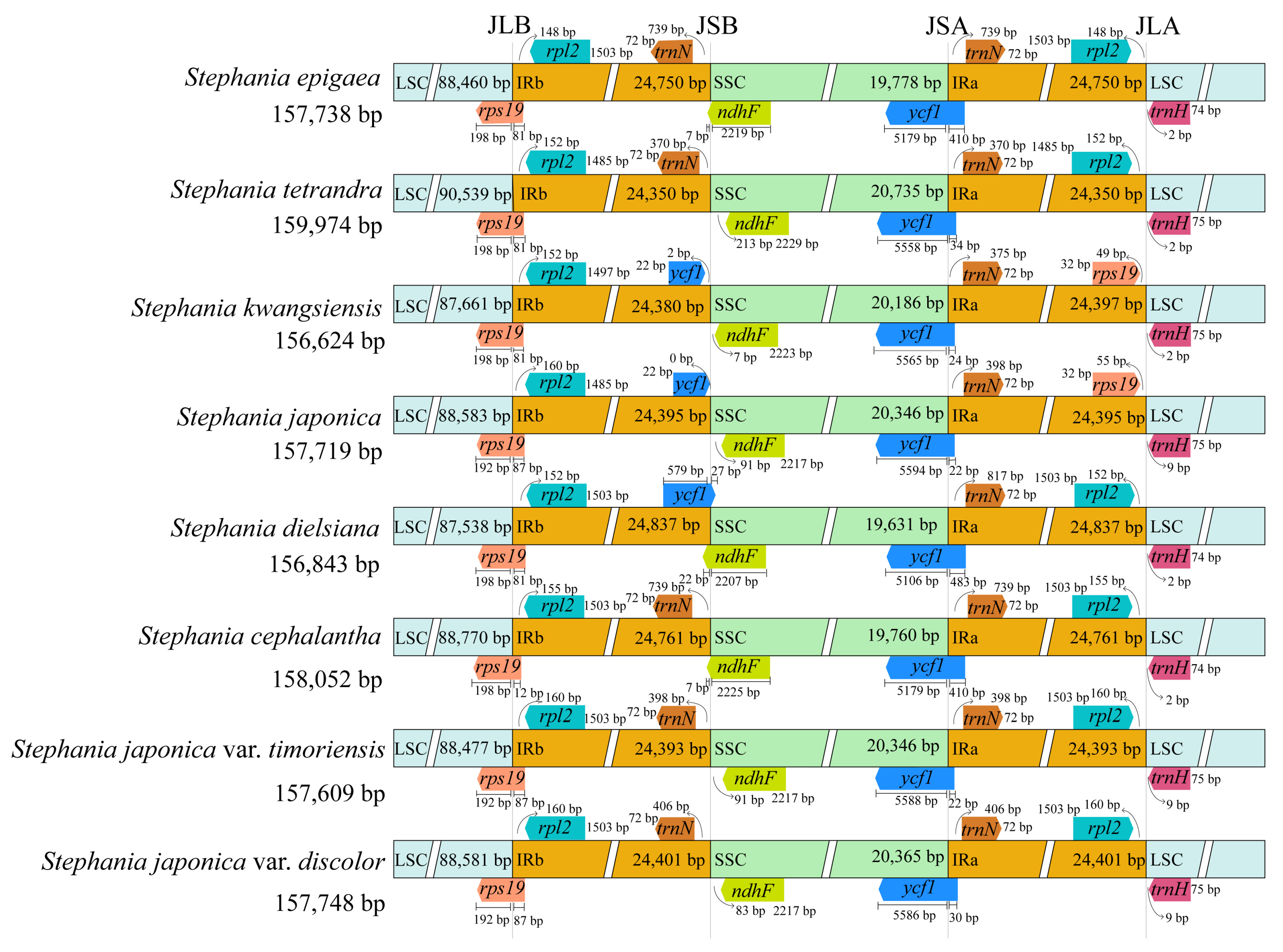
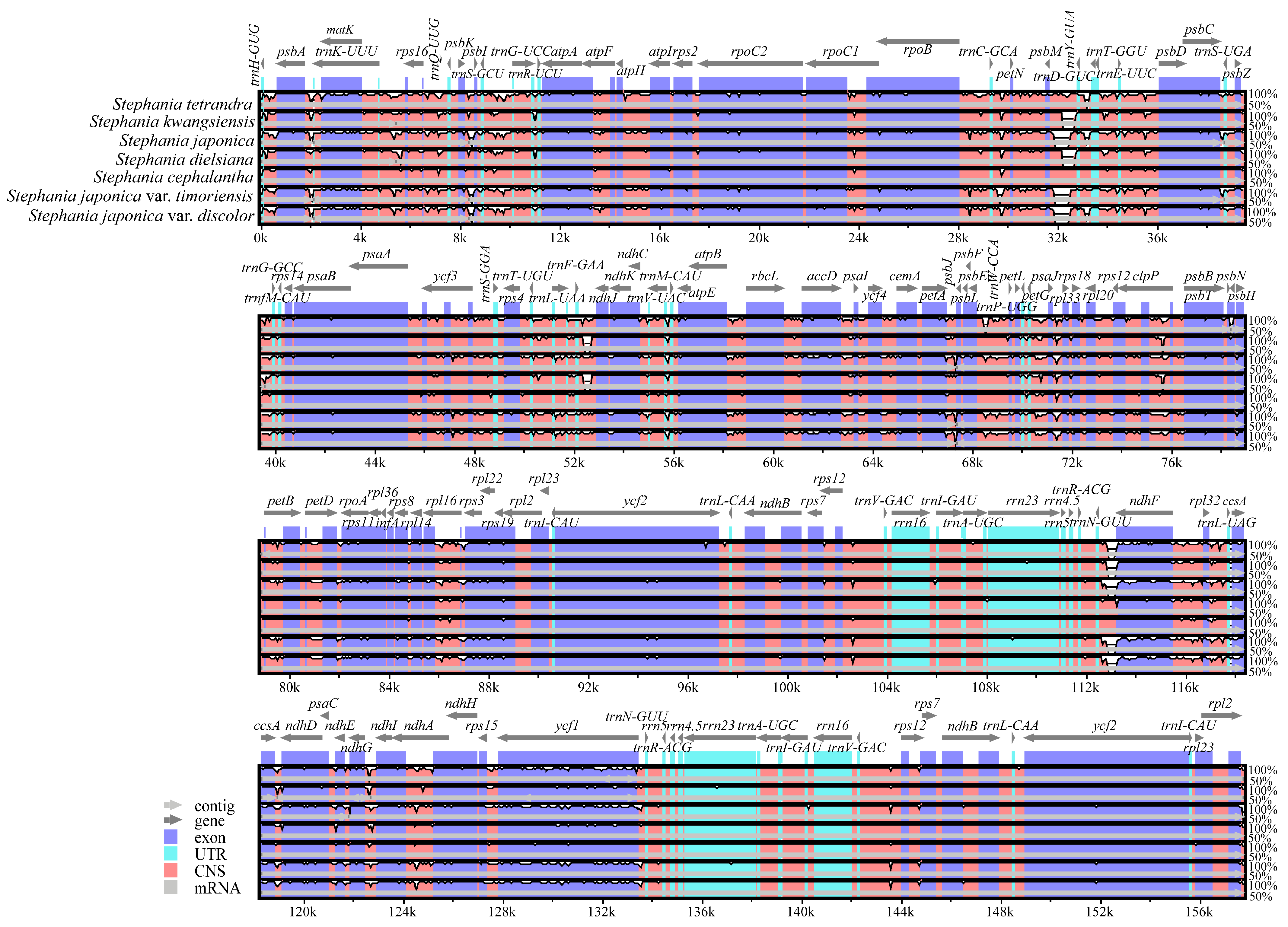
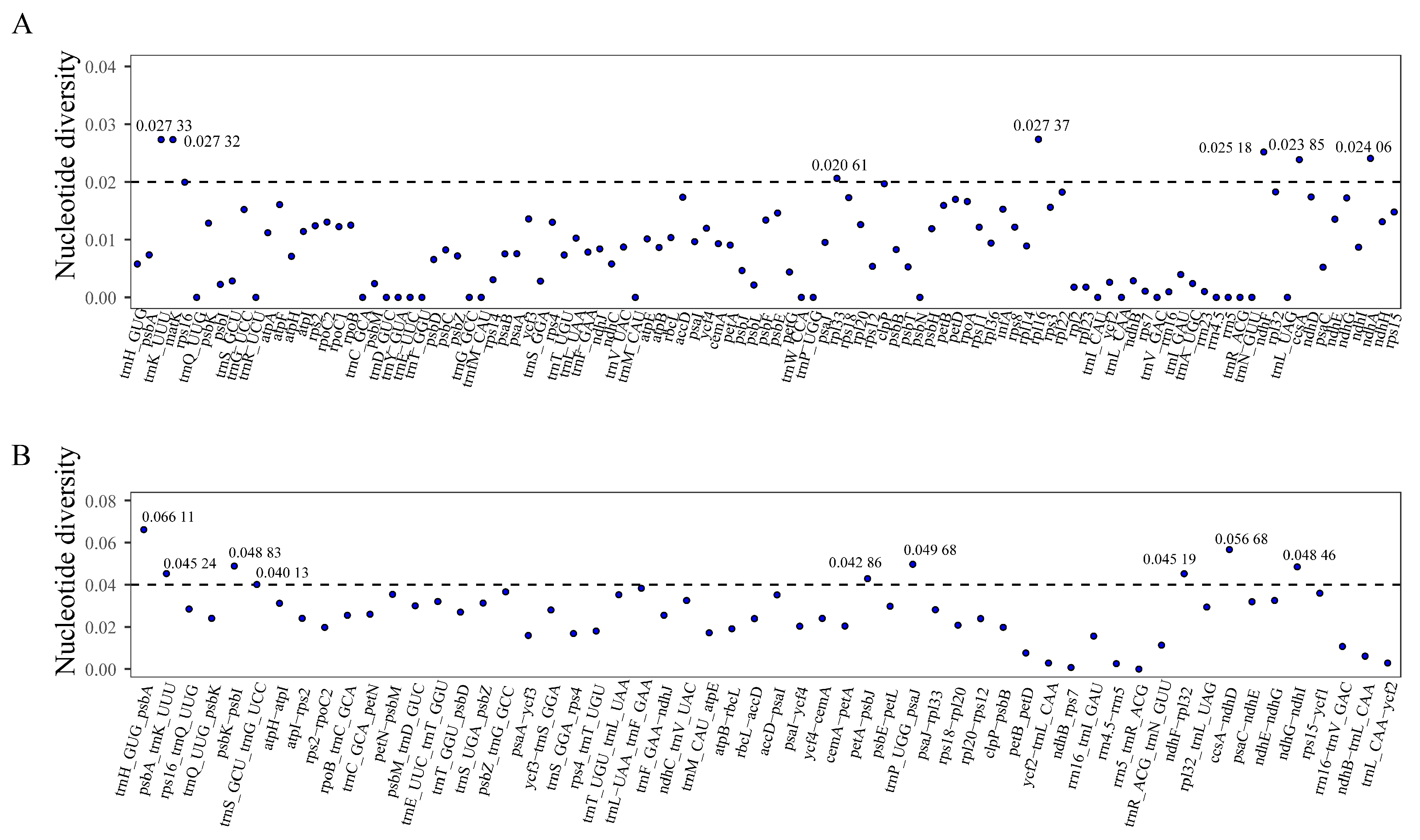
3.4. Comparative Genome Analysis
3.5. Phylogenetic Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Committee for the Flora of China, Chinese Academy of Sciences. Flora of China; Beijing Science Press: Beijing, China, 1996; Volume 30, Part 2; pp. 40–42. [Google Scholar]
- Chen, Z.Q. Supplement to Materia Medica; Wannan Medical College: Wuhu, China, 1983; p. 86. [Google Scholar]
- Dai, Z.H.; Wei, W.; Fan, L.Y. Microscopic identification of Stephaniae Longae Herba of Zhuang medicine. Res. Zhuang Yao Ethn. Med. 2020, 4, 63–67. (In Chinese) [Google Scholar]
- Li, L.; Zuo, A.X.; Rao, G.X. Studies on alkaloids from the Stephania epigaea of Dai traditional medicine. J. Yunnan Univ. Chin. Med. 2012, 35, 14–19. (In Chinese) [Google Scholar]
- An, C.; Chen, M.; Yang, C.Z.; Meng, J.; Zhuang, Y.X. Pharmacognostic Examination and Nomenclature Organization of ‘Jinbuhuan’. J. Chin. Med. Mater. 2019, 42, 2449–2452. (In Chinese) [Google Scholar]
- Luo, X.R. A preliminary study on the genus Stephania in China. Acta Phytotaxon. Sin. 1978, 16, 10–40. (In Chinese) [Google Scholar]
- Feng, Y.X.; Zhu, Z.Y.; Chen, H. Comparative Morphological and Histological Study of Medicinal Plants in the Genus Stephania. Acta Pharm. Sin. 1983, 11, 849–861. (In Chinese) [Google Scholar]
- Tang, C.F.; Song, S.Q. Woody lianas of the genus Stephania. Life World 2022, 11, 70–71. (In Chinese) [Google Scholar]
- Luo, X.R. An Outline of the Classification of the Genus Stephania in China. Bull. Bot. Res. 1982, 1, 33–59. (In Chinese) [Google Scholar]
- Zhu, Z.Y.; Feng, Y.X.; He, L.Y.; Wang, Y.C. Studies on the utilization of medicinal plant resources of the genus Stephania of the family Menispermaceae in China. Acta Pharm. Sin. 1983, 6, 460–467. (In Chinese) [Google Scholar]
- National Pharmacopoeia Committee. Pharmacopoeia of the People’s Republic of China; China Medical Science and Technology Press: Beijing, China, 2020; Volume I, p. 155. [Google Scholar]
- Yang, H.M.; Luo, X.R. Research on the ‘shanwugui’. Acta Pharm. Sin. 1980, 11, 674–683+705–707. (In Chinese) [Google Scholar]
- Yang, H.M.; Chen, Y. An overview of the chemical classification of plants of the genus Stephania in China. J. Integr. Plant Biol. 1994, 11, 14–22. (In Chinese) [Google Scholar]
- Li, X.J.; Liu, L.; Li, G.J.; Dong, J.W. Progress in chemical constituents, pharmacology and fermentation modification of radix Stephaniae Epigaeae. Shandong Chem. Ind. 2023, 52, 89–94. (In Chinese) [Google Scholar]
- Li, S.Z.; Wu, F. A review on the identification of radix Stephania Tetrandrae and the fake. Chin. J. Ethnomed. Ethnopharm. 2020, 29, 57–62. (In Chinese) [Google Scholar]
- Guo, M.; Zhang, R.Y.; Cai, F. Research progress on pharmacological activities of tetrandrine. China Mod. Med. 2018, 25, 30–33. (In Chinese) [Google Scholar]
- He, D.H.; Liu, J.; Fang, D.M.; Wang, X.L.; Li, L.M. Study on alkaloid constituents of Stephania tetrandra S. Moore. Acta Pharm. Sin. 2021, 56, 3503–3510. (In Chinese) [Google Scholar]
- Mo, L.Y.; Zhang, F.; Hao, E.W.; Deng, J.G.; Chen, F.; Li, R.L.; Mo, Y.M.; Zhang, L.Q.; Hou, X.T.; Du, Z.C. Predictive analysis of quality markers (Q-Marker) of Stephania tetrandra based on chemical components and pharmacological and network pharmacology. Chin. Tradit. Herb. Drugs 2022, 53, 6283–6295. (In Chinese) [Google Scholar]
- Shen, X.J.; Dong, J.W.; Mei, R.F.; Cai, L.; Ding, Z.T. Study on alkaloids of Stephania epigaea. Chem. Res. Appl. 2016, 28, 1730–1734. (In Chinese) [Google Scholar]
- Sun, Y.; Moore, M.J.; Zhang, S.; Soltis, P.S.; Soltis, D.E.; Zhao, T.; Meng, A.; Li, X.; Li, J.; Wang, H. Phylogenomic and structural analyses of 18 complete plastomes across nearly all families of early-diverging eudicots, including an angiosperm-wide analysis of IR gene content evolution. Mol. Phylogenet. Evol. 2016, 96, 93–101. [Google Scholar] [CrossRef]
- Wang, L.; Dong, W.P.; Zhou, S.L. Structural mutations and reorganizations in chloroplast genomes of flowering plants. Acta Bot. Boreali-Occident. Sin. 2012, 32, 1282–1288. (In Chinese) [Google Scholar]
- Freudenthal, J.A.; Pfaff, S.; Terhoeven, N.; Korte, A.; Ankenbrand, M.J.; Förster, F. A systematic comparison of chloroplast genome assembly tools. Genome Biol. 2020, 21, 254. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Du, C.H.; Zhan, H.X.; Shang, C.L.; Li, R.F.; Yuan, S.J. Comparative and phylogeny analysis of Platycodon grandiflorus complete chloroplast genomes. Chin. Tradit. Herb. Drugs 2023, 54, 4981–4991. (In Chinese) [Google Scholar]
- Zhang, W.X.; Liu, H.; Wang, L.; Zhang, S.H. Chloroplast genome comparison and phylogenetic analysis of four medicinal plants of evodia. Guid. J. Tradit. Chin. Med. Pharm. 2023, 29, 44–50+65. (In Chinese) [Google Scholar]
- Yang, X.Y.; Zhang, M.J.; Yin, T.; Han, P.C.; Du, C.J.; Zhang, H.Y. Chloroplast genome characteristics and codon preferences analysis in Syzygium. Southwest China J. Agric. Sci. 2023, 36, 1869–1880. (In Chinese) [Google Scholar]
- Schloss, P.D.; Jenior, M.L.; Koumpouras, C.C.; Westcott, S.L.; Highlander, S.K. Sequencing 16S rRNA gene fragments using the PacBio SMRT DNA sequencing system. PeerJ 2016, 4, e1869. [Google Scholar] [CrossRef] [PubMed]
- Lian, Y.; Huang, F.; Zhu, W.T.; Liu, X.F.; Wu, H.; Jiang, G.H.; Yin, X.M. Chloroplast genome structure of Stemona tuberosa and phylogenetic analysis based on PacBio sequencing. Chin. J. Exp. Tradit. Med. Formulae 2023, 29, 123–132. (In Chinese) [Google Scholar]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.J.; Yu, W.B.; Yang, J.B.; Song, Y.; de Pamphilis, C.W.; Yi, T.S.; Li, D.Z. GetOrganelle: A fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 2020, 21, 241. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Sharp, P.M.; Li, W.H. Codon usage in regulatory genes in Escherichia coli does not reflect selection for rare codons. Nucleic Acids Res. 1986, 14, 7737–7749. [Google Scholar] [CrossRef]
- Dai, J.P.; Cai, Y.M.; Liu, Q.Z.; Gao, X.X.; Zhu, S. Analysis of codon usage bias of chloroplast genome in seven Glycyrrhiza species. Chin. Tradit. Herb. Drugs 2023, 54, 2907–2916. (In Chinese) [Google Scholar]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [PubMed]
- Amiryousefi, A.; Hyvönen, J.; Poczai, P. IRscope: An online program to visualize the junction sites of chloroplast genomes. Bioinformatics 2018, 34, 3030–3031. [Google Scholar] [CrossRef] [PubMed]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef] [PubMed]
- Rozas, J.; Ferrer-Mata, A.; Sánchez-DelBarrio, J.C.; Guirao-Rico, S.; Librado, P.; Ramos-Onsins, S.E.; Sánchez-Gracia, A. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Quang Minh, B. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Dong, S.J.; Ying, Z.Q.; Yu, S.S.; Wang, Q.R.; Liao, G.H.; Ge, Y. Complete chloroplast genome of Stephania tetrandra (Menispermaceae) from Zhejiang Province: Insights into molecular structures, comparative genome analysis, mutational hotspots and phylogenetic relationships. BMC Genom. 2021, 22, 880. [Google Scholar] [CrossRef]
- Guan, Q.; Feng, D.; Fan, M. The complete chloroplast genome sequence of the medicinal plant Stephania epigaea H. S. Lo, 1978 (Menispermaceae) from Yunnan, China. Mitochondrial DNA B Resour. 2022, 7, 1403–1405. [Google Scholar] [CrossRef] [PubMed]
- Song, S.; Cameron, K.M.; Wang, Y.; Wang, S.; Jin, X.; Hina, F.; Yang, Z.; Li, P. Phylogenomics and phylogeography of Menispermum (Menispermaceae). Front. Plant Sci. 2023, 14, 1116300. [Google Scholar] [CrossRef]
- Kim, J.; Lee, J.; Um, S.; Choi, S.; Kim, H.; Chun, H.S.; Nah, G. The complete chloroplast genome of Sinomenium acutum (Menispermaceae). Mitochondrial DNA Part B Resour. 2020, 5, 2992–2993. [Google Scholar] [CrossRef]
- Zheng, L.P.; Feng, R. Characterization of the complete chloroplast genome of Fibraurea recisa Pierre 1885 (Menispermaceae), an important medicinal herb from Yunnan, China. Mitochondrial DNA Part B Resour. 2022, 7, 501–502. [Google Scholar] [CrossRef] [PubMed]
- Ni, Z.; Ye, Y.; Bai, T.; Xu, M.; Xu, L.A. Complete Chloroplast Genome of Pinus massoniana (Pinaceae): Gene Rearrangements, Loss of ndh Genes, and Short Inverted Repeats Contraction, Expansion. Molecules 2017, 22, 1528. [Google Scholar] [CrossRef] [PubMed]
- Zhao, P.Y.; Zhao, W.; Hou, Z.Y.; Li, X.; Liu, B.K.; Du, Y.; Xie, R.Q.; Cao, Z.L.; Wang, K.K. Comparison of chloroplast genome characteristics and phylogenetic analysis between Themeda japonica and two species of Themeda. Chin. Tradit. Herb. Drugs 2023, 54, 3261–3272. (In Chinese) [Google Scholar]
- Wang, X.L.; Xue, J.Y.; Zhang, Y.Y.; Xie, H.; Wang, Y.Q.; Weng, W.; Kang, Y.; Huang, J. DNA barcodes for the identification of Stephania (Menispermaceae) species. Mol. Biol. Rep. 2020, 47, 2197–2203. [Google Scholar] [CrossRef]
- Xie, D.T.; He, J.Y.; Huang, J.M.; Xie, H.; Wang, Y.Q.; Kang, Y.; Jabbour, F.; Guo, J. Molecular phylogeny of Chinese Stephania (Menispermaceae) and reassessment of the subgeneric and sectional classifications. Aust. Syst. Bot. 2015, 28, 246–255. [Google Scholar] [CrossRef]


| No. | Species | From | GenBank Accession No. |
|---|---|---|---|
| 1 | Stephania japonica var. timoriensis | This study | PP175340 |
| 2 | Stephania japonica var. discolor | This study | PP175339 |
| 3 | Stephania cephalantha | GenBank | NC_067079 |
| 4 | Stephania dielsiana | GenBank | NC_054337 |
| 5 | Stephania epigaea | GenBank | NC_058988 |
| 6 | Stephania japonica | GenBank | NC_029432 |
| 7 | Stephania kwangsiensis | GenBank | NC_048524 |
| 8 | Stephania tetrandra | GenBank | NC_050924 |
| 9 | Pericampylus glaucus | GenBank | NC_046846 |
| 10 | Menispermum canadense | GenBank | NC_048451 |
| 11 | Menispermum dauricum | GenBank | NC_042371 |
| 12 | Sinomenium acutum | GenBank | MT040976 |
| 13 | Fibraurea recisa | GenBank | NC_060536 |
| 14 | Akebia quinata | GenBank | KX611091 |
| 15 | Decaisnea insignis | GenBank | KY200671 |
| 16 | Thalictrum baicalense | GenBank | MW133265 |
| 17 | Thalictrum minus | GenBank | NC_041544 |
| 18 | Berberis amurensis | GenBank | KM057374 |
| 19 | Nandina domestica | GenBank | DQ923117 |
| 20 | Sinopodophyllum hexandrum | GenBank | KR779994 |
| 21 | Epimedium lishihchenii | GenBank | KU522472 |
| 22 | Vancouveria planipetala | GenBank | MH337373 |
| Species | Genome | LSC | IR | SSC | ||||
|---|---|---|---|---|---|---|---|---|
| Length/bp | GC/% | Length/bp | GC/% | Length/bp | GC/% | Length/bp | GC/% | |
| S. japonica var. timoriensis | 157,609 | 38.25 | 88,477 | 36.46 | 24,393 | 43.72 | 20,346 | 32.94 |
| S. japonica var. discolor | 157,748 | 38.26 | 88,581 | 36.46 | 24,401 | 43.73 | 20,365 | 32.97 |
| Gene Function | Gene Group | Gene Name | Number |
|---|---|---|---|
| Gene for photosynthesis | Photosystem I | psaA, psaB, psaC, psaI, psaJ | 5 |
| Photosystem II | psbA, psbB, psbC, psbD, psbE, psbF, psbH, psbI, psbJ, psbK, psbL, psbM, psbN, psbT, psbZ | 15 | |
| Cytochrome b/f complex | petA, petB, petD, petG, petL, petN | 6 | |
| NADH dehydrogenase | ndhA, ndhB a, ndhC, ndhD, ndhE, ndhF, ndhG, ndhH, ndhI, ndhJ, ndhK | 12 | |
| Subunits of ATP synthase | atpA, atpB, atpE, atpF, atpH, atpI | 6 | |
| Rubisco large subunits | rbcL | 1 | |
| Self-replication | Lange subunits of ribosome | rpl2 a, rpl14, rpl16, rpl20, rpl22, rpl23 a, rpl32, rpl33, rpl36 | 11 |
| Small subunits of ribosome | rps2, rps3, rps4, rps7 a, rps8, rps11, rps12 a, rps14, rps15, rps16, rps18, rps19 | 14 | |
| RNA polymerase | rpoA, rpoB, rpoC1, rpoC2 | 4 | |
| Ribosomal RNA genes | rrn4.5S a, rrn5S a, rrn16S a, rrn23S a | 8 | |
| Transfer RNA genes | trnA-UGC a, trnC-GCA, trnD-GUC, trnE-UUC, trnF-GAA, trnfM-CAU, trnG-GCC, trnG-UCC, trnH-GUG, trnI-CAU a, trnI-GAU a, trnK-UUU, trnL-CAA a, trnL-UAA, trnL-UAG, trnM-CAU, trnN-GUU a, trnP-UGG, trnQ-UUG, trnR-ACG a, trnR-UCU, trnS-GCU, trnS-GGA, trnS-UGA, trnT-GGU, trnT-UGU, trnV-GAC a, trnV-UAC, trnW-CCA, trnY-GUA | 37 | |
| Other genes | Protease | clpP | 1 |
| Maturase | matK | 1 | |
| Translation initiation factor | infA | 1 | |
| Envelop membrane protein | cemA | 1 | |
| Subunits of acety-CoA-carboxylase | accD | 1 | |
| C-type cytochrome synthesis | ccsA | 1 | |
| Unknow gene | Hypothetical chloroplast reading frames | ycf1, ycf2 a, ycf3, ycf4 | 5 |
| Total | 130 |
| Types and Regions | Name | S. japonica var. timoriensis | S. japonica var. discolor |
|---|---|---|---|
| Types | mononucleotide | 168 | 176 |
| dinucleotide | 18 | 18 | |
| trinucleotide | 7 | 6 | |
| tetranucleotide | 9 | 7 | |
| pentanucleotide | 5 | 4 | |
| Regions | LSC | 142 | 143 |
| SSC | 45 | 48 | |
| IR | 20 | 20 | |
| Total | 207 | 211 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, L.-L.; Geng, Y.-M.; Zheng, L.-P. Characterization and Phylogenetic Analysis of the Chloroplast Genomes of Stephania japonica var. timoriensis and Stephania japonica var. discolor. Genes 2024, 15, 877. https://doi.org/10.3390/genes15070877
Wu L-L, Geng Y-M, Zheng L-P. Characterization and Phylogenetic Analysis of the Chloroplast Genomes of Stephania japonica var. timoriensis and Stephania japonica var. discolor. Genes. 2024; 15(7):877. https://doi.org/10.3390/genes15070877
Chicago/Turabian StyleWu, Li-Li, Ying-Min Geng, and Lan-Ping Zheng. 2024. "Characterization and Phylogenetic Analysis of the Chloroplast Genomes of Stephania japonica var. timoriensis and Stephania japonica var. discolor" Genes 15, no. 7: 877. https://doi.org/10.3390/genes15070877




