Comparison of the Fecal Microbiota in Feral and Domestic Goats
Abstract
:1. Introduction
2. Results and Discussion
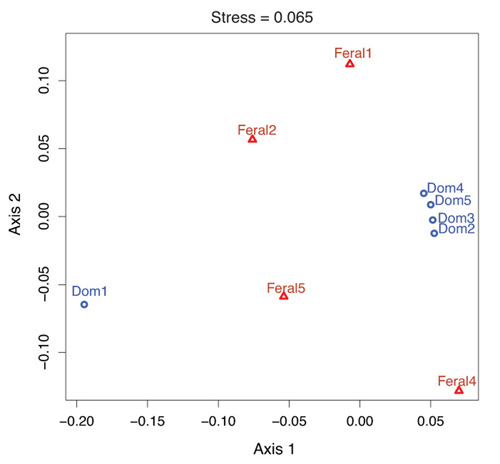
2.1. Fecal Bacterial Community Structures Differ Between Domestic and Feral Goats
| Taxonomic level | Feral goats (n = 4) | Domestic goats (n = 5) | Total N |
|---|---|---|---|
| Phylum | 38 ± 1 | 40 ± 1 | 42 |
| Class | 45 ± 1 | 48 ± 1 | 52 |
| Order | 73 ± 2 | 76 ± 3 | 92 |
| Family | 157 ± 1 | 132 ± 4 | 153 |
| Subfamily | 290 ± 28 | 335 ± 46 | 548 |
| OTUs | 1,121 ± 47 | 1,268 ± 74 | 1,982 |



2.2. Diet and Selective Breeding Might Affect Gut Bacterial Community Structure
2.3. Goat’s Gut Bacteria as a Reservoir of Antibiotic Resistance Genes
| Goats | tet(M) | tet(S) | tet(O) | tet(Q) | tet(W) |
|---|---|---|---|---|---|
| Feral (n = 5) | 0 | 0 | 5 | 0 | 5 |
| Domestic (n = 5) | 0 | 0 | 5 | 5 | 5 |
3. Experimental Section
3.1. Animals and Samples
3.2. DNA Extraction and Amplification
3.3. DNA Array Hybridization
3.4. Data and Statistical Analysis
3.5. Detection of Tetracycline Resistance Genes
4. Conclusions
Acknowledgments
References
- Ley, R.E.; Hamady, M.; Lozupone, C.; Turnbaugh, P.J.; Ramey, R.R.; Bircher, J.S.; Schlegel, M.L.; Tucker, T.A.; Schrenzel, M.D.; Knight, R.; Gordon, J.I. Evolution of mammals and their gut microbes. Science 2008, 320, 1647–1651. [Google Scholar] [CrossRef]
- Backhed, F.; Ding, H.; Wang, T.; Hooper, L.V.; Koh, G.Y.; Nagy, A.; Semenkovich, C.F.; Gordon, J.I. The gut microbiota as an environmental factor that regulates fat storage. Proc. Natl. Acad. Sci. USA 2004, 101, 15718–15723. [Google Scholar] [CrossRef]
- Dillon, R.J.; Vennard, C.T.; Buckling, A.; Charnley, A.K. Diversity of locust gut bacteria protects against pathogen invasion. Ecol. Lett. 2005, 8, 1291–1298. [Google Scholar] [CrossRef]
- Cebra, J.J. Influences of microbiota on intestinal immune system development. Am. J. Clin. Nutr. 1999, 69, 1046–1051. [Google Scholar]
- Konturek, J.W.; Konturek, S.J.; Kwiecien, N.; Bielanski, W.; Pawlik, T.; Rembiasz, K.; Domschke, W. Leptin in the control of gastric secretion and gut hormones in humans infected with Helicobacter pylori. Scand. J. Gastroenterol. 2001, 36, 1148–1154. [Google Scholar] [CrossRef]
- Nwokolo, C.U.; Freshwater, D.A.; O’Hare, P.; Randeva, H.S. Plasma ghrelin following cure of Helicobacter pylori. Gut 2003, 52, 637–640. [Google Scholar] [CrossRef]
- Zoetendal, E.G.; Akkermans, A.D.L.; Akkermans-van Vliet, W.M.; de Visser, J.A.G.M.; de Vos, W.M. The host genotype affects the bacterial community in the human gastrointestinal tract. Microb. Ecol. Health Dis. 2001, 13, 129–134. [Google Scholar] [CrossRef]
- Macdonald, T.T.; Monteleone, G. Immunity, inflammation, and allergy in the gut. Science 2005, 307, 1920–1925. [Google Scholar] [CrossRef]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology: Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef]
- Rawls, J.F.; Samuel, B.S.; Gordon, J.I. Gnotobiotic zebrafish reveal evolutionarily conserved responses to the gut microbiota. Proc. Natl. Acad. Sci. USA 2004, 101, 4596–4601. [Google Scholar] [CrossRef]
- Zeder, M.A.; Hesse, B. The initial domestication of goats (Capra hircus) in the zagros mountains 10,000 years ago. Science 2000, 287, 2254–2257. [Google Scholar] [CrossRef]
- Shrestha, J.N.B.; Fahmy, M.H. Breeding goats for meat production: A review: 1. Genetic resources, management and breed evaluation. Small Rumin. Res. 2005, 58, 93–106. [Google Scholar] [CrossRef]
- Glazko, V. An attempt at understanding the genetic basis of domestication. Anim. Sci. Papers Rep. 2003, 21, 109–120. [Google Scholar]
- Khachatourians, G. Agricultural use of antibiotics and the evolution and transfer of antibiotic resistant bacteria. Can. Med. Assoc. 1998, 159, 1129–1136. [Google Scholar]
- Butaye, P.; Devriese, L.A.; Haesebrouck, F. Antimicrobial growth promoters used in animal feed: A review of the less well known antibiotics and their effects on Gram-positive bacteria. Clin. Microbiol. Rev. 2003, 16, 175–188. [Google Scholar] [CrossRef]
- Heatwoke, H.; Levins, R.; Byer, M. Biogeography of the puerto rican bank. Atoll Res. Bull. 1981, 251, 1042–1055. [Google Scholar] [CrossRef]
- Sponenberg, D.P. Colonial spanish sheep, goats, hogs, and asses in the United States. Zootecnia 1992, 41, 415–419. [Google Scholar]
- Yang, G.; Bao, B.; Peatman, E.; Li, H.; Huang, L.; Ren, D. Analysis of the composition of the bacterial community in puffer fish Takifugu obscurus. Aquaculture 2007, 262, 183–191. [Google Scholar] [CrossRef]
- Croswell, A.; Amir, E.; Teggatz, P.; Barman, M.; Salzman, N. Prolonged impact of antibiotics on intestinal microbial ecology and susceptibility to enteric salmonella infection. Infect. Immun. 2009, 77, 2741–2753. [Google Scholar] [CrossRef]
- Leser, T.D.; Lindecrona, R.H.; Jensen, T.K.; Jensen, B.B.; Moller, K. Changes in bacterial community structure in the colon of pigs fed different experimental diets and after infection with Brachyspira hyodysenteriae. Appl. Environ. Microbiol. 2000, 66, 3290–3296. [Google Scholar] [CrossRef]
- Meléndez-Ackerman, E.J.; Cortés, C.; Sustache, J.; Aragón, S.; Morales-Vargas, M.; García-Bermúdez, M.; Fernández, D.S. Diet of feral goats in Mona Island Reserve, Puerto Rico. Caribb. J. Sci. 2008, 44, 199–205. [Google Scholar]
- Aminov, R.I.; Garrigues-Jeanjean, N.; Mackie, R.I. Molecular ecology of tetracycline resistance: Development and validation of primers for detection of tetracycline resistance genes encoding ribosomal protection proteins. Appl. Environ. Microbiol. 2001, 67, 22–32. [Google Scholar] [CrossRef]
- Aires, J.; Doucet Populaire, F.; Butel, M.J. Tetracycline resistance mediated by tet(W), tet(M), and tet(O) genes of Bifidobacterium isolates from humans. Appl. Environ. Microbiol. 2007, 73, 2751–2754. [Google Scholar] [CrossRef]
- Ready, D.; Pratten, J.; Roberts, A.P.; Bedi, R.; Mullany, P.; Wilson, M. Potential role of Veillonella spp. as a reservoir of transferable tetracycline resistance in the oral cavity. Antimicrob. Agents Chemother. 2006, 50, 2866–2868. [Google Scholar] [CrossRef]
- Stanton, T.B.; Humphrey, S.B.; Stoffregen, W.C. Chlortetracycline-resistant intestinal bacteria in organically-raised and feral swine. Appl. Environ. Microbiol. 2011, 77. [Google Scholar] [CrossRef]
- Osterblad, M.; Norrdahl, K.; Korpimaki, E.; Huovinen, P. Antibiotic resistance. How wild are wild mammals? Nature 2001, 409, 37–38. [Google Scholar]
- Gardner, P.; Smith, D.H.; Beer, H.; Moellering, R.C., Jr. Recovery of resistance (R) factors from a drug-free community. Lancet 1969, 2, 774–776. [Google Scholar] [CrossRef]
- Routman, E.; Miller, R.D.; Phillips-Conroy, J.; Hartl, D.L. Antibiotic resistance and population structure in Escherichia coli from free-ranging African yellow baboons. Appl. Environ. Microbiol. 1985, 50, 749–754. [Google Scholar]
- Gilliver, M.A.; Bennett, M.; Begon, M.; Hazel, S.M.; Hart, C.A. Antibiotic resistance found in wild rodents. Nature 1999, 401, 233–234. [Google Scholar]
- Berg, J.; Tom-Petersen, A.; Nybroe, O. Copper amendment of agricultural soil selects for bacterial antibiotic resistance in the field. Lett. Appl. Microbiol. 2005, 40, 146–151. [Google Scholar] [CrossRef]
- Coates, M.D.; MK.; Kon, S.K. The effect of antibiotics on the intestine of the chick. Br. J. Nutr. 1955, 9, 110–119. [Google Scholar] [CrossRef]
- Guillemot, D.; Carbon, C.; Balkau, B.; Geslin, P.; Lecoeur, H.; Vauzelle-Kervroëdan, F.; Bouvenot, G.; Eschwége, E. Low dosage and long treatment duration of beta-lactam: Risk factors for carriage of penicillin-resistant Streptococcus pneumoniae. J. Am. Med. Assoc. 1998, 279, 365–370. [Google Scholar] [CrossRef]
- DeSantis, T.Z.; Brodie, E.L.; Moberg, J.P.; Zubieta, I.X.; Piceno, Y.M.; Andersen, G.L. High-density universal 16S rRNA microarray analysis reveals broader diversity than typical clone library when sampling the environment. Microb. Ecol. 2007, 53, 371–383. [Google Scholar] [CrossRef]
- Brodie, E.L.; Desantis, T.Z.; Joyner, D.C.; Baek, S.M.; Larsen, J.T.; Andersen, G.L.; Hazen, T.C.; Richardson, P.M.; Herman, D.J.; Tokunaga, T.K.; et al. Application of a high-density oligonucleotide microarray approach to study bacterial population dynamics during uranium reduction and reoxidation. Appl. Environ. Microbiol. 2006, 72, 6288–6298. [Google Scholar] [CrossRef]
- Lozupone, C.; Hamady, M.; Knight, R. UniFrac-An online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinforma. 2006, 7. [Google Scholar] [CrossRef]
- Schatz, M.C.; Phillippy, A.M.; Gajer, P.; DeSantis, T.Z.; Andersen, G.L.; Ravel, J. Integrated microbial survey analysis of Prokaryotic communities for the PhyloChip microarray. Appl. Environ. Microbiol. 2010, 76, 5636–5638. [Google Scholar] [CrossRef]
- PhyloTrac: Environmental Sample Analysis. Available online: http://www.phylotrac.org/Home.html (accessed on 28 November 2011).
- The R Project for Statistical Computing. Available online: http://www.R-project.org (accessed on 28 November 2011).
- Development Core Team, R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2008.
- Dudoit, S.; Yang, Y.H.; Callow, M.J.; Speed, T.P. Statistical methods for identifying differentially expressed genes in replicated cDNA microarray experiments. Tech. Rep. 2000, 12, 111–139. [Google Scholar]
- Schnappinger, D.; Hillen, W. Tetracyclines: Antibiotic action, uptake, and resistance mechanisms. Arch. Microbiol. 1996, 165, 359–369. [Google Scholar] [CrossRef]
- Roberts, M.C. Update on acquired tetracycline resistance genes. FEMS Microbiol. Lett. 2005, 245, 195–203. [Google Scholar] [CrossRef]
- Villedieu, A.; Diaz-Torres, M.L.; Hunt, N.; McNab, R.; Spratt, D.A.; Wilson, M.; Mullany, P. Prevalence of tetracycline resistance genes in oral bacteria. Antimicrob. Agents Chemother. 2003, 47, 878–882. [Google Scholar] [CrossRef]
Supplementary Materials
| Phylum | Phylum Ranking | N Families | Average N OTUs | |||
|---|---|---|---|---|---|---|
| Domestic | Feral | Domestic | Feral | Domestic (N = 5) | Feral (N = 4) | |
| Firmicutes | 1 | 1 | 25 | 25 | 485 ± 23.9 | 421 ± 21.9 |
| Proteobacteria | 2 | 2 | 70 | 59 | 340 ± 47.5 | 326 ± 31.0 |
| Actinobacteria | 3 | 3 | 31 | 29 | 115 ± 10.5 | 93 ± 2.1 |
| Bacteroidetes | 4 | 4 | 11 | 11 | 88 ± 4.8 | 82 ± 6.9 |
| Acidobacteria | 5 | 5 | 2 | 2 | 40 ± 1.7 | 37 ± 3.7 |
| Spirochaetes | 6 | 11 | 2 | 2 | 28 ± 2.1 | 9 ± 4.1 |
| Chloroflexi | 7 | 6 | 1 | 1 | 25 ± 1.4 | 22 ± 1.8 |
| Unclassified | 8 | 7 | 1 | 1 | 23 ± 1.3 | 20 ± 1.0 |
| Verrucomicrobia | 9 | 8 | 5 | 5 | 21 ± 0.6 | 19 ± 1.0 |
| Cyanobacteria | 10 | 9 | 1 | 1 | 14 ± 2.0 | 16 ± 4.6 |
| Planctomycetes | 11 | 10 | 4 | 4 | 12 ± 0.7 | 13 ± 0.6 |
| Gemmatimonadetes | 12 | 12 | 1 | 1 | 7 ± 0.7 | 6 ± 1.1 |
| Chlorobi | 13 | 15 | 2 | 2 | 6 ± 0.8 | 4 ± 0.7 |
| OP10 | 14 | 14 | 1 | 1 | 5 ± 0.6 | 4 ± 0.3 |
| TM7 | 15 | 16 | 1 | 1 | 5 ± 0.4 | 4 ± 0.9 |
| Synergistes | 16 | 19 | 1 | 1 | 5 ± 0.4 | 3 ± 0.3 |
| Natronoanaerobium | 17 | 13 | 1 | 1 | 4 ± 0.7 | 5 ± 0.5 |
| Deinococcus-Thermus | 18 | 17 | 1 | 1 | 4 ± 0.0 | 4 ± 0.0 |
| OP9/JS1 | 19 | 22 | 1 | 1 | 4 ± 0.6 | 3 ± 0.5 |
| NC10 | 20 | 18 | 1 | 1 | 4 ± 0.2 | 4 ± 0.3 |
| Nitrospira | 21 | 24 | 1 | 1 | 3 ± 0.6 | 3 ± 0.6 |
| Lentisphaerae | 22 | 20 | 1 | 1 | 3 ± 0.0 | 3 ± 0.0 |
| OP3 | 23 | 21 | 1 | 1 | 3 ± 0.0 | 3 ± 0.0 |
| BRC1 | 24 | 25 | 1 | 1 | 3 ± 0.0 | 3 ± 0.5 |
| Chlamydiae | 25 | 23 | 3 | 3 | 2 ± 0.2 | 3 ± 0.3 |
| Aquificae | 26 | 27 | 2 | 2 | 2 ± 0.4 | 2 ± 0.5 |
| marine group A | 27 | 29 | 1 | 1 | 2 ± 0.5 | 1 ± 0.3 |
| Caldithrix | 28 | 26 | 1 | 1 | 2 ± 0.0 | 2 ± 0.0 |
| WS3 | 29 | 28 | 1 | 1 | 2 ± 0.2 | 2 ± 0.3 |
| SPAM | 30 | 30 | 1 | 1 | 1 ± 0.2 | 1 ± 0.0 |
| DSS1 | 31 | 31 | 1 | 1 | 1 ± 0.2 | 1 ± 0.0 |
| TM6 | 32 | 35 | 1 | 1 | 1 ± 0.2 | 1 ± 0.3 |
| LD1PA group | 33 | 32 | 1 | 1 | 1 ± 0.0 | 1 ± 0.0 |
| OP8 | 34 | 33 | 1 | 1 | 1 ± 0.0 | 1 ± 0.0 |
| Thermodesulfobacteria | 35 | 34 | 1 | 1 | 1 ± 0.0 | 1 ± 0.0 |
| AD3 | 36 | 36 | 1 | 1 | 1 ± 0.0 | 1 ± 0.3 |
| Thermotogae | 37 | 37 | 1 | 1 | 1 ± 0.2 | 1 ± 0.3 |
| WS5 | 38 | 38 | 1 | 1 | 1 ± 0.2 | 1 ± 0.3 |
| Coprothermobacteria | 39 | 39 | 1 | 1 | 1 ± 0.2 | 1 ± 0.3 |
| OD1 | 40 | 41 | 1 | 0 | 1 ± 0.2 | 0 ± 0.0 |
| Deferribacteres | 41 | 40 | 1 | 1 | 1 ± 0.2 | 0 ± 0.3 |
| Termite group 1 | 42 | 42 | 1 | 0 | 0 ± 0.4 | 0 ± 0.0 |




| Taxa | p-value Holm correction |
|---|---|
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6562 | 3.51E-11 |
| Bacteria;Proteobacteria;Deltaproteobacteria;Unclassified;Unclassified;sf_9;9890 | 8.45E-05 |
| Bacteria;Proteobacteria;Epsilonproteobacteria;Campylobacterales;Helicobacteraceae;sf_3;10467 | 0.0006201 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Thermomonosporaceae;sf_1;1546 | 0.00126733 |
| Bacteria;Firmicutes;gut clone group;Unclassified;Unclassified;sf_1;4400 | 0.00747837 |
| Bacteria;OP3;Unclassified;Unclassified;Unclassified;sf_2;349 | 0.00775145 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6506 | 0.00825169 |
| Bacteria;Bacteroidetes;Bacteroidetes;Bacteroidales;Unclassified;sf_15;6233 | 0.00850866 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Rhodobacterales;Rhodobacteraceae;sf_1;7508 | 0.00853058 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6571 | 0.0087866 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6476 | 0.00905065 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6568 | 0.00914326 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6490 | 0.00938643 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Micromonosporaceae;sf_1;1910 | 0.00962793 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Micromonosporaceae;sf_1;1488 | 0.00962793 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Micromonosporaceae;sf_1;1633 | 0.00962793 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Micromonosporaceae;sf_1;1760 | 0.00962793 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6507 | 0.00962793 |
| Bacteria;Synergistes;Unclassified;Unclassified;Unclassified;sf_3;117 | 0.00962793 |
| Bacteria;Firmicutes;Catabacter;Unclassified;Unclassified;sf_4;4517 | 0.00962793 |
| Bacteria;OD1;OP11-5;Unclassified;Unclassified;sf_1;515 | 0.00962793 |
| Bacteria;Proteobacteria;Gammaproteobacteria;Alteromonadales;Alteromonadaceae;sf_1;8222 | 0.00962883 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6489 | 0.00963896 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Kineosporiaceae;sf_1;1581 | 0.00966581 |
| Bacteria;Proteobacteria;Gammaproteobacteria;Unclassified;Unclassified;sf_3;8606 | 0.00966755 |
| Bacteria;Firmicutes;Clostridia;Clostridiales;Lachnospiraceae;sf_5;4324 | 0.00967552 |
| Bacteria;Firmicutes;Clostridia;Clostridiales;Lachnospiraceae;sf_5;2810 | 0.00967554 |
| Bacteria;Firmicutes;Clostridia;Clostridiales;Lachnospiraceae;sf_5;3223 | 0.00967787 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6487 | 0.00968213 |
| Bacteria;Actinobacteria;Actinobacteria;Acidimicrobiales;Acidimicrobiaceae;sf_1;2014 | 0.00970127 |
| Bacteria;Firmicutes;gut clone group;Unclassified;Unclassified;sf_1;4579 | 0.00970475 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Gordoniaceae;sf_1;1654 | 0.00971853 |
| Bacteria;Firmicutes;Clostridia;Clostridiales;Unclassified;sf_17;3099 | 0.00977429 |
| Bacteria;Proteobacteria;Unclassified;Unclassified;Unclassified;sf_8;8247 | 0.00978004 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;7477 | 0.00979143 |
| Bacteria;Proteobacteria;Betaproteobacteria;Nitrosomonadales;Nitrosomonadaceae;sf_1;7770 | 0.00979751 |
| Bacteria;Proteobacteria;Deltaproteobacteria;Myxococcales;Polyangiaceae;sf_3;9671 | 0.00980123 |
| Bacteria;Proteobacteria;Epsilonproteobacteria;Campylobacterales;Helicobacteraceae;sf_3;10417 | 0.00981157 |
| Bacteria;Actinobacteria;Actinobacteria;Acidimicrobiales;Microthrixineae;sf_12;1721 | 0.00987931 |
| Bacteria;Proteobacteria;Deltaproteobacteria;EB1021 group;Unclassified;sf_4;9741 | 0.00990536 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Gordoniaceae;sf_1;1184 | 0.01015274 |
| Bacteria;Proteobacteria;Gammaproteobacteria;Aeromonadales;Succinivibrionaceae;sf_1;8822 | 0.01020072 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6508 | 0.01036306 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6488 | 0.0104335 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;7044 | 0.01047121 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6523 | 0.01062729 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6494 | 0.01082364 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Promicromonosporaceae;sf_1;1711 | 0.01086604 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;6799 | 0.01118413 |
| Bacteria;Firmicutes;Mollicutes;Mycoplasmatales;Mycoplasmataceae;sf_1;3929 | 0.01120143 |
| Bacteria;Proteobacteria;Gammaproteobacteria;Enterobacteriales;Enterobacteriaceae;sf_1;8607 | 0.01120143 |
| Bacteria;Proteobacteria;Gammaproteobacteria;Thiotrichales;Thiotrichaceae;sf_3;8477 | 0.01120143 |
| Bacteria;Lentisphaerae;Unclassified;Unclassified;Unclassified;sf_5;10330 | 0.01120143 |
| Bacteria;Lentisphaerae;Unclassified;Unclassified;Unclassified;sf_5;9704 | 0.01120143 |
| Bacteria;Unclassified;Unclassified;Unclassified;Unclassified;sf_160;2385 | 0.01120143 |
| Bacteria;Firmicutes;Catabacter;Unclassified;Unclassified;sf_4;4325 | 0.01120143 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Rhodobacterales;Rhodobacteraceae;sf_1;7511 | 0.01120154 |
| Bacteria;Nitrospira;Nitrospira;Nitrospirales;Nitrospiraceae;sf_3;833 | 0.01126881 |
| Bacteria;Firmicutes;Clostridia;Clostridiales;Lachnospiraceae;sf_5;4489 | 0.01129829 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Rhizobiales;Bradyrhizobiaceae;sf_1;7029 | 0.01138694 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6479 | 0.01144975 |
| Bacteria;Firmicutes;Clostridia;Clostridiales;Lachnospiraceae;sf_5;3075 | 0.01145728 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;6636 | 0.01173191 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;7522 | 0.01173191 |
| Bacteria;Actinobacteria;Actinobacteria;Actinomycetales;Micromonosporaceae;sf_1;1931 | 0.01174823 |
| Bacteria;Proteobacteria;Gammaproteobacteria;Enterobacteriales;Enterobacteriaceae;sf_1;8362 | 0.01178839 |
| Bacteria;Bacteroidetes;Flavobacteria;Flavobacteriales;Flavobacteriaceae;sf_1;6274 | 0.01184388 |
| Bacteria;Bacteroidetes;Bacteroidetes;Bacteroidales;Prevotellaceae;sf_1;5398 | 0.01200674 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Beijerinck/Rhodoplan/Methylocyst; sf_3;7219 | 0.01219961 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;6768 | 0.01267711 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;6867 | 0.01287698 |
| Bacteria;Spirochaetes;Spirochaetes;Spirochaetales;Spirochaetaceae;sf_1;6580 | 0.01365949 |
| Bacteria;Firmicutes;Mollicutes;Unclassified;Unclassified;sf_1;4000 | 0.01386526 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Rhizobiales;Rhizobiaceae;sf_1;6770 | 0.0145046 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;7316 | 0.01458135 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Bradyrhizobiales;Bradyrhizobiaceae;sf_1;7333 | 0.01458135 |
| Bacteria;Firmicutes;Clostridia;Clostridiales;Clostridiaceae;sf_12;4384 | 0.0149426 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Rhodobacterales;Rhodobacteraceae;sf_1;6652 | 0.01867578 |
| Bacteria;Proteobacteria;Epsilonproteobacteria;Campylobacterales;Helicobacteraceae;sf_23;10443 | 0.02022574 |
| Bacteria;Proteobacteria;Epsilonproteobacteria;Campylobacterales;Helicobacteraceae;sf_3;10576 | 0.02022574 |
| Bacteria;Proteobacteria;Deltaproteobacteria;Myxococcales;Polyangiaceae;sf_4;9733 | 0.02218193 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Sphingomonadales;Sphingomonadaceae;sf_1;7215 | 0.0467565 |
| Bacteria;Proteobacteria;Alphaproteobacteria;Sphingomonadales;Sphingomonadaceae;sf_1;7100 | 0.04689578 |
| Bacteria;Proteobacteria;Gammaproteobacteria;Alteromonadales;Alteromonadaceae;sf_1;8978 | 0.04825366 |
| Phylum | Family | Domestic | Feral |
|---|---|---|---|
| Actinobacteria | Acidimicrobiaceae | 1 | 0 |
| Gordoniaceae | 2 | 0 | |
| Kineosporiaceae | 1 | 0 | |
| Micromonosporaceae | 5 | 0 | |
| Microthrixineae | 1 | 0 | |
| Promicromonosporaceae | 1 | 0 | |
| Thermomonosporaceae | 1 | 0 | |
| Bacteroidetes | Flavobacteriaceae | 1 | 0 |
| Prevotellaceae | 0 | 1 | |
| Unclassified | 1 | 0 | |
| Firmicutes | Clostridiaceae | 1 | 0 |
| Lachnospiraceae | 4 | 1 | |
| Mycoplasmataceae | 1 | 0 | |
| Unclassified | 5 | 1 | |
| Lentisphaerae | Unclassified | 2 | 0 |
| Nitrospira | Nitrospiraceae | 0 | 1 |
| OD1 | Unclassified | 1 | 0 |
| OP3 | Unclassified | 1 | 0 |
| Proteobacteria | Alteromonadaceae | 2 | 0 |
| Beijerinck/Rhodoplan/Methylocyst | 0 | 1 | |
| Bradyrhizobiaceae | 0 | 10 | |
| Enterobacteriaceae | 1 | 1 | |
| Helicobacteraceae | 4 | 0 | |
| Nitrosomonadaceae | 1 | 0 | |
| Polyangiaceae | 2 | 0 | |
| Rhizobiaceae | 0 | 1 | |
| Rhodobacteraceae | 2 | 1 | |
| Sphingomonadaceae | 2 | 0 | |
| Succinivibrionaceae | 1 | 0 | |
| Thiotrichaceae | 0 | 1 | |
| Unclassified | 4 | 0 | |
| Spirochaetes | Spirochaetaceae | 15 | 0 |
| Synergistes | Unclassified | 1 | 0 |
| Unclassified | Unclassified | 1 | 0 |
| TOTAL | 65 | 19 | |
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
De Jesús-Laboy, K.M.; Godoy-Vitorino, F.; Piceno, Y.M.; Tom, L.M.; Pantoja-Feliciano, I.G.; Rivera-Rivera, M.J.; Andersen, G.L.; Domínguez-Bello, M.G. Comparison of the Fecal Microbiota in Feral and Domestic Goats. Genes 2012, 3, 1-18. https://doi.org/10.3390/genes3010001
De Jesús-Laboy KM, Godoy-Vitorino F, Piceno YM, Tom LM, Pantoja-Feliciano IG, Rivera-Rivera MJ, Andersen GL, Domínguez-Bello MG. Comparison of the Fecal Microbiota in Feral and Domestic Goats. Genes. 2012; 3(1):1-18. https://doi.org/10.3390/genes3010001
Chicago/Turabian StyleDe Jesús-Laboy, Kassandra M., Filipa Godoy-Vitorino, Yvette M. Piceno, Lauren M. Tom, Ida G. Pantoja-Feliciano, Michelle J. Rivera-Rivera, Gary L. Andersen, and María G. Domínguez-Bello. 2012. "Comparison of the Fecal Microbiota in Feral and Domestic Goats" Genes 3, no. 1: 1-18. https://doi.org/10.3390/genes3010001