The Usefulness of Cell-Based and Liquid-Based Urine Tests in Clarifying the Diagnosis and Monitoring the Course of Urothelial Carcinoma. Identification of Novel, Potentially Actionable, RB1 and ERBB2 Somatic Mutations
Abstract
:1. Introduction
1.1. Background
1.2. Patient’s Medical History
2. Materials and Methods
2.1. Cytourofish(+)
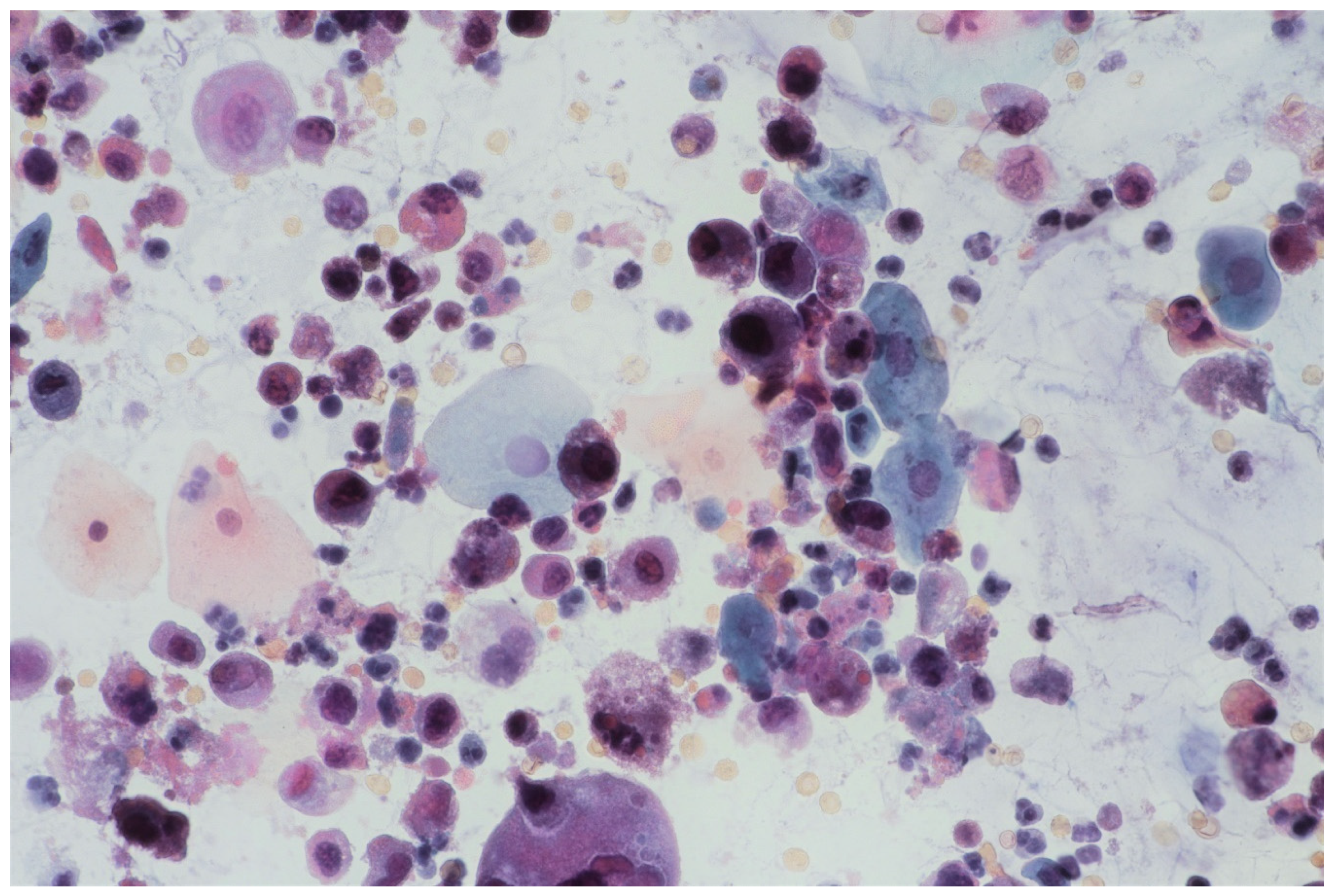
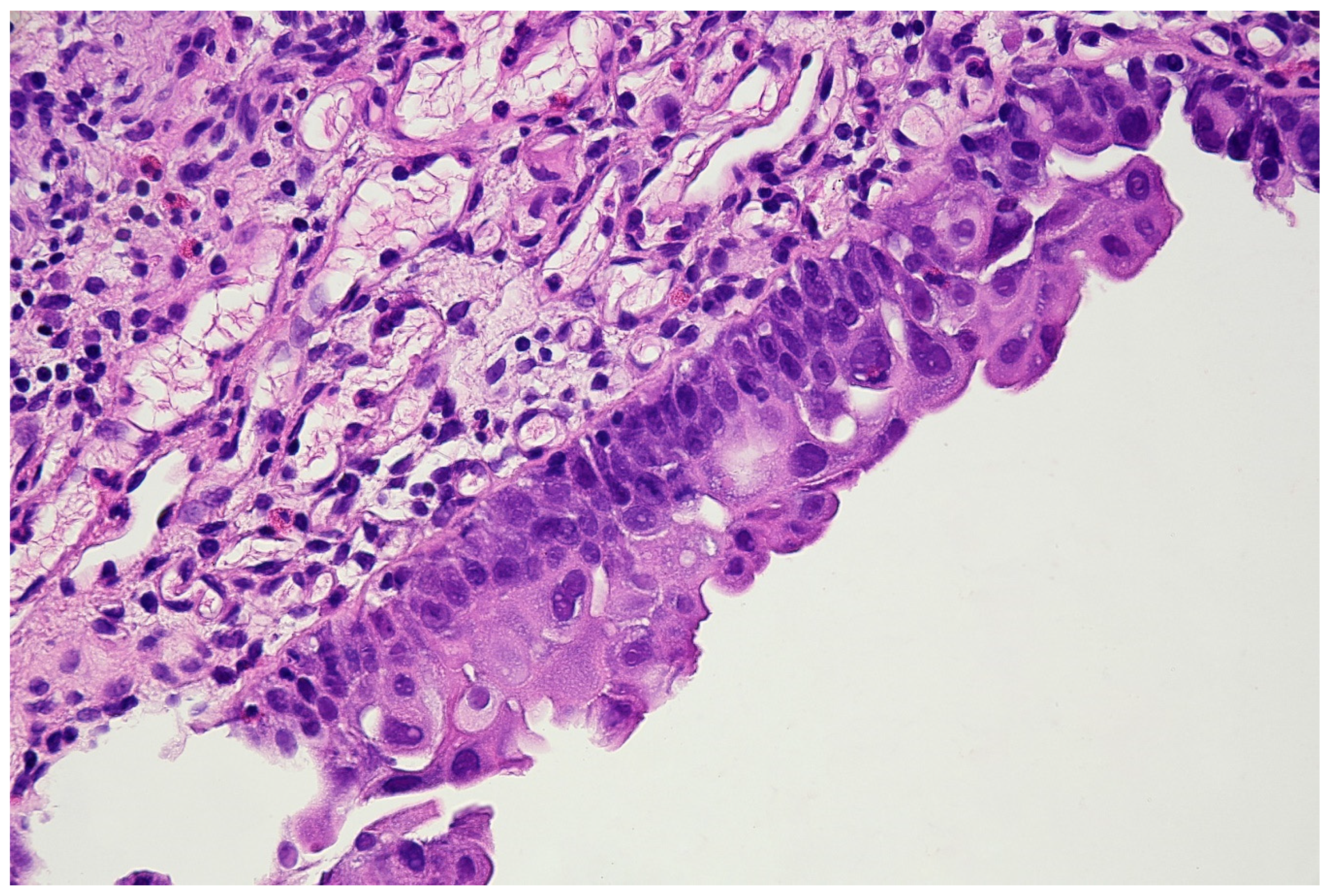
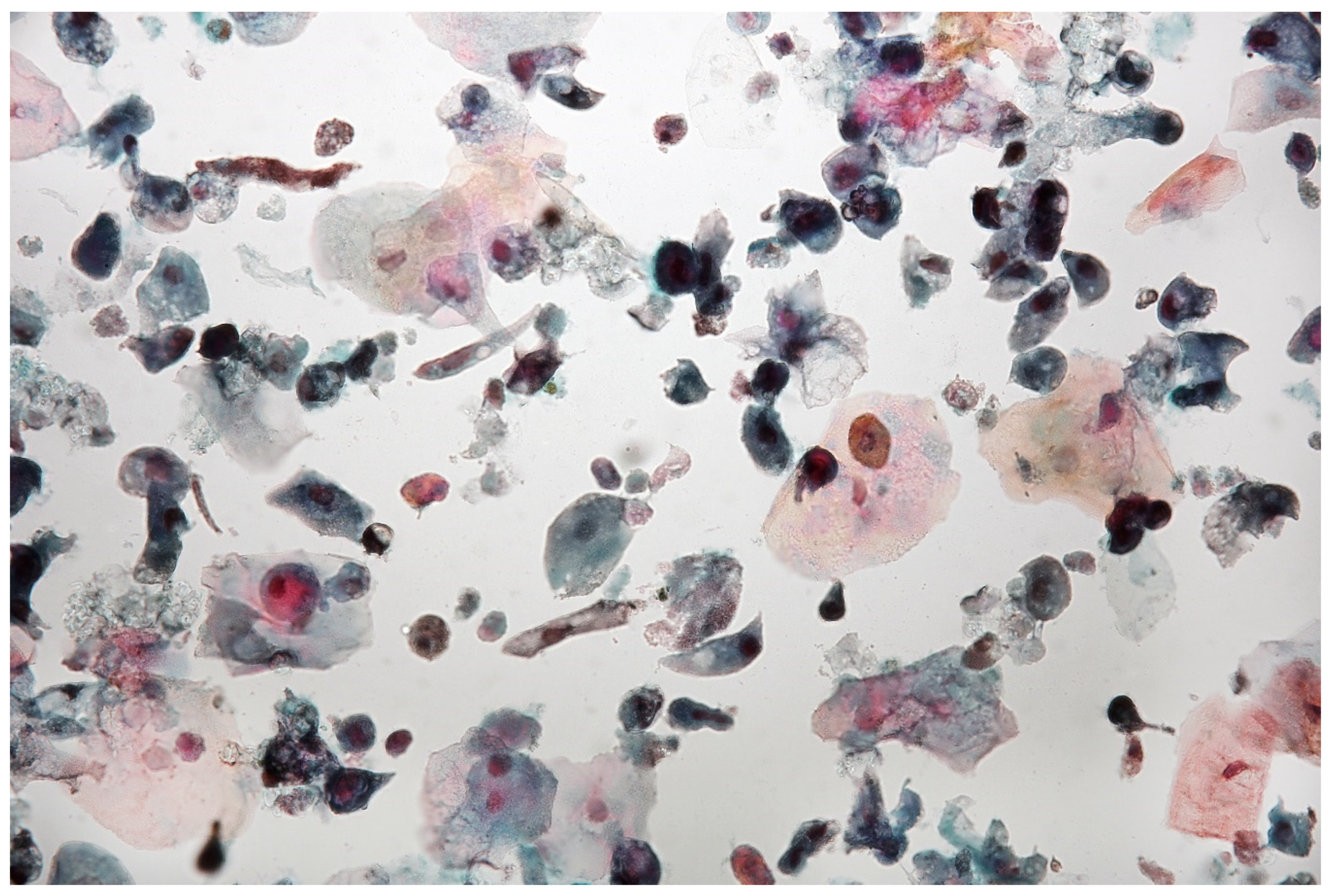
2.1.1. Urinary Cytology
2.1.2. FISH
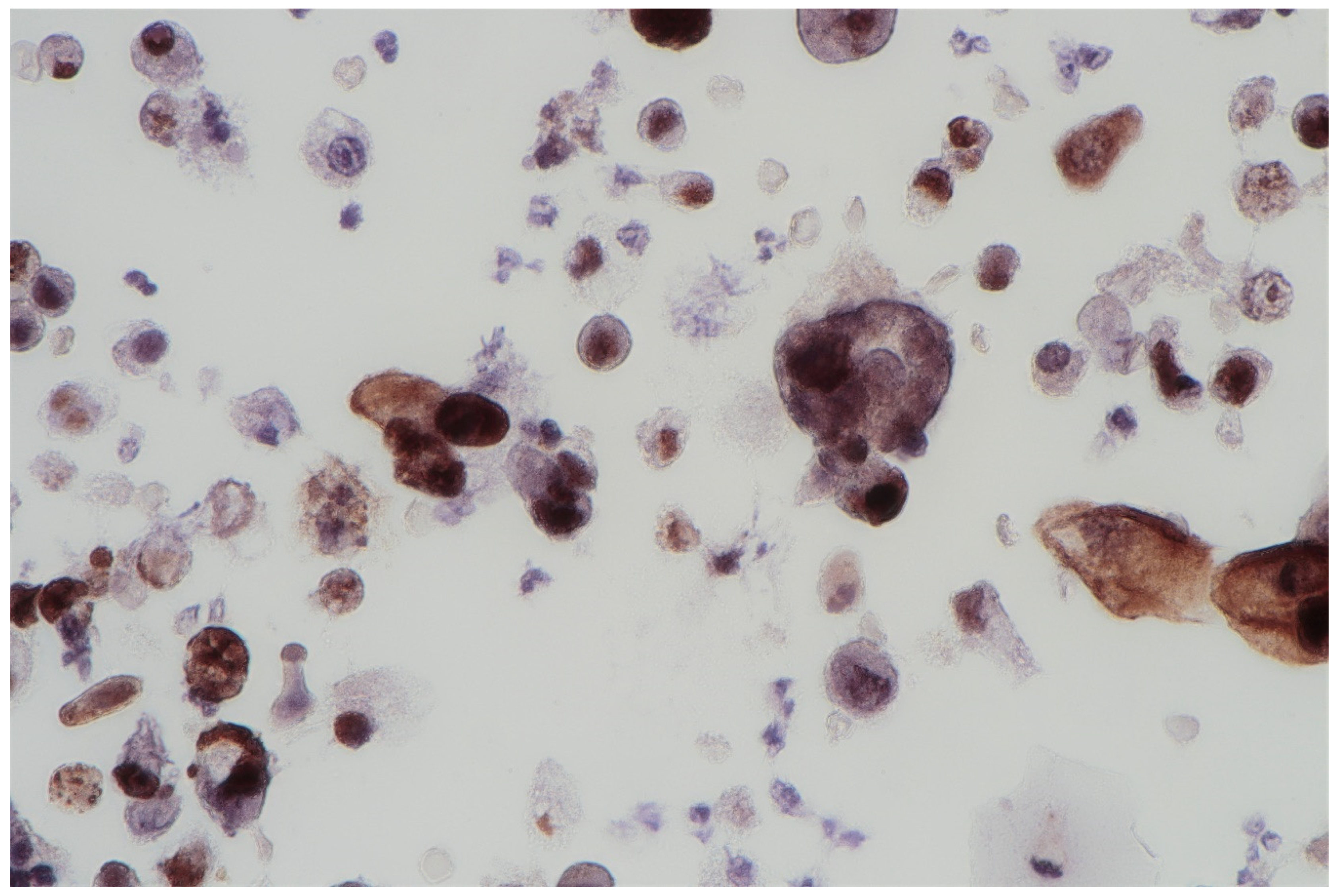
2.1.3. Immunocytochemical Staining
2.2. Bladder EpiCheck
2.3. DNA Isolation and Whole Genome Sequencing (WGS)
2.4. Bioinformatic Analysis
3. Results
3.1. Cytourofish(+)
3.2. Bladder EpiCheck
3.3. Whole Genome Sequencing (WGS)
3.3.1. Structural Variations (SV)
3.3.2. Single Nucleotide Variations, Deletions and Insertions
3.4. Further Proceeding
3.5. Non-Invasive Monitoring
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A. FISH Procedure
Appendix A.1. Sample Collection and Slides Preparation
- Sample collection: 700 mL of fresh urine, placed in a 50 mL tip centrifuge tube and centrifuged at room temperature with 2000 rpm for 10 min.
- Cell harvest: The supernatant was carefully discarded, leaving a few milliliters.
- Step 1 and 2 were repeated until the entire delivered urine sample was concentrated.
- The cytospin technique was used to perform a thin-layer cytological preparation.
- The slide was fixed in 95% ethyl alcohol (POCH; Gliwice; Poland).
Appendix A.2. Slide Processing
- The slide was soaked for 2 min in the 2 × SSC buffer (Abbott; Wiesbaden; Germany) at 73 °C.
- The slide was incubated for 10 min in pepsin solution (Pan Path; Budel; The Netherlands) at 37 °C.
- The slide was washed for 1 × 5 min in PBS (Biological Industries; Kibbutz Beit Haemek; Israel).
- The slide was washed for 1 × 5 min in 1% formaldehyde (POL-AURA; Olsztyn; Poland) at room temperature.
- The slide was washed for 1 × 5 min in PBS at room temperature.
- The slide was dehydrated in gradient alcohol (70%, 85%, 100%) for 1 min each.
- The slide was allowed to air dry.
Appendix A.3. Denaturation and Hybridisation
- The probe was taken out from a freezer and placed at room temperature for 5 min.
- The probe was mixed and briefly centrifuged.
- 5 µL of the probe was taken and dropped onto the slide, which immediately was covered with a 22 × 22 mm coverslip (Knittel Glass; Braunschweig; Germany), avoiding air bubbles.
- The slide was placed in a hybridiser: denaturation at 80 °C for 10 min; hybridisation at 37 °C for 24 h.
Appendix A.4. Posthybridisation Washes
- The slide was immersed in 2xSSC/0.3% NP-40 buffer (Pan Path; Budel; Netherlands) for about 5 s to gently remove the cover slide.
- The slide was placed in a Coplin jar with 2xSSC/0.3% NP-40 buffer in a 72 °C water bath for 2 min.
- The slide was washed again in 2xSSC/0.1% NP-40 buffer at room temperature for 1 min.
Appendix A.5. Counterstaining
- 10 µL of Mounting Medium Fluorescence with DAPI (Pan Path; Budel; the Netherlands) was placed on a dry glass slide and immediately covered and sealed with nail polish.
- The slide was stored at 4 °C in the dark for 1 day and then was analysed.
Appendix B. Imunnocytochemical Procedure
Appendix B.1. Sample Collection and Slide Preparation
- Sample collection: 700 mL of fresh urine was placed in a 50 mL tip centrifuge tube and centrifuged at room temperature with 2000 rpm for 10 min.
- Cell harvest: The supernatant was carefully discarded, leaving a few milliliters.
- Step a and b were repeated until the entire delivered urine sample was concentrated.
- The cytospin technique was used to obtain a thin-layer cytological preparation.
- The slide was fixed in 95% ethyl alcohol (POCH; Gliwice; Poland).
Appendix B.2. IHC Procedure
- Formalin (POL-AURA; Olsztyn; Poland) fixation at room temperature for 15 min.
- Block endogenous peroxidase by incubating the slide with 3% hydrogen peroxide (POCH; Gliwice; Poland) at room temperature for 5 min.
- It was rinsed in tap water for 5 min.
- The antigen was retrieved by incubating the slide in TRS buffer (Leica; Wetzlar; Germany) and preheated to 99 °C in water bath for 40 min.
- The slide was cooled in tap water.
- The slide was washed for 1 × 5 min in TBS buffer (Mar-Four; Konstantynow Lodzki; Poland).
- In order to minimise false-positive staining, the slide was preincubated with Normal Goat Antibody (Vector Laboratories; Burlingame; United States) at room temperature for 20 min at a moist chamber.
- The excess of Normal Goat Antibody was shaken off.
- The primary antibody was added (GenomeMe; Geel; Belgium) and the slide was incubated at a moist chamber for 30 min.
- The slide was washed for 1 × 5 min in TBS buffer.
- The secondary antibody (GenomeMe; Geel; Belgium) was added and the slide was incubated at the moist chamber for 30 min.
- The slide was washed for 1 × 5 min in TBS buffer.
- A substrate—DAB (Vector Laboratories; Burlingame; United States) was added and the slide was incubated at the moist chamber for 5 min.
- It was then rinsed in tap water for 2 min.
- Haematoxylin (Sigma-Aldrich; Missouri; United States) staining was carried out for 5 to 10 min.
- The slide was dehydrated in gradient alcohol (70%, 80%, 100%) for 5 s each.
- A coverslip with Entellan (Merck; Darmstadt; Germany) was used.
Appendix C
| Gene | Localisation hg19 | Reference Sequence | Variant |
|---|---|---|---|
| TP53 | chr17:7,578,406 | NM_001126114.2 | c.524G > T (p.Arg175Leu) |
| KDM6B | chr17:7,752,523 | NM_001080424.2 | c.2917A > C (p.Lys973Gln) |
| FGFR3 | - | - | - |
| RXRA | - | - | - |
| ERBB2 | chr17:37,879,885 | NM_001382805.1 | c.2180G > C (p.Gly727Ala) |
| HRAS | - | - | - |
| RB1 | chr13:49,054,146 | NM_000321.3 | c.2726C > T (p.Thr909Ile) |
| chr13:48,942,667 | NM_000321.3 | c.1054G > C (p.Glu352Gln) | |
| CDKN2A | - | - | - |
| KRAS | - | - | - |
| IDH1 | - | - | - |
| MLL2 (KMT2D) | - | - | - |
| MDM2 | - | - | - |
| CCND1 | - | - | - |
| CCNE1 | - | - | - |
| E2F3 | - | - | - |
| SETD2 | - | - | - |
| KDM6A | - | - | - |
| CREBBP | - | - | - |
| EP300 | - | - | - |
| ARID1A | - | - | - |
| ARID1B | - | - | - |
| EGFR | - | - | - |
| ERBB3 | - | - | - |
| PIK3CA | - | - | - |
| NRAS | - | - | - |
| NF1 | - | - | - |
| TSC1 | - | - | - |
| TSC2 | - | - | - |
| PTEN | - | - | - |
| AKT3 | - | - | - |
| STAG2 | - | - | - |
| ERCC2 | - | - | - |
| NFE2L2 | - | - | - |
| TXNIP | - | - | - |
| RHOA | - | - | - |
| RHOB | - | - | - |
| TERT | - | - | - |
| SMARCA2 | - | - | - |
| SMARCA4 | chr19:11,118,640 | NM_001128849.3 | c.2064G > A (p.Lys688=) |
| chr19:11,169,514 | NM_001128849.3 | c.4680C > T (p.Asp1560=) | |
| SMARCC1 | - | - | - |
| SMARCC2 | - | - | - |
| PPARG | - | - | - |
| NECTIN4 | - | - | - |
| MYC | - | - | - |
| YWHAZ | - | - | - |
| GATA3 | - | - | - |
| FOXA1 | - | - | - |
| UPK3A | - | - | - |
| KRT14 | - | - | - |
| KRT5 | - | - | - |
| KRT6A | - | - | - |
| BRAF | - | - | - |
| RYR1 | - | - | - |
| STXBP1 | chr9:130,438,099 | NM_003165.6 | c.1127C > T (p.Thr376Ile) |
| CSRP1 | - | - | - |
| SLC26A4 | - | - | - |
| MTOR | - | - | - |
| FBN1 | - | - | - |
| PEX7 | - | - | - |
| FANCA | - | - | - |
| B9D2 | - | - | - |
| LAMA3 | - | - | - |
| LTBP2 | chr14:74,991,855 | NM_000428.3 | c.2502T > C (p.Thr834=) |
| LARS2 | - | - | - |
| FGFR2 | - | - | - |
| RP1 | - | - | - |
| IDUA | chr4:997,162 | NM_000203.5 | c.1554A > C (p.Leu518Phe) |
| CDKN1A | - | - | - |
| FBXW7 | - | - | - |
| ELF3 | - | - | - |
| KLF5 | - | - | - |
| FOXQ1 | - | - | - |
| PAIP1 | - | - | - |
| BTG2 | - | - | - |
| ZFP36L1 | - | - | - |
| CCND3 | - | - | - |
| NCOR1 | chr17:16,097,825 | NM_006311.4 | c.59A > C (p.Tyr20Ser) |
| YAP1 | - | - | - |
| ZNF703 | - | - | - |
| MYCL | - | - | - |
| MLL (KMT2A) | - | - | - |
| MLL3 (KMT2C) | - | - | - |
| MLL4 (KMT2B) | - | - | - |
| BCL2L1 | - | - | - |
Appendix D
| Frequently Altered Genes and Pathways in Bladder Cancer | |||||
|---|---|---|---|---|---|
| Pathway | Occurrence | Gene | Mechanism | Frequency | Presence |
| TP53/RB1 pathway | >90% | TP53 | Inactivating mutations, deletion | >50% | Yes |
| CKDN2A | Deletion | ~50% | Yes | ||
| RB1 | Inactivating mutations, deletion | ~10% | Effect of detected RB1 mutation is unknown | ||
| MDM2 | Amplification | ~10% | Yes | ||
| CCND1 | Amplification | ~10% | Yes | ||
| CCNE1 | Amplification | ~10% | Yes | ||
| E2F3 | Amplification | ~20% | Yes | ||
| Chromatin remodelling | >90% | MLL/MLL2/MLL3/MLL4 | Inactivating mutations, deletion | >30% | - |
| SETD2 | Inactivating mutations, deletion | ~10% | - | ||
| KDM6A/KDM6B | Inactivating mutations, deletion | >30% | Yes | ||
| CREBBP | Inactivating mutations, deletion | >10% | Yes | ||
| EP300 | Inactivating mutations, deletion | >10% | Yes | ||
| ARID1A/ARID1B | Inactivating mutations, deletion | >30% | Yes | ||
| SMARCA2/SMARCA4 | Inactivating mutations, deletion | >20% | Yes | ||
| SMARCC1/SMARCC2 | Inactivating mutations, deletion | >20% | - | ||
| RTK/RAS/PI(3)K pathway | >70% | FGFR3 | Activating mutations, gene fusions, amplification | ~15% | Yes |
| EGFR | Amplification | ~10% | - | ||
| ERBB2 | Activating mutations, gene fusions, amplification | ~10% | Yes | ||
| ERBB3 | Activating mutations | ~10% | - | ||
| PIK3CA | Activating mutations, Amplification | >10% | Yes | ||
| NRAS/HRAS/KRAS | Activating mutations | ~10% | - | ||
| NF1 | Inactivating mutations | ~5% | - | ||
| TSC1/TSC2 | Inactivating mutations, deletion | ~10% | Yes | ||
| PTEN | Inactivating mutations, deletion | ~10% | - | ||
| AKT3 | Overexpression | 10% | - | ||
| Miscellaneous | N/A | STAG2 | Inactivating mutations, large deletion, hypermethylation | >10% | - |
| ERCC2 | Inactivating missense mutation | ~10% | - | ||
| RXRA | Activating mutations | ~10% | - | ||
| NFE2L2 | Inactivating missense mutation | ~10% | - | ||
| TXNIP | Inactivating mutations | ~7% | - | ||
| RHOA/RHOB | Missense mutation | ~10% | - | ||
| Description | Characteristic in Details | Pathology | Presence |
|---|---|---|---|
| Focally amplified | Focal somatic amplifications (E2F2/SOX4, EGFR, PPARG, PVRL4, YWHAZ, MYC) and MLL2 mutations | - | Average probability |
| CDKN2A-deficient FGFR3 mutant | Copy number loss of CDKN2A, activating mutations of FGFR3 | Enriched in papillary histology | Average probability |
| TP53/cell-cycle mutant | TP53 mutations, RB1 mutations, amplifications of E2F3 and CCNE1 | - | High probability |
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021. [Google Scholar] [CrossRef]
- Wojciechowska, U.; Didkowska, J.; Michałek, I.; Olasek, P.; Ciuba, A. Nowotwory Złośliwe w Polsce w 2018 Roku|Cancer in Poland. In Nowotwory złośliwe w Polsce w … roku; Narodoway Instytut Onkologii: Warsaw, Poland, 2019; ISSN 0867-8251. [Google Scholar]
- Bosetti, C.; Bertuccio, P.; Chatenoud, L.; Negri, E.; La Vecchia, C.; Levi, F. Trends in Mortality from Urologic Cancers in Europe, 1970–2008. Eur. Urol. 2011, 60, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Rosenthal, D.L.; Wojcik, E.M.; Kurtycz, D.F.I. (Eds.) The Paris System for Reporting Urinary Cytology; Springer International Publishing: Cham, Switzerland, 2016; ISBN 978-3-319-22863-1. [Google Scholar]
- Mancini, M.; Righetto, M.; Zumerle, S.; Montopoli, M.; Zattoni, F. The Bladder EpiCheck Test as a Non-Invasive Tool Based on the Identification of DNA Methylation in Bladder Cancer Cells in the Urine: A Review of Published Evidence. Int. J. Mol. Sci. 2020, 21, 6542. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative Genomics Viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koboldt, D.C.; Chen, K.; Wylie, T.; Larson, D.E.; McLellan, M.D.; Mardis, E.R.; Weinstock, G.M.; Wilson, R.K.; Ding, L. VarScan: Variant Detection in Massively Parallel Sequencing of Individual and Pooled Samples. Bioinforma. Oxf. Engl. 2009, 25, 2283–2285. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup the Sequence Alignment/Map Format and SAMtools. Bioinforma. Oxf. Engl. 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rausch, T.; Zichner, T.; Schlattl, A.; Stütz, A.M.; Benes, V.; Korbel, J.O. DELLY: Structural Variant Discovery by Integrated Paired-End and Split-Read Analysis. Bioinformatics 2012, 28, i333–i339. [Google Scholar] [CrossRef] [PubMed]
- Ramírez, F.; Ryan, D.P.; Grüning, B.; Bhardwaj, V.; Kilpert, F.; Richter, A.S.; Heyne, S.; Dündar, F.; Manke, T. DeepTools2: A next Generation Web Server for Deep-Sequencing Data Analysis. Nucleic Acids Res. 2016, 44, W160–W165. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An Information Aesthetic for Comparative Genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clark, P.E. Bladder Cancer. Curr. Opin. Oncol. 2007, 19, 241–247. [Google Scholar] [CrossRef] [PubMed]
- Seok Ju, Y. The Mutational Signatures and Molecular Alterations of Bladder Cancer. Transl. Cancer Res. 2017, 6, S689–S701. [Google Scholar] [CrossRef]
- Lamm, D.L.; Blumenstein, B.A.; Crissman, J.D.; Montie, J.E.; Gottesman, J.E.; Lowe, B.A.; Sarosdy, M.F.; Bohl, R.D.; Grossman, H.B.; Beck, T.M.; et al. Maintenance Bacillus Calmette-Guerin Immunotherapy for Recurrent TA, T1 and Carcinoma in Situ Transitional Cell Carcinoma of the Bladder: A Randomized Southwest Oncology Group Study. J. Urol. 2000, 163, 1124–1129. [Google Scholar] [CrossRef]
- Babjuk, M.; Burger, M.; Compérat, E.M.; Gontero, P.; Mostafid, A.H.; Palou, J.; van Rhijn, B.W.G.; Rouprêt, M.; Shariat, S.F.; Sylvester, R.; et al. European Association of Urology Guidelines on Non-Muscle-Invasive Bladder Cancer (TaT1 and Carcinoma In Situ)—2019 Update. Eur. Urol. 2019, 76, 639–657. [Google Scholar] [CrossRef] [PubMed]
- Daneshmand, S.; Bazargani, S.T.; Bivalacqua, T.J.; Holzbeierlein, J.M.; Willard, B.; Taylor, J.M.; Liao, J.C.; Pohar, K.; Tierney, J.; Konety, B.; et al. Blue Light Cystoscopy for the Diagnosis of Bladder Cancer: Results from the US Prospective Multicenter Registry. Urol. Oncol. 2018, 36, 361.e1–361.e6. [Google Scholar] [CrossRef] [PubMed]
- Kaluzewski, T.; Morzuch, L.; Jedrzejczyk, A.; Marks, P.; Rozniecki, M.; Bednarek, M. Advantages of Fluorescence in Situ Hybridisation Technique in Urothelial Carcinogenesis Identification. Retrospective Evaluation of 512 Urovysion Test Results. MEDtube Sci. 2018, 7, 8–13. [Google Scholar]
- Cousin, S.; Khalifa, E.; Crombe, A.; Laizet, Y.; Lucchesi, C.; Toulmonde, M.; Le Moulec, S.; Auzanneau, C.; Soubeyran, I.; Italiano, A. Targeting ERBB2 Mutations in Solid Tumors: Biological and Clinical Implications. J. Hematol. Oncol. 2018, 11, 86. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murphree, A.L.; Benedict, W.F. Retinoblastoma: Clues to Human Oncogenesis. Science 1984, 223, 1028–1033. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, J.N.; Akbani, R.; Broom, B.M.; Wang, W.; Verhaak, R.G.W.; McConkey, D.; Lerner, S.; Morgan, M.; Creighton, C.J.; Smith, C.; et al. Comprehensive Molecular Characterization of Urothelial Bladder Carcinoma. Nature 2014, 507, 315–322. [Google Scholar] [CrossRef] [Green Version]
- Choi, W.; Porten, S.; Kim, S.; Willis, D.; Plimack, E.R.; Hoffman-Censits, J.; Roth, B.; Cheng, T.; Tran, M.; Lee, I.-L.; et al. Identification of Distinct Basal and Luminal Subtypes of Muscle-Invasive Bladder Cancer with Different Sensitivities to Frontline Chemotherapy. Cancer Cell 2014, 25, 152–165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Witjes, J.A.; Morote, J.; Cornel, E.B.; Gakis, G.; van Valenberg, F.J.P.; Lozano, F.; Sternberg, I.A.; Willemsen, E.; Hegemann, M.L.; Paitan, Y.; et al. Performance of the Bladder EpiCheckTM Methylation Test for Patients Under Surveillance for Non-Muscle-Invasive Bladder Cancer: Results of a Multicenter, Prospective, Blinded Clinical Trial. Eur. Urol. Oncol. 2018, 1, 307–313. [Google Scholar] [CrossRef]
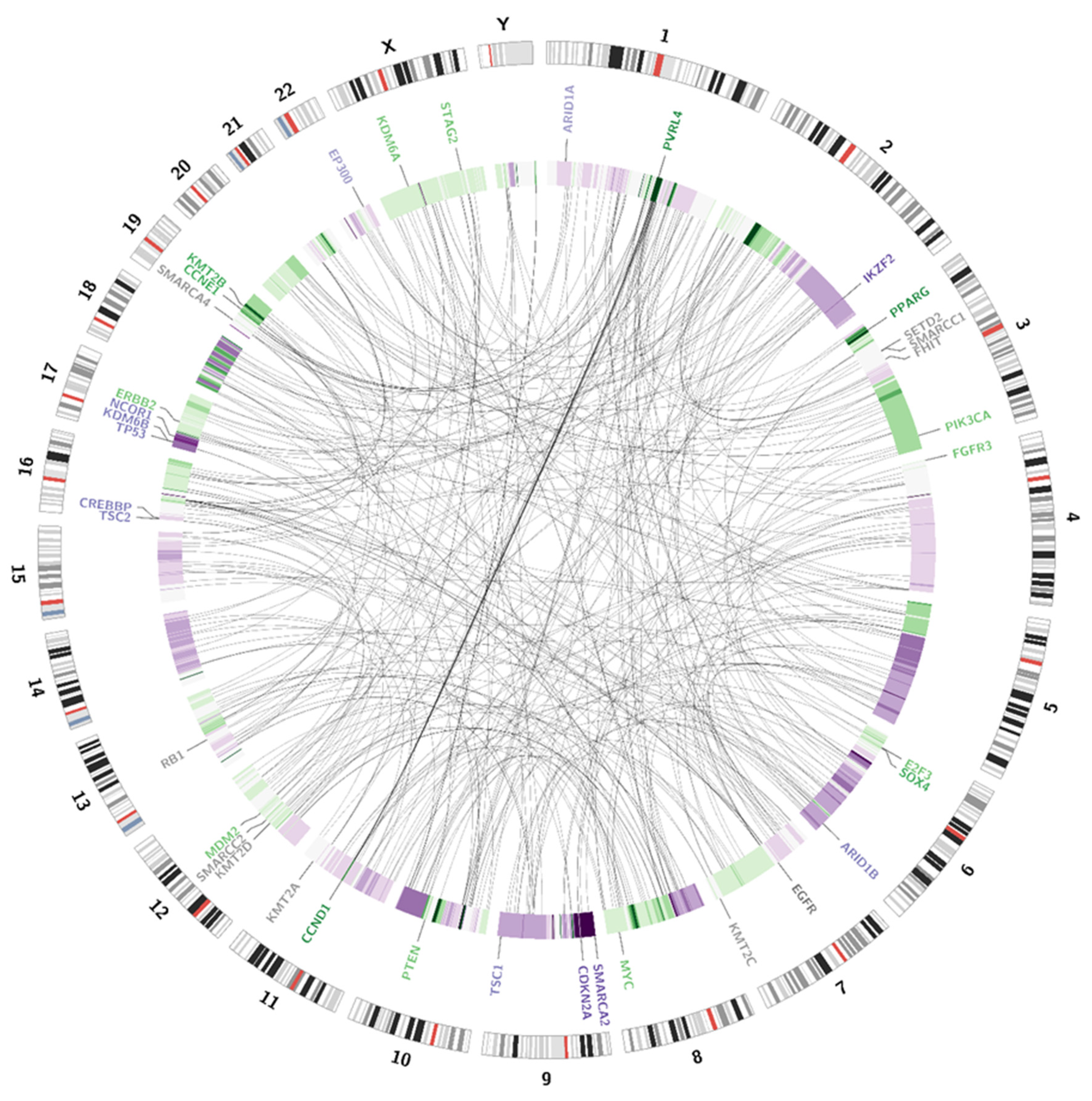

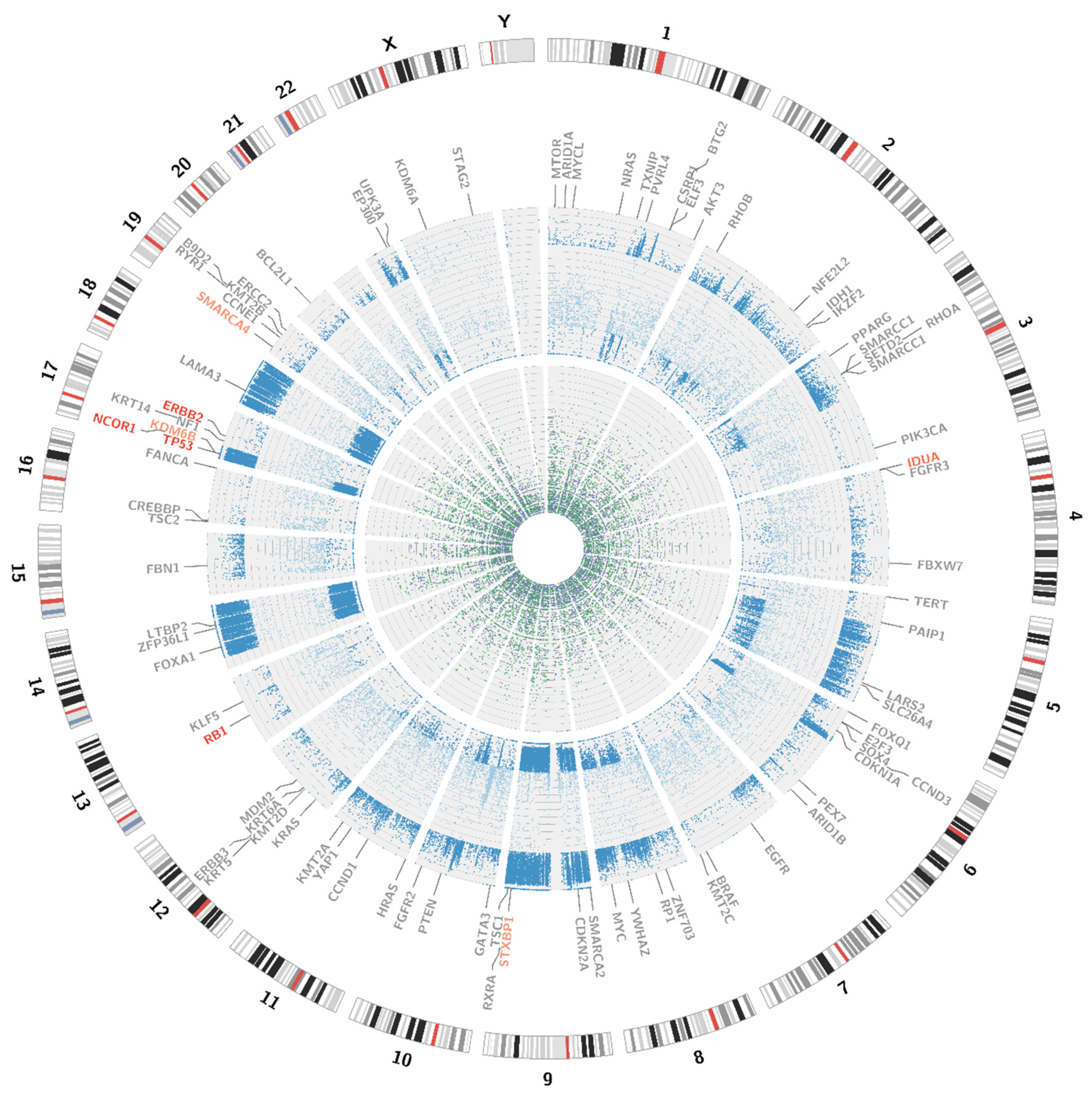


| Amplifications | ||
|---|---|---|
| Chromosomal Locus | Gene | Presence in the Evaluated Case |
| 6p22.3 | E2F3 | Yes |
| 6p22.3 | SOX4 | Yes |
| 11q13.3 | CCND1 | Yes |
| 7p11.2 | EGFR | - |
| 17q12 | ERBB2 | Yes |
| 19q12 | CCNE1 | Yes |
| 3p25.2 | PPARG | Yes |
| 12q15 | MDM2 | Yes |
| 8q24.21 | MYC | Yes |
| 4p16.3 | FGFR3 | Yes |
| 1q23.3 | PVRL4 | Yes |
| 3q26.32 | PIK3CA | Yes |
| Deletions | ||
| 9p21.3 | CDKN2A | Yes |
| 2q34 | IKZF2 | Yes |
| 13q14.2 | RB1 | - |
| 3p14.2 | FHIT | - |
| 16p13.3 | CREBBP | Yes |
| 17p12 | NCOR1 | Yes |
| 12q13.12 | KMT2D | - |
| 11q23.3 | KMT2A | - |
| 7q36.1 | KMT2C | - |
| 19q13.12 | KMT2B | - |
| 22q13.2 | EP300 | Yes |
| 1p36.11 | ARID1A | Yes |
| 6q25.3 | ARID1B | Yes |
| 9p24.3 | SMARCA2 | Yes |
| 19p13.2 | SMARCA4 | - |
| 3p21.31 | SMARCC1 | - |
| 12q13.2 | SMARCC2 | - |
| 9q34.13 | TSC1 | Yes |
| 16p13.3 | TSC2 | Yes |
| 10q23.31 | PTEN | - |
| 3p21.31 | SETD2 | - |
| 17p13.1 | KDM6B | Yes |
| Xp11.3 | KDM6A | - |
| Xq25 | STAG2 | - |
| 17p13.1 | TP53 | Yes |
| Gene | Variant | Percentage of Reads Blood Sample | Percentage of Reads Urine Sample | Clinical Significance |
|---|---|---|---|---|
| TP53 | c.524G > T (p.Arg175Leu) | 0% | 78% | Pathogenic (LOH) |
| ERBB2 | c.2180G > C (p.Gly727Ala) | 0% | 28% | Likely pathogenic |
| RB1 | c.2726C > T (p.Thr909Ile) | 0% | 33% | Uncertain significance |
| c.1054G > C (p.Glu352Gln) | 0% | 36% | Uncertain significance | |
| KDM6B | c.2917A > C (p.Lys973Gln) | 59% | 89% | Benign (LOH) |
| STXBP1 | c.1127C > T (p.Thr376Ile) | 50% | 93% | Uncertain significance (LOH) |
| IDUA | c.1554A > C (p.Leu518Phe) | 14% | 29% | Uncertain significance |
| SMARCA4 | c.2064G > A (p.Lys688=) | 0% | 38% | Benign |
| c.4680C > T (p.Asp1560=) | 35% | 74% | Likely benign (LOH) | |
| LTBP2 | c.2502T > C (p.Thr834=) | 48% | 89% | Benign (LOH) |
| NCOR1 | c.59A > C (p.Tyr20Ser) | 14% | 29% | Uncertain significance |
| Primary Result | Follow-Up Result after 3 Months | Follow-Up Result after 6 Months | Follow-Up Result after 12 Months |
|---|---|---|---|
| Positive, EpiScore: 92 | Negative, EpiScore: 21 | Negative, EpiScore: 17 | Negative, EpiScore: 7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kałużewski, T.; Przybylski, G.K.; Bednarek, M.; Glazar, S.; Grabiec, M.; Jędrzejczyk, A.; Kępczyński, Ł.; Kubiak, I.; Kucharska, D.; Morel, A.; et al. The Usefulness of Cell-Based and Liquid-Based Urine Tests in Clarifying the Diagnosis and Monitoring the Course of Urothelial Carcinoma. Identification of Novel, Potentially Actionable, RB1 and ERBB2 Somatic Mutations. J. Pers. Med. 2021, 11, 362. https://doi.org/10.3390/jpm11050362
Kałużewski T, Przybylski GK, Bednarek M, Glazar S, Grabiec M, Jędrzejczyk A, Kępczyński Ł, Kubiak I, Kucharska D, Morel A, et al. The Usefulness of Cell-Based and Liquid-Based Urine Tests in Clarifying the Diagnosis and Monitoring the Course of Urothelial Carcinoma. Identification of Novel, Potentially Actionable, RB1 and ERBB2 Somatic Mutations. Journal of Personalized Medicine. 2021; 11(5):362. https://doi.org/10.3390/jpm11050362
Chicago/Turabian StyleKałużewski, Tadeusz, Grzegorz K. Przybylski, Michał Bednarek, Sławomir Glazar, Magdalena Grabiec, Adam Jędrzejczyk, Łukasz Kępczyński, Izabela Kubiak, Dorota Kucharska, Agnieszka Morel, and et al. 2021. "The Usefulness of Cell-Based and Liquid-Based Urine Tests in Clarifying the Diagnosis and Monitoring the Course of Urothelial Carcinoma. Identification of Novel, Potentially Actionable, RB1 and ERBB2 Somatic Mutations" Journal of Personalized Medicine 11, no. 5: 362. https://doi.org/10.3390/jpm11050362








