Multi-Omics Analysis of SOX4, SOX11, and SOX12 Expression and the Associated Pathways in Human Cancers
Abstract
:1. Introduction
2. Results
2.1. mRNA Expression of the SOX Gene Family in Various Cancers
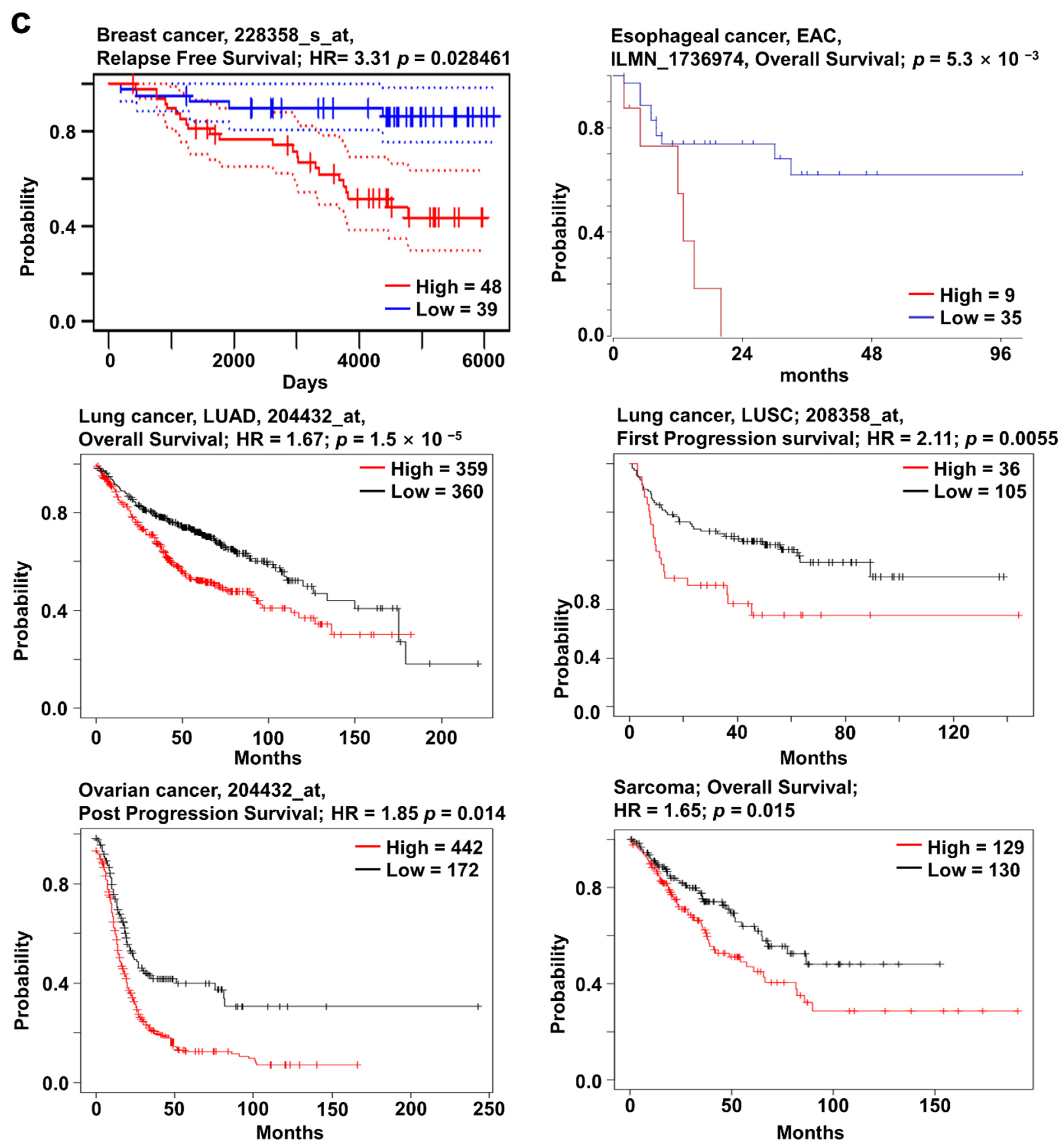
2.2. SOX4 Expression and Its Prognostic Value in Various Cancers
2.3. SOX11 Expression Pattern and Patient Survival in Various Cancers
2.4. SOX12 Expression Pattern and Patient Survival in Various Cancers
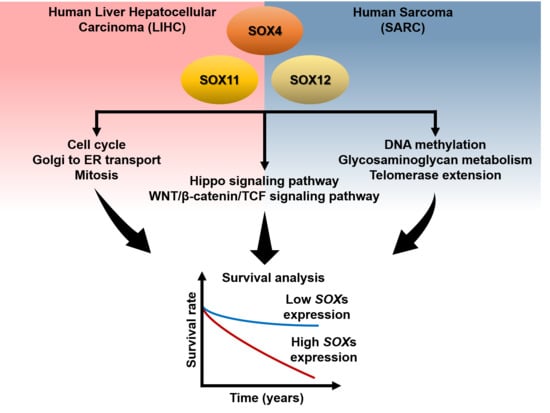
2.5. Clinical Prognosis of the Three SOX Genes in Liver Cancer
2.6. Clinical Prognosis of the Three SOX Genes in Sarcoma
2.7. Functional Protein Partners and Their Predicted Signaling Pathways
3. Discussion
4. Materials and Methods
4.1. Oncomine Database Analysis
4.2. Molecular Evolutionary Genetics Analysis
4.3. GEPIA2 Database Analysis
4.4. cBioPortal Database Analysis
4.5. Kaplan–Meier Plotter
4.6. R2: Genomic Analysis and Visualization Platform
4.7. Gene Correlation Analysis of SOX4, SOX11, and SOX12
4.8. Identification of Functional Protein Partners of SOX4, SOX11, and SOX12, and Signaling Pathway Analysis
4.9. Analysis of SOX Protein Expression Pattern in Various Types of Cancer
4.10. Analysis of Gene Network with SOX4, SOX11, and SOX12
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Global Burden of Disease Cancer Collaboration; Fitzmaurice, C.; Abate, D.; Abbasi, N.; Abbastabar, H.; Abd-Allah, F.; Abdel-Rahman, O.; Abdelalim, A.; Abdoli, A.; Abdollahpour, I.; et al. Global, Regional, and National Cancer Incidence, Mortality, Years of Life Lost, Years Lived With Disability, and Disability-Adjusted Life-Years for 29 Cancer Groups, 1990 to 2017: A Systematic Analysis for the Global Burden of Disease Study. JAMA Oncol. 2019, 5, 1749–1768. [Google Scholar] [CrossRef] [Green Version]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [Green Version]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lefebvre, V.; Dumitriu, B.; Penzo-Méndez, A.; Han, Y.; Pallavi, B. Control of cell fate and differentiation by Sry-related high-mobility-group box (Sox) transcription factors. Int. J. Biochem. Cell Biol. 2007, 39, 2195–2214. [Google Scholar] [CrossRef] [Green Version]
- Kamachi, Y.; Kondoh, H. Sox proteins: Regulators of cell fate specification and differentiation. Development 2013, 140, 4129–4144. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, H.; Song, C.; Liu, P.; Wang, J.; Chen, B.; Huang, X.; Pei, X.; Liu, L. SOX2 promotes cell proliferation and metastasis in triple negative breast cancer. Front. Pharmacol. 2018, 9, 942. [Google Scholar]
- Yang, F.; Gao, Y.; Geng, J.; Qu, D.; Han, Q.; Qi, J.; Chen, G. Elevated expression of SOX2 and FGFR1 in correlation with poor prognosis in patients with small cell lung cancer. Int. J. Clin. Exp. Pathol. 2013, 6, 2846. [Google Scholar] [PubMed]
- Gu, W.; Wang, B.; Wan, F.; Wu, J.; Lu, X.; Wang, H.; Zhu, Y.; Zhang, H.; Shi, G.; Dai, B. SOX2 and SOX12 are predictive of prognosis in patients with clear cell renal cell carcinoma. Oncol. Lett. 2018, 15, 4564–4570. [Google Scholar] [CrossRef]
- Wang, X.; Ji, X.; Chen, J.; Yan, D.; Zhang, Z.; Wang, Q.; Xi, X.; Feng, Y. SOX2 enhances the migration and invasion of ovarian cancer cells via Src kinase. PLoS ONE 2014, 9, e99594. [Google Scholar] [CrossRef]
- Sun, C.; Sun, L.; Li, Y.; Kang, X.; Zhang, S.; Liu, Y. Sox2 expression predicts poor survival of hepatocellular carcinoma patients and it promotes liver cancer cell invasion by activating Slug. Med. Oncol. 2013, 30, 503. [Google Scholar] [CrossRef]
- Shen, H.; Blijlevens, M.; Yang, N.; Frangou, C.; Wilson, K.E.; Xu, B.; Zhang, Y.; Zhang, L.; Morrison, C.D.; Shepherd, L. Sox4 expression confers bladder cancer stem cell properties and predicts for poor patient outcome. Int. J. Biol. Sci. 2015, 11, 1363. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vervoort, S.J.; de Jong, O.G.; Roukens, M.G.; Frederiks, C.L.; Vermeulen, J.F.; Lourenço, A.R.; Bella, L.; Vidakovic, A.T.; Sandoval, J.L.; Moelans, C. Global transcriptional analysis identifies a novel role for SOX4 in tumor-induced angiogenesis. Elife 2018, 7, e27706. [Google Scholar] [CrossRef] [PubMed]
- Song, G.-D.; Sun, Y.; Shen, H.; Li, W. SOX4 overexpression is a novel biomarker of malignant status and poor prognosis in breast cancer patients. Tumor Biol. 2015, 36, 4167–4173. [Google Scholar] [CrossRef] [PubMed]
- Ramezani-Rad, P.; Geng, H.; Hurtz, C.; Chan, L.N.; Chen, Z.; Jumaa, H.; Melnick, A.; Paietta, E.; Carroll, W.L.; Willman, C.L. SOX4 enables oncogenic survival signals in acute lymphoblastic leukemia. Blood 2013, 121, 148–155. [Google Scholar] [CrossRef] [Green Version]
- Wan, Y.-P.; Xi, M.; He, H.-C.; Wan, S.; Hua, W.; Zen, Z.-C.; Liu, Y.-L.; Zhou, Y.-L.; Mo, R.-J.; Zhuo, Y.-J. Expression and clinical significance of SOX9 in renal cell carcinoma, bladder cancer and penile cancer. Oncol. Res. Treat. 2017, 40, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Richtig, G.; Aigelsreiter, A.; Schwarzenbacher, D.; Ress, A.L.; Adiprasito, J.B.; Stiegelbauer, V.; Hoefler, G.; Schauer, S.; Kiesslich, T.; Kornprat, P. SOX9 is a proliferation and stem cell factor in hepatocellular carcinoma and possess widespread prognostic significance in different cancer types. PLoS ONE 2017, 12, e0187814. [Google Scholar] [CrossRef] [Green Version]
- Lü, B.; Fang, Y.; Xu, J.; Wang, L.; Xu, F.; Xu, E.; Huang, Q.; Lai, M. Analysis of SOX9 expression in colorectal cancer. Am. J. Clin. Pathol. 2008, 130, 897–904. [Google Scholar] [CrossRef] [Green Version]
- Oliemuller, E.; Kogata, N.; Bland, P.; Kriplani, D.; Daley, F.; Haider, S.; Shah, V.; Sawyer, E.J.; Howard, B.A. SOX11 promotes invasive growth and ductal carcinoma in situ progression. J. Pathol. 2017, 243, 193–207. [Google Scholar] [CrossRef]
- Shepherd, J.H.; Uray, I.P.; Mazumdar, A.; Tsimelzon, A.; Savage, M.; Hilsenbeck, S.G.; Brown, P.H. The SOX11 transcription factor is a critical regulator of basal-like breast cancer growth, invasion, and basal-like gene expression. Oncotarget 2016, 7, 13106. [Google Scholar] [CrossRef]
- Ding, H.; Quan, H.; Yan, W.; Han, J. Silencing of SOX12 by shRNA suppresses migration, invasion and proliferation of breast cancer cells. Biosci. Rep. 2016, 36, e00389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.; Li, Z.; Zhu, J. Negative regulation of SOX11 in hepatocellular carcinoma. Int J. Clin. Exp. Med. 2017, 10, 2809–2817. [Google Scholar]
- Huang, W.; Chen, Z.; Shang, X.; Tian, D.; Wang, D.; Wu, K.; Fan, D.; Xia, L. Sox12, a direct target of FoxQ1, promotes hepatocellular carcinoma metastasis through up-regulating Twist1 and FGFBP1. Hepatology 2015, 61, 1920–1933. [Google Scholar] [CrossRef]
- Walter, R.F.H.; Mairinger, F.D.; Werner, R.; Ting, S.; Vollbrecht, C.; Theegarten, D.; Christoph, D.C.; Zarogoulidis, K.; Schmid, K.W.; Zarogoulidis, P. SOX4, SOX11 and PAX6 mRNA expression was identified as a (prognostic) marker for the aggressiveness of neuroendocrine tumors of the lung by using next-generation expression analysis (NanoString). Future Oncol. 2015, 11, 1027–1036. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Hu, F.; Shen, S.; Xiao, H.; Li, G.; Wang, M.; Mei, J. Knockdown of SOX12 expression inhibits the proliferation and metastasis of lung cancer cells. Am. J. Transl. Res. 2017, 9, 4003. [Google Scholar]
- Thu, K.; Radulovich, N.; Becker-Santos, D.; Pikor, L.; Pusic, A.; Lockwood, W.; Lam, W.; Tsao, M. SOX15 is a candidate tumor suppressor in pancreatic cancer with a potential role in Wnt/β-catenin signaling. Oncogene 2014, 33, 279. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.; Jiang, S.; Lu, C.; Ji, T.; Yang, W.; Li, T.; Lv, J.; Hu, W.; Yang, Y.; Jin, Z. SOX11: Friend or foe in tumor prevention and carcinogenesis? Ther. Adv. Med Oncol. 2019, 11, 1758835919853449. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Ji, E.H.; Zhao, X.; Cui, L.; Misuno, K.; Guo, M.; Huang, Z.; Chen, X.; Hu, S. Sox11 promotes head and neck cancer progression via the regulation of SDCCAG8. J. Exp. Clin. Cancer Res. 2019, 38, 138. [Google Scholar] [CrossRef]
- Du, F.; Chen, J.; Liu, H.; Cai, Y.; Cao, T.; Han, W.; Yi, X.; Qian, M.; Tian, D.; Nie, Y.; et al. SOX12 promotes colorectal cancer cell proliferation and metastasis by regulating asparagine synthesis. Cell Death Dis. 2019, 10, 1–19. [Google Scholar] [CrossRef]
- Du, F.; Feng, W.; Chen, S.; Wu, S.; Cao, T.; Yuan, T.; Tian, D.; Nie, Y.; Yu, K.; Fan, D.; et al. Sex determining region Y-box 12 (SOX12) promotes gastric cancer metastasis by upregulating MMP7 and IGF1. Cancer Lett. 2019, 452, 103–118. [Google Scholar] [CrossRef] [PubMed]
- Guth, S.; Wegner, M. Having it both ways: Sox protein function between conservation and innovation. Cell. Mol. Life Sci. 2008, 65, 3000–3018. [Google Scholar] [CrossRef]
- Dy, P.; Penzo-Mendez, A.; Wang, H.; Pedraza, C.E.; Macklin, W.B.; Lefebvre, V. The three SoxC proteins—Sox4, Sox11 and Sox12—exhibit overlapping expression patterns and molecular properties. Nucleic Acids Res. 2008, 36, 3101–3117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grimm, D.; Bauer, J.; Wise, P.; Krüger, M.; Simonsen, U.; Wehland, M.; Infanger, M.; Corydon, T.J. The Role of SOX Family Members in Solid Tumours and Metastasis. In Seminars in Cancer Biology; Elsevier: Amsterdam, The Netherlands, 2019. [Google Scholar]
- Hargrave, M.; Wright, E.; Kun, J.; Emery, J.; Cooper, L.; Koopman, P. Expression of the Sox11 gene in mouse embryos suggests roles in neuronal maturation and epithelio-mesenchymal induction. Dev. Dyn. 1997, 210, 79–86. [Google Scholar] [CrossRef]
- Sock, E.; Rettig, S.D.; Enderich, J.; Bösl, M.R.; Tamm, E.R.; Wegner, M. Gene targeting reveals a widespread role for the high-mobility-group transcription factor Sox11 in tissue remodeling. Mol. Cell. Biol. 2004, 24, 6635–6644. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoser, M.; Potzner, M.R.; Koch, J.M.; Bösl, M.R.; Wegner, M.; Sock, E. Sox12 deletion in the mouse reveals nonreciprocal redundancy with the related Sox4 and Sox11 transcription factors. Mol. Cell. Biol. 2008, 28, 4675–4687. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, C.; Wilhelm, D.; Koopman, P. Sox genes and cancer. Cytogenet. Genome Res. 2004, 105, 442–447. [Google Scholar] [CrossRef]
- Leber, M.F.; Efferth, T. Molecular principles of cancer invasion and metastasis. Int. J. Oncol. 2009, 34, 881–895. [Google Scholar]
- Foulkes, W.D.; Flanders, T.Y.; Pollock, P.M.; Hayward, N.K. The CDKN2A (p16) gene and human cancer. Mol. Med. 1997, 3, 5–20. [Google Scholar] [CrossRef] [Green Version]
- Yuan, X.; Larsson, C.; Xu, D. Mechanisms underlying the activation of TERT transcription and telomerase activity in human cancer: Old actors and new players. Oncogene 2019, 38, 6172–6183. [Google Scholar] [CrossRef] [Green Version]
- Jančík, S.; Drábek, J.; Radzioch, D.; Hajdúch, M. Clinical relevance of KRAS in human cancers. J. Biomed. Biotechnol. 2010, 2010, 150960. [Google Scholar] [CrossRef] [Green Version]
- Soussi, T.; Wiman, K. TP53: An oncogene in disguise. Cell Death Differ. 2015, 22, 1239. [Google Scholar] [CrossRef]
- Porru, M.; Pompili, L.; Caruso, C.; Biroccio, A.; Leonetti, C. Targeting KRAS in metastatic colorectal cancer: Current strategies and emerging opportunities. J. Exp. Clin. Cancer Res. 2018, 37, 57. [Google Scholar] [CrossRef] [Green Version]
- Waters, A.M.; Der, C.J. KRAS: The critical driver and therapeutic target for pancreatic cancer. Cold Spring Harb. Perspect. Med. 2018, 8, a031435. [Google Scholar] [CrossRef] [PubMed]
- Samuels, Y.; Velculescu, V.E. Oncogenic mutations of PIK3CA in human cancers. Cell Cycle 2004, 3, 1221–1224. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shlien, A.; Malkin, D. Copy number variations and cancer. Genome Med. 2009, 1, 62. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mehta, G.A.; Parker, J.S.; Silva, G.O.; Hoadley, K.A.; Perou, C.M.; Gatza, M.L. Amplification of SOX4 promotes PI3K/Akt signaling in human breast cancer. Breast Cancer Res. Treat. 2017, 162, 439–450. [Google Scholar] [CrossRef]
- Medina, P.P.; Castillo, S.D.; Blanco, S.; Sanz-Garcia, M.; Largo, C.; Alvarez, S.; Yokota, J.; Gonzalez-Neira, A.; Benitez, J.; Clevers, H.C. The SRY-HMG box gene, SOX4, is a target of gene amplification at chromosome 6p in lung cancer. Hum. Mol. Genet. 2009, 18, 1343–1352. [Google Scholar] [CrossRef] [PubMed]
- Vervoort, S.J.; van Boxtel, R.; Coffer, P.J. The role of SRY-related HMG box transcription factor 4 (SOX4) in tumorigenesis and metastasis: Friend or foe? Oncogene 2013, 32, 3397–3409. [Google Scholar] [CrossRef] [Green Version]
- Rhodes, D.R.; Yu, J.; Shanker, K.; Deshpande, N.; Varambally, R.; Ghosh, D.; Barrette, T.; Pandey, A.; Chinnaiyan, A.M. ONCOMINE: A cancer microarray database and integrated data-mining platform. Neoplasia 2004, 6, 1–6. [Google Scholar] [CrossRef] [Green Version]
- Rhodes, D.R.; Kalyana-Sundaram, S.; Mahavisno, V.; Varambally, R.; Yu, J.; Briggs, B.B.; Barrette, T.R.; Anstet, M.J.; Kincead-Beal, C.; Kulkarni, P.; et al. Oncomine 3.0: Genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia 2007, 9, 166–180. [Google Scholar] [CrossRef] [Green Version]
- Tang, Z.; Li, C.; Kang, B.; Gao, G.; Li, C.; Zhang, Z. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017, 45, W98–W102. [Google Scholar] [CrossRef] [Green Version]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef] [Green Version]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nagy, A.; Lanczky, A.; Menyhart, O.; Gyorffy, B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci. Rep. 2018, 8, 9227. [Google Scholar] [CrossRef] [PubMed]
- Jassal, B.; Matthews, L.; Viteri, G.; Gong, C.; Lorente, P.; Fabregat, A.; Sidiropoulos, K.; Cook, J.; Gillespie, M.; Haw, R. The reactome pathway knowledgebase. Nucleic Acids Res. 2020, 48, D498–D503. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [Green Version]
- Chandrashekar, D.S.; Bashel, B.; Balasubramanya, S.A.H.; Creighton, C.J.; Ponce-Rodriguez, I.; Chakravarthi, B.V.; Varambally, S. UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 2017, 19, 649–658. [Google Scholar] [CrossRef]
- Warde-Farley, D.; Donaldson, S.L.; Comes, O.; Zuberi, K.; Badrawi, R.; Chao, P.; Franz, M.; Grouios, C.; Kazi, F.; Lopes, C.T. The GeneMANIA prediction server: Biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010, 38, W214–W220. [Google Scholar] [CrossRef]










| #Term ID | Term Description | False Discovery Rate |
|---|---|---|
| hsa05213 | Endometrial cancer | 2.20 × 10−20 |
| hsa05215 | Prostate cancer | 4.75 × 10−18 |
| hsa05200 | Pathways in cancer | 2.24 × 10−17 |
| hsa04310 | Wnt signaling pathway | 2.61 × 10−16 |
| hsa05226 | Gastric cancer | 2.92 × 10−16 |
| hsa04520 | Adherens junction | 3.34 × 10−16 |
| hsa05225 | Hepatocellular carcinoma | 7.38 × 10−16 |
| hsa05165 | Human papillomavirus infection | 2.76 × 10−15 |
| hsa05217 | Basal cell carcinoma | 5.93 × 10−15 |
| hsa05206 | MicroRNAs in cancer | 8.70 × 10−15 |
| hsa04390 | Hippo signaling pathway | 9.90 × 10−15 |
| hsa05210 | Colorectal cancer | 6.99 × 10−14 |
| hsa05224 | Breast cancer | 2.64 × 10−13 |
| hsa04934 | Cushing’s syndrome | 3.71 × 10−13 |
| hsa05216 | Thyroid cancer | 4.14 × 10−13 |
| hsa04115 | p53 signaling pathway | 4.94 × 10−13 |
| hsa04110 | Cell cycle | 1.55 × 10−12 |
| hsa05166 | HTLV-I infection | 4.71 × 10−11 |
| hsa04916 | Melanogenesis | 4.15 × 10−10 |
| hsa04218 | Cellular senescence | 4.27 × 10−10 |
| #Term ID | Term Description | False Discovery Rate |
|---|---|---|
| HSA-201681 | TCF-dependent signaling in response to WNT | 4.69 × 10−15 |
| HSA-3769402 | Deactivation of the β-catenin transactivating complex | 4.69 × 10−15 |
| HSA-8878159 | Transcriptional regulation by RUNX3 | 1.63 × 10−12 |
| HSA-212436 | Generic Transcription Pathway | 3.64 × 10−12 |
| HSA-2990846 | SUMOylation | 5.69 × 10−12 |
| HSA-74160 | Gene expression (Transcription) | 9.40 × 10−12 |
| HSA-195253 | Degradation of β-catenin by the destruction complex | 1.42 × 10−11 |
| HSA-162582 | Signal Transduction | 3.63 × 10−11 |
| HSA-1640170 | Cell Cycle | 4.88 × 10−11 |
| HSA-201722 | Formation of the β-catenin/TCF transactivating complex | 7.06 × 10−11 |
| HSA-6804760 | Regulation of TP53 activity through methylation | 4.81 × 10−10 |
| HSA-3065678 | SUMO is transferred from E1 to E2 (UBE2I, UBC9) | 6.14 × 10−10 |
| HSA-4411364 | Binding of TCF/LEF: CTNNB1 to target gene promoters | 1.59 × 10−9 |
| HSA-8951430 | RUNX3 regulates WNT signaling | 1.59 × 10−9 |
| HSA-3108232 | SUMO E3 ligases SUMOylate target proteins | 1.86 × 10−9 |
| HSA-69563 | p53-dependent G1 DNA Damage Response | 3.53 × 10−9 |
| HSA-1643685 | Disease | 4.70 × 10−9 |
| HSA-3700989 | Transcriptional Regulation by TP53 | 6.39 × 10−9 |
| HSA-5663202 | Diseases of signal transduction | 6.39 × 10−9 |
| HSA-6804757 | Regulation of TP53 Degradation | 6.39 × 10−9 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Seok, J.; Gil, M.; Dayem, A.A.; Saha, S.K.; Cho, S.-G. Multi-Omics Analysis of SOX4, SOX11, and SOX12 Expression and the Associated Pathways in Human Cancers. J. Pers. Med. 2021, 11, 823. https://doi.org/10.3390/jpm11080823
Seok J, Gil M, Dayem AA, Saha SK, Cho S-G. Multi-Omics Analysis of SOX4, SOX11, and SOX12 Expression and the Associated Pathways in Human Cancers. Journal of Personalized Medicine. 2021; 11(8):823. https://doi.org/10.3390/jpm11080823
Chicago/Turabian StyleSeok, Jaekwon, Minchan Gil, Ahmed Abdal Dayem, Subbroto Kumar Saha, and Ssang-Goo Cho. 2021. "Multi-Omics Analysis of SOX4, SOX11, and SOX12 Expression and the Associated Pathways in Human Cancers" Journal of Personalized Medicine 11, no. 8: 823. https://doi.org/10.3390/jpm11080823