1. Introduction
Mitogen-activated protein kinase kinase 4 (
MKK4) is an important regulatory kinase of the JNK signaling pathway. Activation of the JNK signaling pathway is achieved by dual phosphorylation of Thr and Tyr residues of the conserved Thr–Pro–Tyr (T–Y–P) motif [
1,
2].
MKK4 can alternately activate the two pathways of JNK and p38 [
3]. The first successful cloning of the
MKK4 gene was in
Xenopus laevis, and since then, its role and molecular mechanism functions in both physiological and biochemical areas have been studied more and more deeply. However, the leading research focuses on mammals, such as humans and mice [
4,
5]. In studies on crustaceans, it was only found in
Daphnia pulex,
Fenneropenaeus chinensis,
Litopenaeus vannamei, and
P. monodon [
6,
7,
8]. In a prior study of
P. monodon, the expression of
MKK4 was only studied in spermatozoa and ovary developmental stages. The
MKK4 gene of
L. vannamei shows significant changes under the stimulation of various bacteria and is activated by the phosphorylation of upstream genes and then activated JNK, which plays an important role in the antibacterial response [
3,
9]. The
MKK4 gene of
Ctenopharyngodon idella is involved in the immune stress response to muramyl dipeptide in the intestinal tissue of
C. idella [
10].
In order to further improve the molecular function study of the MKK4 gene in P. monodon, this study cloned the entire length of the MKK4 gene in P. monodon, analyzed the expression of MKK4 in various tissues of P. monodon, and explored the effect of different degrees of pathogen stimulation and salinity stress. It is necessary to explore the molecular function and role of MKK4 in low-salinity stress in P. monodon to provide more basic data to determine the molecular mechanism of the response to salinity stress in P. monodon.
2. Materials and Methods
2.1. Experimental Animals and PmMKK4 cDNA Cloning
P. monodon specimens, each with a body length of 7–10 cm and a body weight of 8–12 g, were selected and cultured for 1 week in a 25–28 °C seawater environment with full aeration at the same time. Based on the unpublished transcriptome of
P. monodon tissues constructed in our laboratory, specific primers, namely,
PmMKK4-F1,
PmPKK4-R1,
PmMKK4-F2, and
PmPKK4-R2 (
Table 1), were designed by Primer Premier 5.0 (RuiBiotech, Guangzhou, China). The 5′ and 3′ ends of
PmMKK4 were acquired by using the rapid amplification of the cDNA end (RACE) method (Clonetech, Tokyo, Japan). Through the 5′ or 3′ RACE-PCR, PCR was performed initially with
PmMKK4-5G1 (
PmMKK4-3G1 for 3′) and universal primers UPM Long and UPM Short, followed by semi-nested PCR with
PmMKK4-5G2 (
PmMKK4-3G2 for 3′) and a nested universal primer NUP (
Table 1). The PCR conditions were designed as follows: one cycle of 94 °C for 3 min, 35 cycles of 94 °C for 30 s, 67 °C for 30 s, and 72 °C for 45 s, followed by a final cycle of 72 °C for 10 min. The PCR products were then gel-purified and sequenced, and the sequences were determined after the analysis.
2.2. Bioinformatic Analysis
2.3. Sample Collection and cDNA Synthesis
After 1 week of seawater culturing, healthy male and female P. monodon were randomly selected. Hepatopancreas, gills, intestines, stomach, lymph, heart, muscle, epidermis, eye stalk nerve, cranial nerve, thoracic nerve, ventral nerve, and ovary (testis) tissue were dissected for collection. The same tissue samples from 3 shrimps were mixed into one tube. Furthermore, during summer, the mating season of P. monodon, samples of shrimp larvae at different stages of development were collected. Those samples including zygote, nauplius, zoea, mysis, and postlarval stages. Zygote stage samples were collected immediately after egg laying. When 80% of the population had reached the objective stage according to their morphologies, as observed using an optical microscope, samples of the larvae of different stages were collected. The samples mentioned above were organized into three parallel groups, stored in RNAlater® RNA stabilization solution (Invitrogen, Carlsbad, CA, USA) at 4 °C overnight, and then stored at −80 °C.
Following the manufacturer’s instructions of Trizol reagent (Invitrogen, USA), the total RNA of all collected samples was extracted. The ratios of ultraviolet absorbance at 260/280 nm were measured using a NanoDrop2000 device (NanoDrop Technologies, Waltham, MA, USA); 1.5% agarose gel electrophoresis was used to ensure the integrity. A template of cDNA was synthesized from the RNA using the PrimeScript II 1st strand cDNA synthesis kit (Takara, Tokyo, Japan). For the sake of real-time quantitative PCR (qRT-PCR), cDNA was synthesized in accordance with the manufacturer’s instructions for the PrimeScript TM RT reagent kit with a gDNA eraser (Perfect Real-time, Takara, Japan) and diluted to 50 ng/μL for use as the template.
2.4. Bacterial Infection Challenge
In total, 200 healthy
P. monodon with an average weight of 15–18 g were selected for immune challenge experiments. Three pathogenic bacteria provided by the Key Laboratory of South China Sea Fishery Resources Exploitation and Utilization were used. According to a previous study [
11], the culture schedule and injection concentration for each strain were determined. Four experimental groups were included: PBS (shrimp specimen injected with sterile phosphate-buffered saline as control),
Staphylococcus aureus,
Vibrio harveyi, and
Vibrio anguillarum groups. Each group of 50 shrimp specimens was injected into the second abdominal segment with 100 μL of sterile phosphate-buffered saline (PBS, pH 7.4) or 100 μL (1.0 × 10
8 cfu/mL) of
S. aureus,
Vibrio harveyi, and
V. anguillarum, respectively. Healthy and intact shrimp were randomly selected at 0 h, 3 h, 6 h, 12 h, 24 h, 48 h, and 72 h after injection for the dissection of hepatopancreas and gill tissues. Hepatopancreas and gill tissues were stored overnight in RNAlater solution at 4 °C and then kept at −80 °C.
2.5. Low-Salinity Stress
The experiment site of the low-salt stress test was in the South China Sea Fisheries Research Institute (Shenzhen City, Guangdong Province, China). A total of 360 shrimps (7–10 cm) was selected for these experiments. The salinity concentration was adjusted to the target salinity by mixing the cultured seawater with freshwater using a salinity meter (AZ8371, Hengxin, Taiwan). According to the pre-test results, the 96 h half-lethal salinity was 3 psu [
12]. Therefore, the stress salinity was set to 3 psu. Another experimental group with salinity of 17 psu was set up, and conventional aquaculture seawater (about 25 psu) was used as the control group [
13]. Three parallel groups (
n = 40/group) were created. The incubation temperature and pH were maintained at 25–28 °C and 7.0 ± 0.5, respectively, and 0 h, 3 h, 6 h, 12 h, 24 h, 48 h, 72 h, and 96 h after exposure to different salinity stresses, shrimp specimens with optimal activity during the intermolt were selected for dissection to collect gills and hepatopancreas tissues, and they were preserved in RNAlater solution. The tissues were then maintained at −80 °C after overnight storage at 4 °C.
2.6. qRT-PCR Analysis of PmMKK4 mRNA Expression
In this study, qRT-PCR was used to detect
PmMKK4 mRNA expression in different tissues at different developmental stages following bacterial challenge and low-salinity stress exposure. Since the reaction component and cycle condition for EF1a are consistent with
PmMKK, the reference gene was chosen to be 1α (EF1a) (
Table 1). The solution in each hole (12.5 μL) was a mixture of 6.25 μL of 2 × TB GreenTM Premix ExTaq (Takara, Beijing, China), 0.5 μL each of
PmMKK4-qF and
PmMKK4-qR (50 μmol/L), 1 μL qRT-PCR diluted cDNA, and 4.25 μL double-distilled water. Green fluorescence measurement qRT-PCR was carried out in the quantitative real-time PCR system, Roche Light Cycler
® 480II. The following four steps were conducted: degeneration for 30 s at 95 °C, a quantitative analysis stage with 40 cycles of 94 °C for 5 s and 60 °C for 30 s, dissolution curve analysis for 5 s at 95 °C and 60 °C and up to 95 °C for 1 min, and an according stage of 30 s at 50 °C. The relative CT method (2
−ΔΔCT) was used to obtain the PCR data. One-way ANOVA was used to work on statistical analysis. SPSS statistics version 23.0 software (IBM, Armonk, New York, USA) was used to carry out the Tukey’s multiple range test. The differences were considered to be significant at
p < 0.05. Tested data were presented as mean ± SD (standard deviation).
2.7. Low-Salinity Stress Testing in P. monodon following RNA Interference (RNAi)-Mediated Knockdown of PmMKK4
Primer Premier 5.0 was used to design the following primers: dsMKK-f, dsMKK-r, dsMKK-T7-f, and dsMKK-T7-r (
Table 1). A DNA fragment containing the T7 promoter was amplified by Ex Taq using normal cDNA as a template. PCR conditions were as follows: 3 min at 94 °C, 35 cycles of 30 s at 94 °C, 30 s at 58 °C, 1 min at 72 °C, and a final cycle of 10 min at 72 °C. Excess bands were clipped after agarose gel electrophoresis, leaving clear and bright bands. DNA fragments were recovered according to the gel recovery kit instructions. dsRNA synthesis was performed according to the T7 RiboMAXTMExpress RNAi System kit instructions. Reaction system: RiboMAXTM Express T7 2X Buffer 10.0 μL, linear DNA template (total 1 μg) 1.0–8.0 μL, Nuclease-Free Water 0–7.0 μL (8 μL-DNA volume), Enzyme Mix-T7 Express 2.0 μL. A final volume of 20.0 μL was incubated at 37 °C for 30 min to obtain single-stranded RNA (ssRNA). Then, equal volumes of complementary ssRNA were mixed and incubated at 70 °C for 10 min, 65 °C for 10 min, and 25 °C for 10 min. Then, 2.0 μL of freshly diluted RNase solution (1 μL RNase solution: 199 μL Nuclease-Free Water) and 2.0 μL RQ1 RNase-Free DNase was added and incubated at 37 °C for 30 min. The obtained dsRNA was purified for later use. dsGFP-F/R primers and pD-GFP recombinant vectors were used to synthesize green fluorescent protein (GFP) double-stranded RNA in the same way.
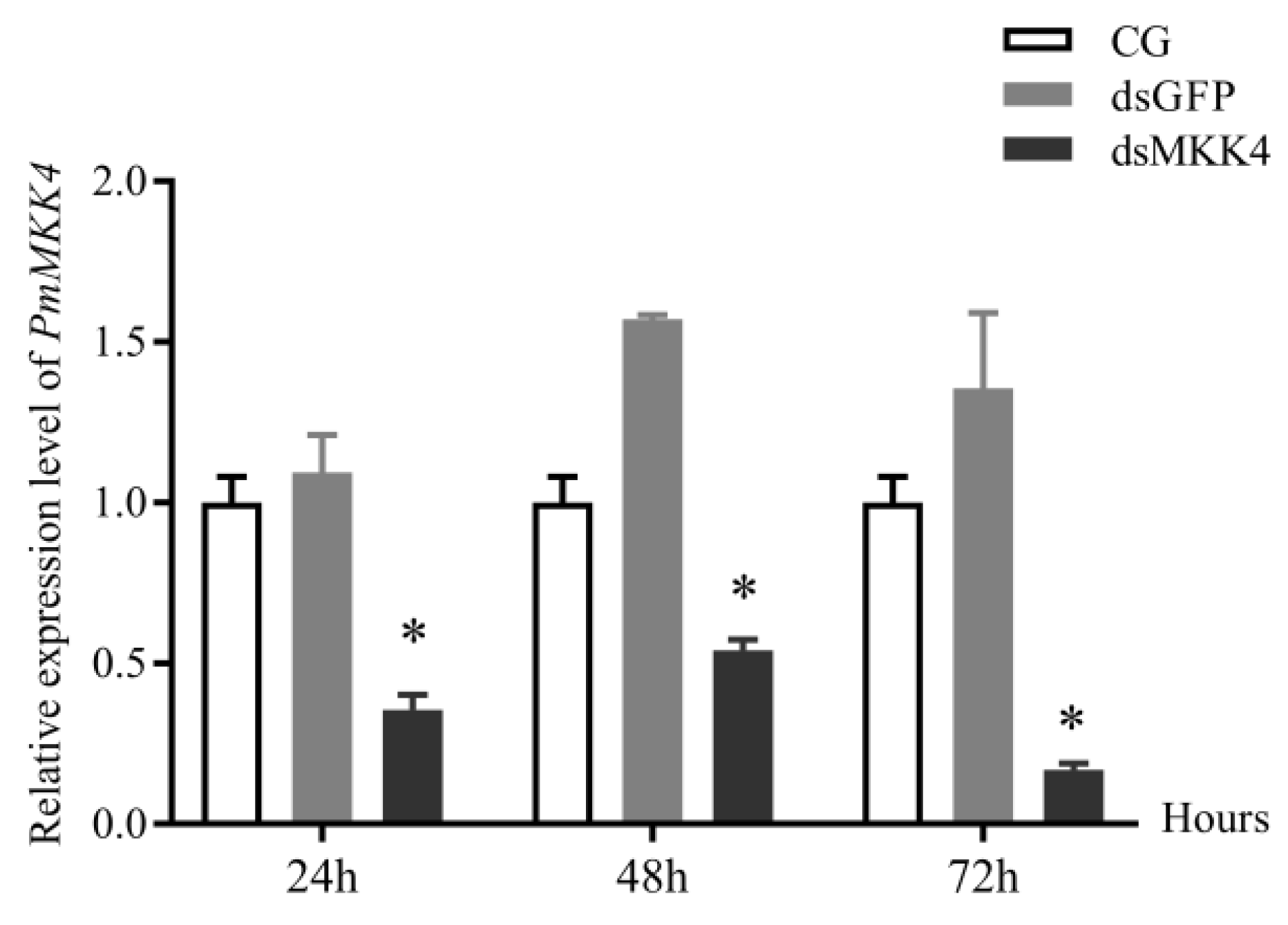
The weight of P. monodon was 5.0 ± 1.0 g, and the dsRNA injection volume was 3–5 μg/g shrimp. Each shrimp was injected intramuscularly in the second abdominal segment. The injection experiments were divided into three groups: PBS group, dsGFP group, and dsMKK group. Before injection, samples of healthy shrimp in the intermolt phase were randomly collected, and their gill tissues were placed in RNAlater as a 0 h sample to check the RNAi efficiency. Twenty-four hours after injection, the three groups of shrimp specimens were transferred to plastic buckets adjusted to salinity of 3 psu. The dead shrimps of each group were collected and recorded every 3 h.
After 3 h, 6 h, 9 h, 12 h, 24 h, 48 h, 72 h, and 96 h exposure to salt stress, healthy shrimp specimens were collected from each treatment group, and their gill tissues were dissected and placed in RNAlater. Furthermore, 10 shrimp injected with dsGFP and dsMKK, respectively, were placed in two separate plastic buckets filled with seawater (about 25 psu). After 24 and 48 h, gill tissues were randomly collected and placed in RNAlater as samples to measure dsRNA interference efficiency. All RNA samples were mixed in RNAlater and stored at 4 °C overnight, then kept at −80 °C.
4. Discussion
The
MKK4 gene of
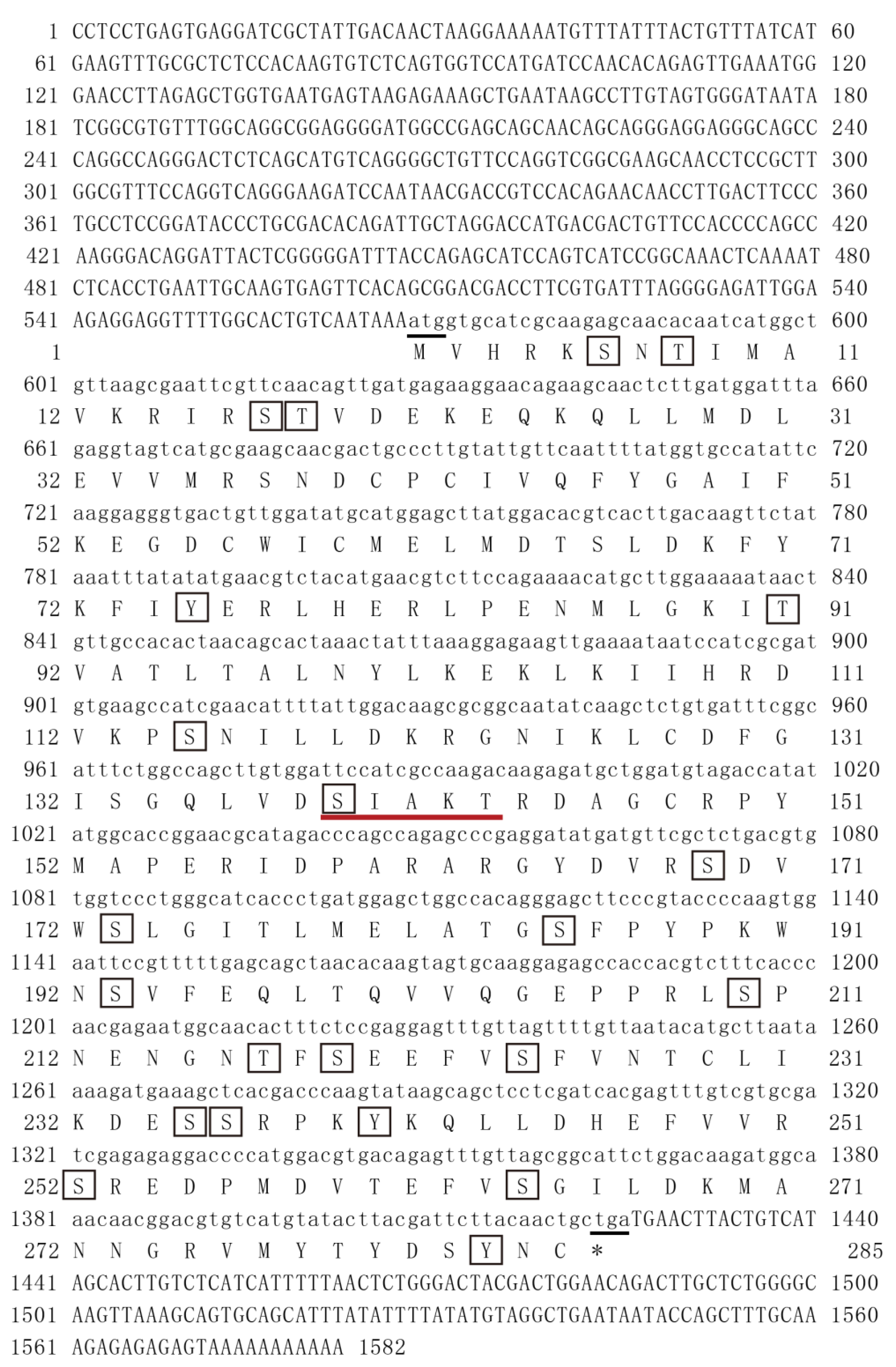
P. monodon was cloned in this study. The results of amino acid sequence analysis showed that
PmMKK4 has 22 phosphorylation sites. The phosphate groups can regulate different functions of the protein and may play a synergistic or reverse role in various reactions [
14]. Structural prediction indicated that
PmMKK4 contains a conserved serine/threonine protein kinase (S–T–T–K–C) region, which is a potential double phosphorylation site. The
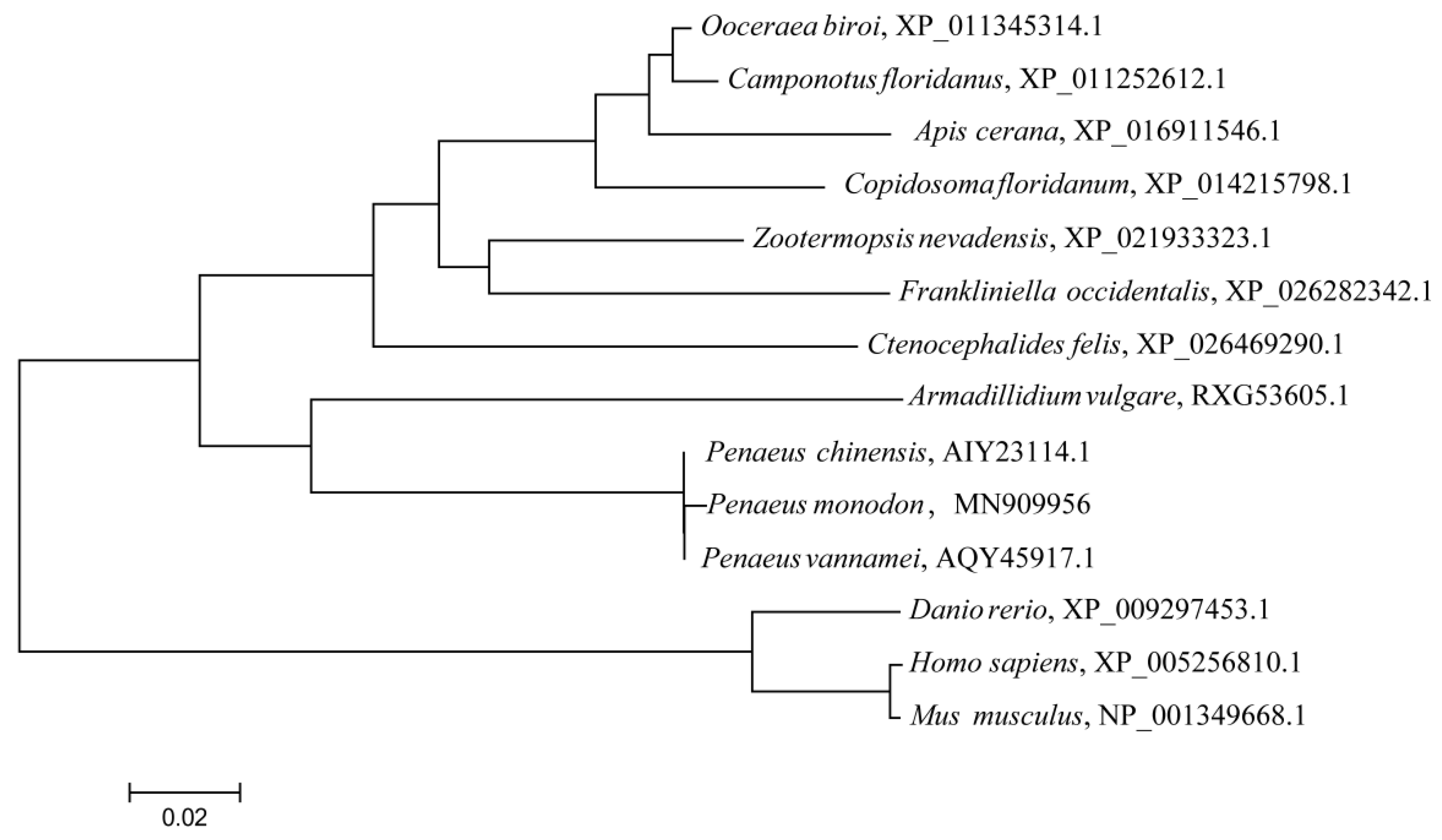
MKK4 gene has two downstream pathways: the JNK signaling pathway, and the p38 signaling pathway. These two branch pathways play important roles in immunity and anti-stress processes in organisms. Phylogenetic tree analysis showed that the
MKK4 gene of
P. monodon was most closely related to
P. vannamei and
P. chinensis, clustered into a clade. The results of multiple sequence alignment showed that
PmMKK4 has a high similarity with
MKK4 genes of other species, among which the similarity with
P. vannamei and
P. chinensis was the highest (99.65%), indicating that
MKK4 genes were relatively similar among closely related species. It is speculated that its conserved protein kinase domain may play an important role in many aspects such as physiology and biochemistry.
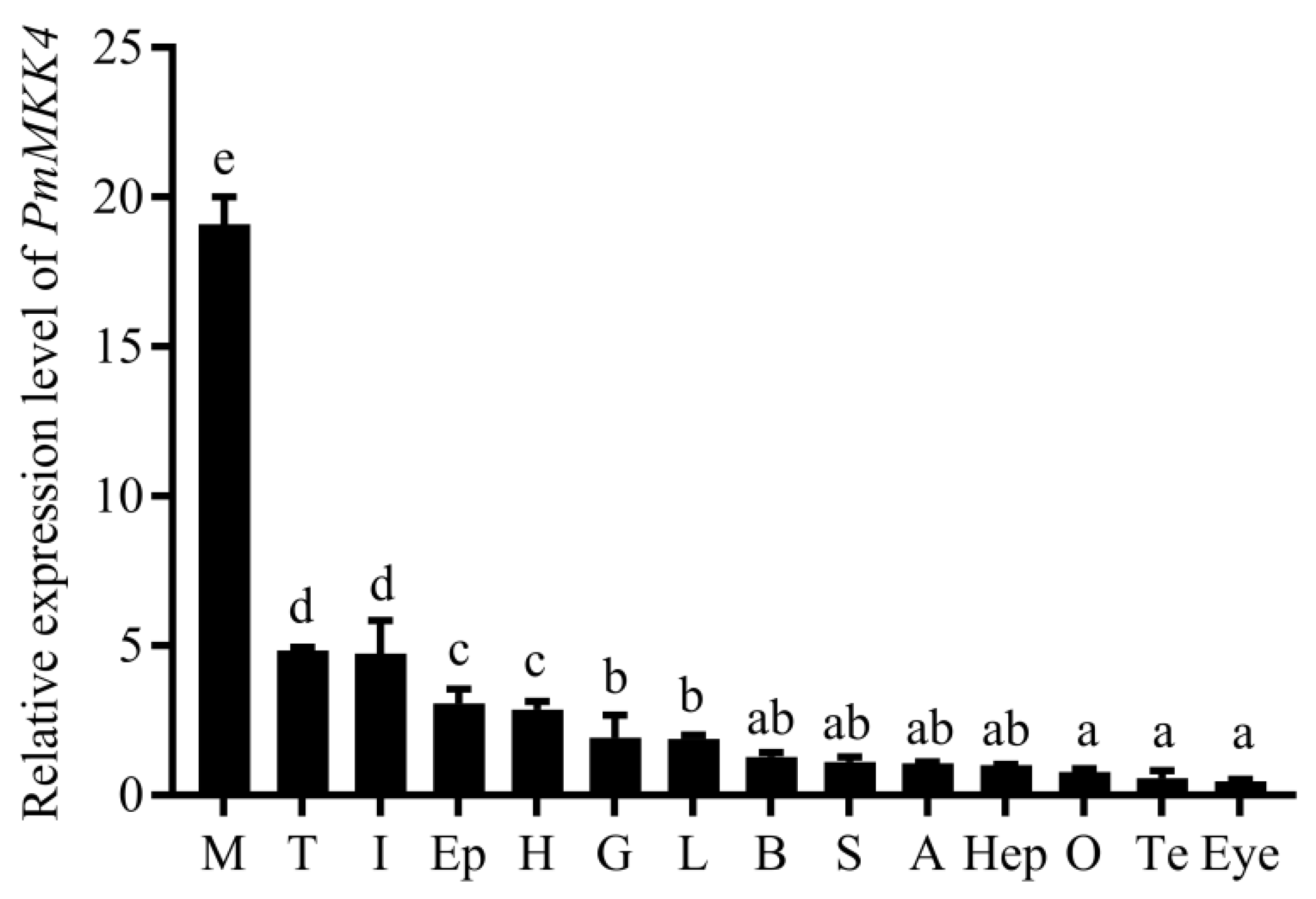
Through quantitative tissue analysis, it was determined that
PmMKK4 was expressed in all tested tissues, which is the same as the tissue distribution results of other species. In the detected tissues, the expression level of Pm
MKK4 in muscle was the highest, which was significantly higher than other tissues. Similarly, in the study of
P. chinensis, the expression level of
MKK4 in muscle tissue was also significantly higher than that in other tissues [
7], suggesting that muscle may be an important tissue for the function of the
MKK4 gene. In addition to muscle,
PmMKK4 was also highly expressed in tissues such as intestine, heart, gill, and lymph. Intestine and lymph were important immune tissues, and gills were important tissues for crustaceans to exchange ions with the environment and regulate osmotic pressure. Therefore, it was speculated that
PmMKK4 plays an important role in the immune and salinity stress responses of
P. monodon.
In order to explore the role of the
PmMKK4 gene in the innate immunity of
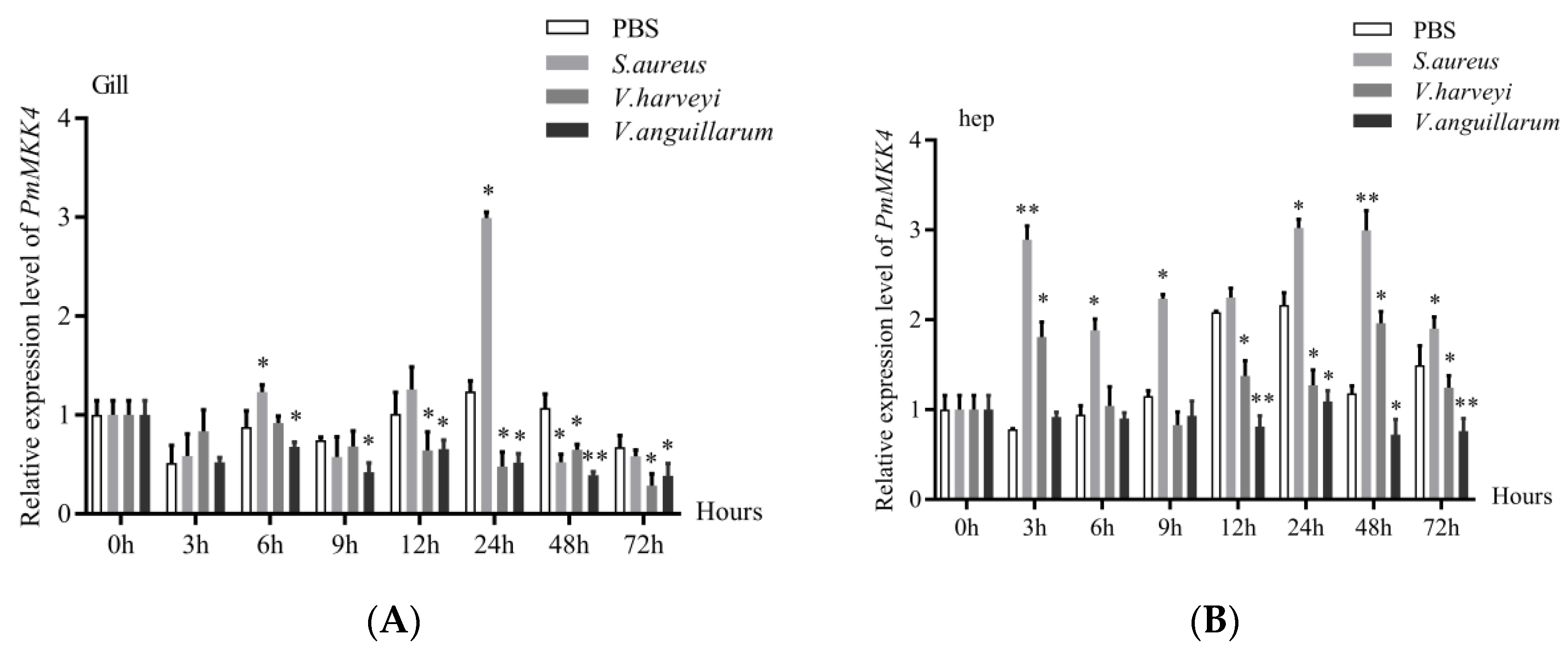
P. monodon, we carried out pathogen infection experiments. Although the expression of
PmMKK4 was the highest in muscle, considering that it had been widely accepted that gill and hepatopancreas were important immune organs in previous studies, while muscle and nerve were rarely considered immune-related tissues, we chose gill and hepatopancreas. Tissue responses to bacterial stimulation were studied. The experimental results showed that
PmMKK4 was up-regulated in hepatopancreas and gill tissues after infection with
S. aureus, which is similar to the
MKK4 gene in
L. vannamei in response to
S. aureus [
3]. After infection with
Vibrio harveyi,
PmMKK4 was down-regulated for a period of time in gill tissue, while fluctuating between up- and down-regulation in hepatopancreas tissue. After
P. monodon was infected with
Vibrio harveyi, the expression levels of three related genes in the JNK pathway were significantly reduced, indicating that some effector molecules of the JNK signaling pathway may be involved in the process of immune regulation, which is consistent with the research results of Shi et al. [
15]. After infection with
V. anguillarum, gill tissue and hepatopancreas tissue showed a continuous downward trend. However, the
MKK4 gene was significantly up-regulated after
L. vannamei infection with the Gram-negative bacteria
Vibrio parahaemolyticus [
3]. The physiological role of the post-
MKK4 gene is different, and therefore the response mode is different. In addition, when
Pinctada fucata are infested by exogenous pathogens, the expression level of
MKK4 gene is significantly changed and phosphorylated, suggesting that it is involved in the self-protection mechanism of
Pinctada fucata to defend against the occurrence of diseases [
16,
17].
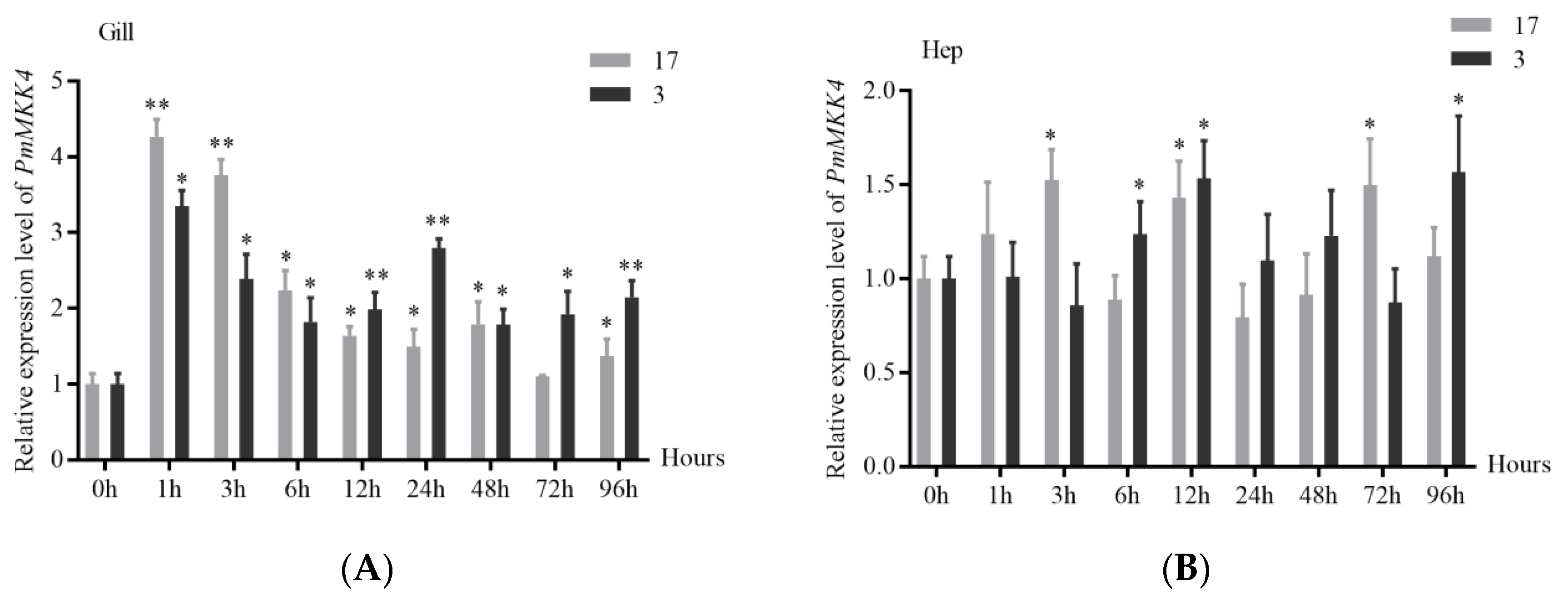
Changes in salinity can cause changes in the osmotic pressure and the activities of various non-specific immune enzymes in the shrimp, affecting the immune defense ability of the shrimp. The experimental results of acute low-salt stress in this study showed that the
PmMKK4 gene in the gill tissue and hepatopancreas tissue of
P. monodon was activated after stress, resulting in different degrees of up-regulation. Among them, the expression change of
PmMKK4 in gill tissue was more significant. It can be speculated that under low salt stress, the gill tissue undertakes more physiological and biochemical reactions related to the
MKK4 gene. In the process of ammonia nitrogen stress in
P. chinensis, the expression of the
MKK4 gene in muscle, hepatopancreas, gill, and other tissues is significantly increased, suggesting that the
MKK4 gene is involved in the stress resistance process of
P. chinensis.
MKK4 can activate downstream JNK signaling pathway. Studies have confirmed that the JNK branch pathway plays an important role in the salinity adaptation process of aquatic animals. Under salinity stress, the expression of
MAPK 8 (JNK1) and
MAPK 9 (JNK 2) was significantly up-regulated [
18,
19,
20]. To further explore the role of the
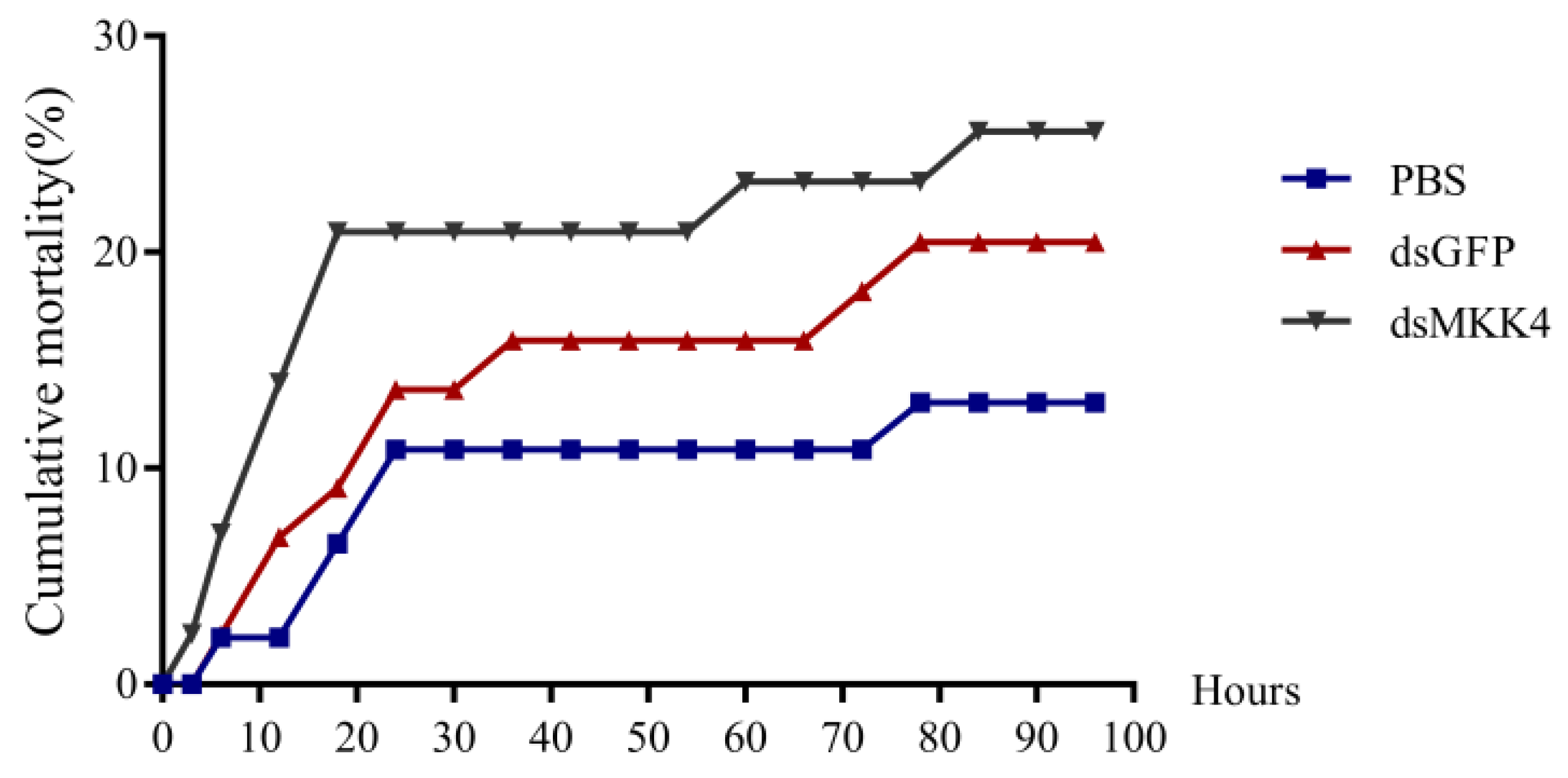
PmMKK4 gene in low-salt stress, we performed double-stranded RNA injection experiments. In a low-salt environment, the mortality rate after knocking down the
PmMKK4 gene increased rapidly in a short period of time, while the mortality rates of the other two groups were lower. It is speculated that the low expression of the
MKK4 gene reduces the adaptability of
P. monodon to the low-salt environment and causes more deaths. The
PmMKK4 gene may play an important role in adaptation to the low-salt environment. After the interference, we further conducted a quantitative study on other genes in the JNK signaling pathway. The results showed that the expression of the
MKK7 gene in
P. monodon showed a significant upward trend as a whole, while the expression of
PmJNK fluctuated in the early stage and stably increased in the later stage. It is speculated that the
JNK signaling pathway is further activated under low-salt stress, and the
PmMKK7 gene maintains high expression, while the
PmMKK4 gene is expressed less to ensure the stable functioning of the JNK signaling pathway.