Abstract
Rose (Rosa chinensis Jacq.) is an important economic ornamental crop and its yield is affected by different biotic and abiotic stresses. Among the biotic stresses, the gray mold disease caused by Botrytis cinerea is a serious threat to rose production. The basic helix-loop-helix (bHLH) is a large transcription factor family involved in several vital plant physiological processes, including growth, development, and stress response. However, no substantial reports exist on bHLH genes in rose. Here, the genome-wide identification, characterization, and expression analysis of the rose bHLH (RcbHLH) genes was carried out. In total, 100 RcbHLHs were identified in the rose genome and mapped onto different rose chromosomes. The gene duplication analysis revealed both tandem and segmental duplications in RcbHLHs. The RcbHLHs among other plant bHLHs were divided into 21 sub-groups on the phylogenetic tree. Additionally, prediction of the different cis-regulatory elements and the gene ontology of the identified RcbHLHs indicated their possible functions in rose plants. The expression analysis of selected RcbHLHs genes in two contrasting rose varieties (A29 = Black Baccara and XS = Sweet Avalanche) under B. cinerea infection provided insights into the involvement of RcbHLHs in rose–B. cinerea interactions. Moreover, this study provided details on the bHLH family genes in rose and their potential roles in rose defense against B. cinerea infection.
1. Introduction
Plants experience different biotic and abiotic stresses during their life span. These harsh conditions affect plant growth and ultimately reduce production []. To reduce the effects of these stresses, plants have developed diverse mechanisms, such as activating target genes through transcription factors (TFs) [,]. Basic helix-loop-helix (bHLH) is a large TF family and its members are found in almost all of the eukaryotes []. In the beginning, bHLHs were reported in animals involved in various processes. However, bHLHs were later identified in several plants, including Arabidopsis (147), rice (167), tomato (152), and wheat (571) [,,]. On the basis of evolution, sequence similarities, and ability of DNA binding, the bHLH is categorized into six main classes (A to F) []. The bHLH family contains a 60-amino-acid DNA binding domain with two functionally different regions; first, 10–17 aa binding site at the N-terminal, and second, 40 aa at the C-terminal known as the helix-loop-helix (HLH) region [,]. The first bHLH member was reported in maize, which regulate the anthocyanin-related genes []. Later, studies on plants showed that bHLH are involved in cell division, development, photo-morphogenesis, hormone signaling, secondary metabolites, light signaling, and biotic and abiotic stress resistance [,,,,,]. Several reports have shown that bHLHs respond to pathogens. For example, in tomato, SlybHLH131 increased the resistance against yellow leaf curl virus through controlling cell death []. Similarly, bHLH132 acts positively in the response to X. auvesicatoria. In soybean, GmPIB1, a bHLH TF, provides defense against Phytophthora root []. Similarly, in banana, several MabHLHs have been reported to confer resistance against Fusarium. oxysporum wilting [].
Rose (Rosa chinensis Jacq.) is an important dicotyledonous plant having high economic, cultural, and medicinal values []. However, the global rose production is challenged by the floral infection caused by Botrytis cinerea, a necrotrophic phytopathogen. This floral infection disease is known as gray mold in rose and is considered to be a devastating factor in rose production. Therefore, deciphering the molecular players involved in the rose–B. cinerea interactions will help in understanding the rose defense mechanisms and in developing strategies to scale rose production. In addition, the availability of genomic and transcriptomic data of rose in the public domain has facilitated the data mining, identification, and characterization of several candidate genes and TF families []. For instance, the genome-wide identification and characterization of important TFs in rose, including WRKYs, C3Hs, Dofs, and YABBYs, have been reported [,,,]. In this study, the genome-wide survey and characterization of the rose bHLH genes (RcbHLHs) was carried out. Subsequently, different protein properties and the subcellular localizations of the identified RcbHLHs were predicted. A phylogenetic analysis was performed to ascertain the evolutionary relationship of RcbHLH among themselves and other plant bHLHs. Other detail characterizations, including gene structure analysis (exon-introns), motif and domain analysis, gene ontology (GO), gene duplication and synteny analysis, and chromosomal mappings, provided further valuable insights. Lastly, the expression analysis of the selected RcbHLHs under B. cinerea infection was evaluated to understand the involvement of RcbHLHs in rose defense.
2. Results
2.1. Identification of bHLH Genes in R. chinensis and Their Chromosomal Distribution
The genome-wide search resulted in the identification of 100 bHLH genes in rose genome and named as RcbHLH1 to RcbHLH100 as per their distribution on the rose chromosomes. The initial searches and sequence retrieval resulted in 118 potential bHLH genes in rose. However, further analysis with CDD and SMART tools for the possession of conserved signature domain reduced the total to 100. The distribution of the RcbHLHs was uneven among the rose chromosomes. All 100 identified bHLHs were mapped onto the seven rose chromosomes (Figure 1). Chromosome 7 contained highest number of the bHLHs (20), whereas chromosome 3 contained the lowest (7).
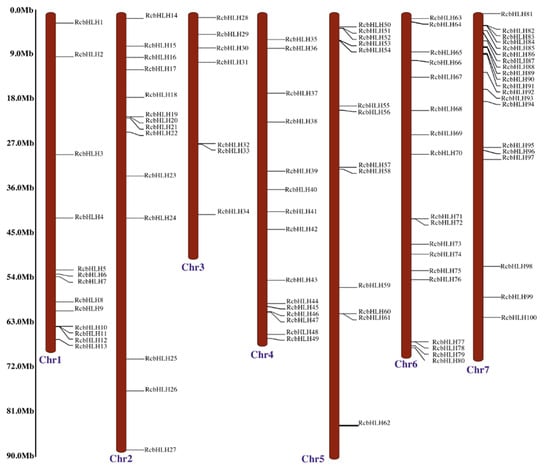
Figure 1.
Distribution of RcbHLHs on rose chromosomes. Chr represents the chromosome. The left side of the figure scale was used to represent the chromosome length and position of each RcbHLH member on the respective chromosome.
Subsequently, the predicted peptide characteristics revealed the individual molecular weight (Mw), isoelectric points (pI), and hydropathy (GRAVY) values for all 100 RcbHLHs (Supplementary Table S1). For instance, the Mw of the RcbHLHs ranged from 10.32 KDa (RcbHLH83) to 78.15 Kda (RcbHLH30). Similarly, the pI values also showed a wide range, varying from 4.66 (RcbHLH16) to 10.02 (RcbHLH30). The subcellular localization predictions suggested that the majority of the RcbHLHs localize in the nucleus, while a couple of them stay in the cytoplasm (Supplementary Table S1).
2.2. Alignment of RcbHLHs, Phylogeny, Motif, and Gene Structure Analysis
A phylogenetic tree was constructed to investigate the ancestral relationship of the identified RcbHLHs within themselves and among other plant bHLHs. The maximum likelihood (ML) tree was constructed by taking all 100 RcbHLH sequences and Arabidopsis bHLHs (AtbHLH) downloaded from the TAIR database. The results revealed that the RcbHLHs were classified into the 21 subfamilies as per the established bHLH classifications along with their homologs from Arabidopsis. The subfamily XII had the maximum number of RcbHLHs (13 genes) among the AtbHLHs, while the subfamilies II, IV(d), VIII(b), and XIV contained lowest numbers (1 gene) (Figure 2). Moreover, the individual numbers of RcbHLHs and AtbHLHs varied in a given subfamily.
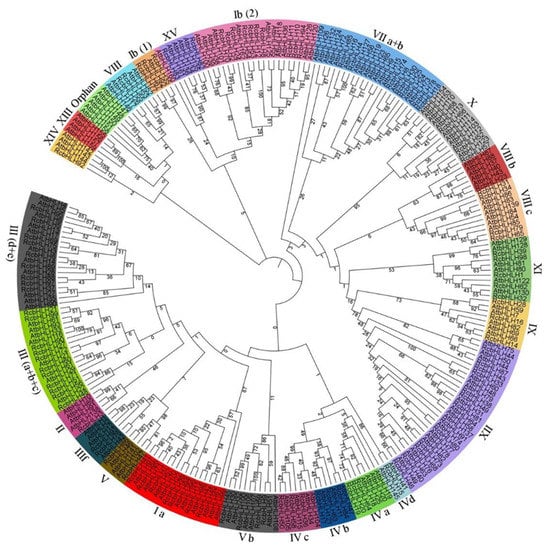
Figure 2.
Phylogenetic analysis of bHLH members of rose and Arabidopsis. Different classes were denoted by different colors.
In addition, the motif analysis of the identified RcbHLHs suggested their evolutionary relationship. In total, 12 conserved motifs were identified in the RcbHLHs (Figure 3). These identified motifs corresponded to the key domains, including the bHLH domain and bHLH-MYC-N-terminal transcription factor domain (Figure 4). Besides, the exon-intron arrangement analysis of the RcbHLHs provided important insights into the gene structures. The results revealed that the number of exons and introns showed great variations among the bHLH genes (Figure 5). For instance, RcbHLH95 has a single exon, while RcbHLH86 had 11 exons. Moreover, all of these results suggested the conserved nature of plant bHLHs and support the reliability of the current results and classifications.
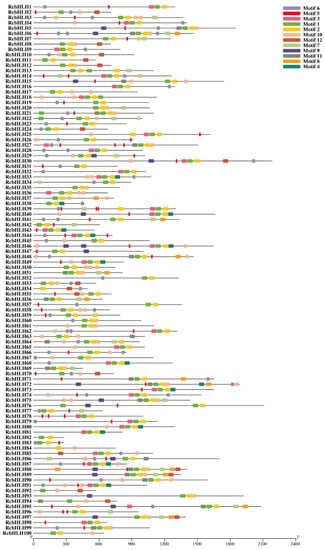
Figure 3.
Prediction of motifs in RcbHLH gene members.
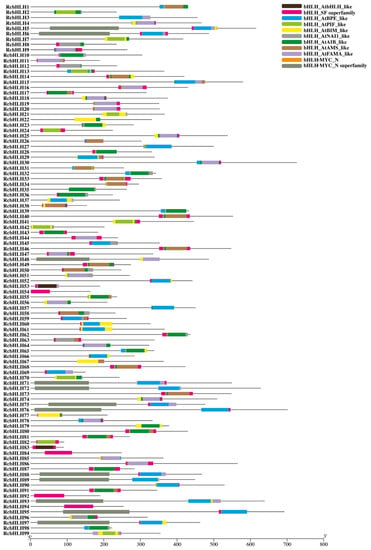
Figure 4.
Domain analysis in RcbHLH gene members.
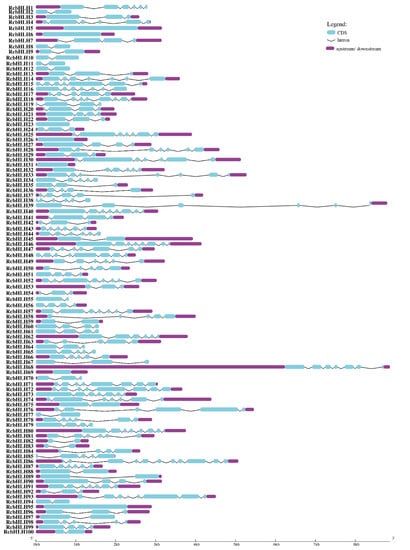
Figure 5.
Gene structure analysis of identified RcbHLH gene members. Cyan shapes represent the CDS region, purple shapes showed the up and down stream regions, and black lines represent the introns.
2.3. Collinearity and Synteny Analysis of RcbHLHs
Furthermore, a comparative collinearity analysis among rose and Arabidopsis was conducted to obtain more insights into the origin and ancestral relationships of the bHLH genes (Figure 6A). Members of RcbHLH showed collinearity with AtbHLH. Besides, the expansion of gene families can happen majorly as a result of gene duplications. Therefore, we conducted a synteny analysis of the RcbHLH genes (Figure 6B). The results revealed that both tandem and segment duplication events have occurred in the rose genome and several pairs of duplicated genes were detected on different chromosomes (Supplementary Table S2). The results suggested that these duplication events could be the major driving force in the RcbHLH evolution.
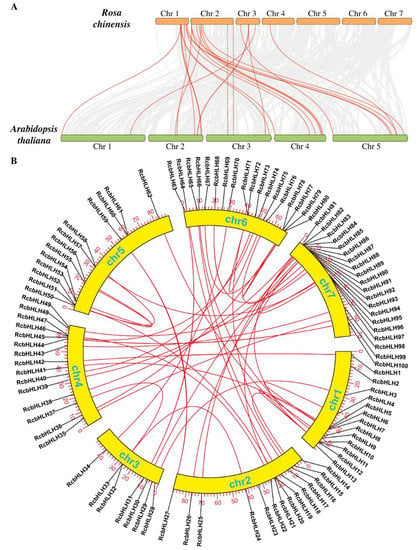
Figure 6.
Collinearity (A) and synteny analysis (B) of RcbHLH genes. Chr represents the chromosomes.
2.4. Prediction of Cis-Elements among RcbHLH Genes
The cis-regulatory elements act as enhancers or silencers of the genes. It has been previously reported that these are located in the promoter region and control the transcription of that specific gene []. In this study, 2 kb regions of each RcbHLH gene member were retrieved and cis-elements were analyzed with the help of the PLACE online database. The results revealed that different growth-related (CAT-box, AT-rich elements, and so on), hormonal regulation (ABRE, CGTCA, and TATC-box), and stress-related (MYC, MYB, and MBS) elements were found (Figure 7).
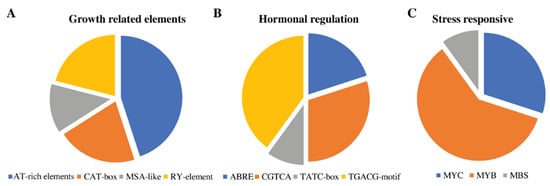
Figure 7.
Cis-regulatory elements in the 2 kb upstream region of each RcbHLH gene member. The cis-elements associated with different physiological processes are represented (A–C).
2.5. Expression Dynamics of RcbHLHs under B. cinerea Infection
To study the expression profiling of RcbHLHs under biotic stress (B. cinerea), two rose varieties, A29 (Black Baccara) and XS (Sweet Avalanche) were used, which were tolerant and susceptible to B. cinerea, respectively. After inoculation with B. cinerea, rose samples were collected at different time points for isolation of total RNA and qRT-PCR-based expression profiling. A total of 14 RcbHLHs were selected for the expression analysis under B. cinerea treatment based on the rose transcriptome data available in the public domain. Out of these 14 selected members, RcbHLH13, RcbHLH35, RcbHLH41, RcbHLH44, and RcbHLH49 were up-regulated in the XS variety as compared with A29 after 48 h of B. cinerea treatment and showed significantly higher expression (Figure 8). RcbHLH30, RcbHLH62, and RcbHLH66 were unchanged after treatment. Such sorts of discoveries connote that RcbHLHs members may be engaged in alleviating the biotic stress and provide helpful enlightening of the future practical portrayal of these genes.
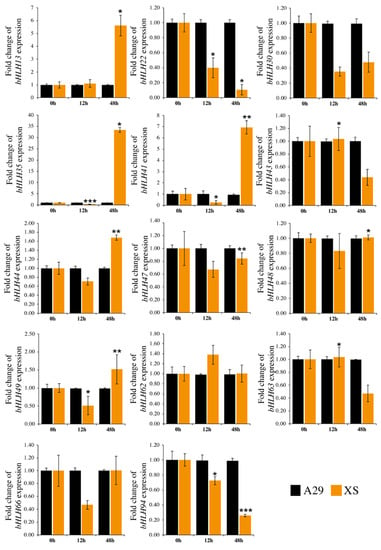
Figure 8.
Quantification of RcbHLH genes in A29 and XS varieties under control and stress conditions. Samplings were taken at 12 h and 48 h after treatment. *, *and *** are significant at p < 0.05, 0.01, and 0.001, respectively.
3. Discussion
Rose is a very important economic ornamental crop and its yield is affected by different biotic and abiotic stresses. Among the biotic stresses, the gray mold disease is a serious threat for rose production, caused by the airborne pathogen Botrytis cinerea []. To cope with these harsh situations, plants have adopted different mechanisms through the regulation of target genes by TFs [,]. The basic helix-loop-helix (bHLH) protein family is a TF family and is involved in vital plant processes, including stress response. In tomato, SlybHLH131 was involved in providing resistance against tomato yellow leaf curl virus and its mutant showed cell death []. In another study, the tomato bHLH, SlbHLH132, showed positive regulation of defense against X. auvesicatoria []. Further, the overexpression of CmbHLH87 in Arabidopsis and tobacco showed resistance against powdery mildew and other bacterial diseases []. The members of bHLH were varied from species to species, such as 41 in Carthamus tinctorius and 230 in Brassica rapa [,]. However, the characterization of bHLH in rose has not yet been explored. In this study, 100 RcbHLHs members were reported, which were mapped onto the seven rose chromosomes (Figure 1). Chromosome 7 contained the highest number of RcbHLHs (20 genes), whereas chromosome 3 contained the lowest (7 genes). These identified 100 RcbHLHs were clustered into 21 subfamilies on the bases of phylogenetic analysis. Subfamily XII had the maximum number of RcbHLHs (13 genes) among the AtbHLHs, while subfamilies II, IV(d), VIII(b), and XIV contained the lowest numbers (1 gene, Figure 2). To characterize a new protein, it is necessary to study the motif and domains in that specific protein. In the evolutionary analysis, in total, 12 conserved motifs were identified in the RcbHLHs (Figure 3). All of the identified members contain the bHLH domain and bHLH-MYC-N-terminal transcription factor domain (Figure 4). Similarly, the expression of any gene is also dependent on its gene structure, which provides clues for the evolution of the genes. For this purpose, our findings also showed the exon-intron arrangement of the RcbHLHs. The number of exons and introns showed great variations among the RcbHLHs genes (Figure 5). For instance, RcbHLH95 has a single exon, while RcbHLH86 had 11 exons. Late examinations expressed that the duplicated genes showed primary disparity, which is extremely pervasive in the age of practically distinct paralogs. This underlying uniqueness had a critical impact on the development of duplicated genes contrasted with non-duplicated genes [,].
In nature, there are two types of evolution. The first is HGT (horizontal gene transfer) and the second is gene duplication [,]. The second method is the most common method. To study the evolutionary relationship of RcbHLHs with AtbHLHs, collinearity and synteny analyses were performed. Gene duplications were calculated through Ka/Ks and the results showed that both tandem and segmental duplications were found in the rose genome (Figure 6). The cis-elements were present in the upstream region of the RcbHLHs. For genes that showed the same evolutionary clusters and co-expression under the unfavorable conditions, this might be because of the presence of cis-elements []. Similarly, it is also reported that genes related to hormones play an essential role in the plant defense system []. So, for the better understanding of the role of RcbHLHs genes in stressed conditions, we checked the cis-elements and we found growth-, hormones-, and stress-related cis-elements (Figure 7).
From the current findings and the available previous literature, it was thought that the identified RcbHLHs might be involved in the rose defense system against biotic stress. To confirm this, expression profiling of selected RcbHLHs genes as analyzed in the two contrasting genotypes of rose (Figure 8). The results showed that five genes (RcbHLH13, RcbHLH35, RcbHLH41, RcbHLH44, and RcbHLH49) were up-regulated after 48 h of B. cinerea treatment and showed significantly higher expression, and RcbHLH22, RcbHLH47, and RcbHLH94 were down-regulated. RcbHLH30, RcbHLH62, and RcbHLH66 were unchanged after treatment. Previous findings in wheat also described similar findings against the tripe rust pathogen, Fusarium pseudograminearum []. Such sorts of discoveries connote that RcbHLHs members may be engaged in alleviating biotic stress and provide helpful enlightening of the future practical portrayal of these genes.
4. Materials and Methods
4.1. Identification of RcbHLH
GDR (Genome Database for Rosaceae, https://www.rosaceae.org) was used for the retrieval of full-length sequences of rose in the form of gff3 files [], while the bHLH members of Arabidopsis were retrieved using Plant Transcription Factor Database (PlantTFDB, http://planttfdb.gao-lab.org/) []. Arabidopsis bHLHs were used as a reference and rose bHLH members were searched using the BLAST method with the help of TBtool and NCBI []. Confirmation of RcbHLH members and Pfam domain (PF00010) among them was confirmed with the help of HMMER and Batch CD-Search tool [,,]. Furthermore, the physiochemical properties of RcbHLH were studied with the help of ExPASy []. The position of RcbHLH at the cellular level was predicted using CELLO 2.5 and WoLF PSORT [,].
4.2. Prediction of Gene Structure, Conserved Motifs, and Cis-Regulatory Elements
For the prediction of genes’ structure pattern, Gene Structure Display Server (GSDS) was utilized []. MEME suite, with a setting of 1 site per sequence and 12 total motifs of width six to hundred amino acids, was utilized for the prediction of motifs in RcbHLH proteins. The visualization of identified motifs was carried out with TBtool [,]. The genome of R. chinensis (AP85-441) was downloaded and 1.5 kb upstream sequences of every RcbHLH member were obtained and used for the identification of cis-elements. For this purpose, PlantCare was utilized [].
4.3. Phylogenetic Analysis
ClustalW and MEGA with default settings were implemented for the alignment of RcbHLHs among them and with bHLH members of A. thaliana []. In the end, a phylogenetic tree was developed by applying the neighbor-joining method (NJ). During these steps, 1000 replicates were adjusted.
4.4. Chromosomal Location and Gene Duplication Analysis
The obtained gff3-files of the R. chinensis genome were analyzed using TBtool and each RcbHLH was mapped onto the respective chromosomes. Gene duplication analysis was performed by setting the following parameters: length of the aligned genes was more than 90%, similarities between the matched region were also set to be greater than 90%, and genes had one duplication process.
4.5. Plant, Fungal Growth, and Inoculation
The Rosa chinensis varieties (A29 (Black Baccara) = tolerant and XS (Sweet Avalanche) = susceptible) were grown under control conditions in different parts of the Yunnan province, China. During this study, light was provided for 14 h and dark for 10 h, while the temperature was maintained at 22 ± 2 °C. Sampling was performed as previously described by []. In detail, fully opened buds of rose plants were selected and their stems were cut and directly rinsed in water. Further cutting up to 20 cm was performed before processing. The discs of rose petals of 12.5 mm in size were prepared and kept on liquid 0.4% agar. Spores of Botrytis cinerea (KMSC-011) were used for inoculation []. Sampling was performed at 0 h, 12 h, and 48 h after inoculation.
4.6. Expression Analysis
For the verifications of RNA-Seq data, fourteen (RcbHLHs) genes were taken and their expression profiling was performed through qRT-PCR (StepOnePlus Real-Time PCR System, Thermo Fisher Scientific). For this purpose, after treatment with Botrytis cinerea, 2 g of inoculated leaves were ground in liquid nitrogen at different time points (0, 12, and 48 h). RNA was extracted by following the instructions of Tiangen kit (DP441, RT411) and three biological same samples were pooled together. Moreover, 500 ng cDNA was synthesized using Takara Reverse Transcriptase M-MLV, and KAPATM SYBRR quantitative PCR kit (Takara) was used. RhActin1 was utilized for normalization []. Gene-specific primers used during this study are provided in Supplementary Table S1.
5. Conclusions
In this study, we found 100 RcbHLH genes in rose genome. Their genome-wide characterization and functional expectations uncovered more about their rationed nature and putative capabilities. Furthermore, the expression analysis of all RcbHLHs added valuable insights into their involvement in rose–Botrytis cinerea interactions. Five identified genes (RcbHLH13, RcbHLH35, RcbHLH41, RcbHLH44, and RcbHLH49) could be involved in the rose resistance against the gray mold pathogen. These identified genes can be further explored in the near future for further functional characterization.
Supplementary Materials
The following supporting information can be downloaded at https://www.mdpi.com/article/10.3390/horticulturae8110989/s1, Table S1: Physiochemical properties of identified RcbHLH genes members; Table S2: Gene duplication events among RcbHLHs genes; Table S3: List of primers used in this study.
Author Contributions
Conceptualization, I.U. and H.W.; methodology, validation, formal analysis, investigation, data curation, I.U., W.Y., M.U., O.U.R. and S.L.; writing—original draft preparation, I.U., W.Y. and M.U.; writing—review and editing, S.N. and H.W.; supervision, H.W. and S.N.; funding acquisition, H.W. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by National Key Research and Development Program of China (Grant No. 2018YFD1000407) and Science and Technology Program of Yunnan Province (Grant No. 202102AE090001).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
I.U. thanks the Govt. of China for providing the fully-funded Master’s scholarship.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Rahim, A.A.; Uzair, M.; Rehman, N.; Rehman, O.U.; Zahra, N.; Khan, M.R. Genome-Wide Identification and Characterization of Receptor-Like Protein Kinase 1 (RPK1) Gene Family in Triticum aestivum under Drought Stress. Front. Genet. 2022, 13, 912251. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.; Xiong, R.; Liu, H.; Wu, M.; Chen, F.; Yan, H.; Xiang, Y. Basic helix-loop-helix gene family: Genome wide identification, phylogeny, and expression in Moso bamboo. Plant Physiol. Biochem. 2018, 132, 104–119. [Google Scholar] [CrossRef] [PubMed]
- Javed, T.; Shabbir, R.; Ali, A.; Afzal, I.; Zaheer, U.; Gao, S.-J. Transcription Factors in Plant Stress Responses: Challenges and Potential for Sugarcane Improvement. Plants 2020, 9, 491. [Google Scholar] [CrossRef] [PubMed]
- Feller, A.; Machemer, K.; Braun, E.L.; Grotewold, E. Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J. 2011, 66, 94–116. [Google Scholar] [CrossRef]
- Ke, Y.-Z.; Wu, Y.-W.; Zhou, H.-J.; Chen, P.; Wang, M.-M.; Liu, M.-M.; Li, P.-F.; Yang, J.; Li, J.-N.; Du, H. Genome-wide survey of the bHLH super gene family in Brassica napus. BMC Plant Biol. 2020, 20, 115. [Google Scholar] [CrossRef]
- Li, X.; Duan, X.; Jiang, H.; Sun, Y.; Tang, Y.; Yuan, Z.; Guo, J.; Liang, W.; Chen, L.; Yin, J.; et al. Genome-Wide Analysis of Basic/Helix-Loop-Helix Transcription Factor Family in Rice and Arabidopsis. Plant Physiol. 2006, 141, 1167–1184. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hu, Z.; Zhao, T.; Yang, Y.; Chen, T.; Yang, M.; Yu, W.; Zhang, B. Genome-wide analysis of bHLH transcription factor and involvement in the infection by yellow leaf curl virus in tomato (Solanum lycopersicum). BMC Genom. 2015, 16, 39. [Google Scholar] [CrossRef] [PubMed]
- Mao, K.; Dong, Q.; Li, C.; Liu, C.; Ma, F. Genome Wide Identification and Characterization of Apple bHLH Transcription Factors and Expression Analysis in Response to Drought and Salt Stress. Front. Plant Sci. 2017, 8, 480. [Google Scholar] [CrossRef] [PubMed]
- Pires, N.; Dolan, L. Origin and Diversification of Basic-Helix-Loop-Helix Proteins in Plants. Mol. Biol. Evol. 2010, 27, 862–874. [Google Scholar] [CrossRef] [PubMed]
- Toledo-Ortiz, G.; Huq, E.; Quail, P.H. The Arabidopsis Basic/Helix-Loop-Helix Transcription Factor Family. Plant Cell 2003, 15, 1749–1770. [Google Scholar] [CrossRef] [PubMed]
- Chandler, V.L.; Radicella, J.P.; Robbins, T.P.; Chen, J.; Turks, D. Two regulatory genes of the maize anthocyanin pathway are homologous: Isolation of B utilizing R genomic sequences. Plant Cell 1989, 1, 1175–1183. [Google Scholar]
- Baillo, E.H.; Kimotho, R.N.; Zhang, Z.; Xu, P. Transcription Factors Associated with Abiotic and Biotic Stress Tolerance and Their Potential for Crops Improvement. Genes 2019, 10, 771. [Google Scholar] [CrossRef]
- Buti, S.; Hayes, S.; Pierik, R. The bHLH network underlying plant shade-avoidance. Physiol. Plant 2020, 169, 312–324. [Google Scholar] [CrossRef] [PubMed]
- Chezem, W.R.; Clay, N.K. Regulation of plant secondary metabolism and associated specialized cell development by MYBs and bHLHs. Phytochemistry 2016, 131, 26–43. [Google Scholar] [CrossRef]
- Dong, H.; Chen, Q.; Dai, Y.; Hu, W.; Zhang, S.; Huang, X. Genome-wide identification of PbrbHLH family genes, and expression analysis in response to drought and cold stresses in pear (Pyrus bretschneideri). BMC Plant Biol. 2021, 21, 86. [Google Scholar] [CrossRef]
- Komatsu, M.; Maekawa, M.; Shimamoto, K.; Kyozuka, J. The LAX1 and FRIZZY PANICLE 2 genes determine the inflorescence architecture of rice by controlling rachis-branch and spikelet development. Dev. Biol. 2001, 231, 364–373. [Google Scholar] [CrossRef] [PubMed]
- Meraj, T.A.; Fu, J.; Raza, M.A.; Zhu, C.; Shen, Q.; Xu, D.; Wang, Q. Transcriptional Factors Regulate Plant Stress Responses Through Mediating Secondary Metabolism. Genes 2020, 11, 346. [Google Scholar] [CrossRef]
- Wang, Z.; Jia, C.; Wang, J.-Y.; Miao, H.-X.; Liu, J.-H.; Chen, C.; Yang, H.-X.; Xu, B.; Jin, Z. Genome-Wide Analysis of Basic Helix-Loop-Helix Transcription Factors to Elucidate Candidate Genes Related to Fruit Ripening and Stress in Banana (Musa acuminata L. AAA Group, cv. Cavendish). Front. Plant Sci. 2020, 11, 650. [Google Scholar] [CrossRef]
- Liu, X.; Li, D.; Zhang, S.; Xu, Y.; Zhang, Z. Genome-wide characterization of the rose (Rosa chinensis) WRKY family and role of RcWRKY41 in gray mold resistance. BMC Plant Biol. 2019, 19, 522. [Google Scholar] [CrossRef] [PubMed]
- Jung, S.; Ficklin, S.; Lee, T.; Cheng, C.-H.; Blenda, A.; Zheng, P.; Yu, J.; Bombarely, A.; Cho, I.; Ru, S.; et al. The Genome Database for Rosaceae (GDR): Year 10 update. Nucleic Acids Res. 2014, 42, D1237–D1244. [Google Scholar] [CrossRef]
- Ullah, I.; Abbas, A.; Hussain, S.; Nanda, S. Genome-wide identification and expression analysis of the RcYABBY s reveals their potential functions in rose (Rosa chinensis Jacq.). J. Hortic. Sci. Biotechnol. 2022, 97, 593–602. [Google Scholar] [CrossRef]
- Nan, H.; Ludlow, R.A.; Lu, M.; An, H. Genome-Wide Analysis of Dof Genes and Their Response to Abiotic Stress in Rose (Rosa chinensis). Front. Genet. 2021, 12, 538733. [Google Scholar] [CrossRef]
- Li, C.-H.; Fang, Q.-X.; Zhang, W.-J.; Li, Y.-H.; Zhang, J.-Z.; Chen, S.; Yin, Z.-G.; Li, W.-J.; Liu, W.-D.; Yi, Z. Genome-wide identification of the CCCH gene family in rose (Rosa chinensis Jacq.) reveals its potential functions. Biotechnol. Biotechnol. Equip. 2021, 35, 517–526. [Google Scholar] [CrossRef]
- Rasool, F.; Uzair, M.; Naeem, M.K.; Rehman, N.; Afroz, A.; Shah, H.; Khan, M.R. Phenylalanine ammonia-lyase (PAL) genes family in wheat (Triticum aestivum L.): Genome-wide characterization and expression profiling. Agronomy 2021, 11, 2511. [Google Scholar] [CrossRef]
- Williamson, B.; Tudzynski, B.; Tudzynski, P.; Van Kan, J.A.L. Botrytis cinerea: The cause of grey mould disease. Mol. Plant Pathol. 2007, 8, 561–580. [Google Scholar] [CrossRef] [PubMed]
- Hao, Y.; Zong, X.; Ren, P.; Qian, Y.; Fu, A. Basic Helix-Loop-Helix (bHLH) Transcription Factors Regulate a Wide Range of Functions in Arabidopsis. Int. J. Mol. Sci. 2021, 22, 7152. [Google Scholar] [CrossRef] [PubMed]
- Rout, S.S.; Rout, P.; Uzair, M.; Kumar, G.; Nanda, S. Genome-wide identification and expression analysis of CRK gene family in chili pepper (Capsicum annuum L.) in response to Colletotrichum truncatum infection. J. Hortic. Sci. Biotechnol. 2022, 11, 118. [Google Scholar] [CrossRef]
- Kim, J.-G.; Mudgett, M.B. Tomato bHLH132 Transcription Factor Controls Growth and Defense and Is Activated by Xanthomonas euvesicatoria Effector XopD During Pathogenesis. Mol. Plant-Microbe Interactions 2019, 32, 1614–1622. [Google Scholar] [CrossRef]
- Guo, W.-L.; Chen, B.-H.; Guo, Y.-Y.; Chen, X.-J.; Li, Q.-F.; Yang, H.-L.; Li, X.-Z.; Zhou, J.-G.; Wang, G.-Y. Expression of Pumpkin CmbHLH87 Gene Improves Powdery Mildew Resistance in Tobacco. Front. Plant Sci. 2020, 11, 163. [Google Scholar] [CrossRef]
- Song, X.-M.; Huang, Z.-N.; Duan, W.-K.; Ren, J.; Liu, T.-K.; Li, Y.; Hou, X.-L. Genome-wide analysis of the bHLH transcription factor family in Chinese cabbage (Brassica rapa ssp. pekinensis). Mol. Genet. Genom. 2014, 289, 77–91. [Google Scholar] [CrossRef]
- Hong, Y.; Ahmad, N.; Tian, Y.; Liu, J.; Wang, L.; Wang, G.; Liu, X.; Dong, Y.; Wang, F.; Liu, W.; et al. Genome-Wide Identification, Expression Analysis, and Subcellular Localization of Carthamus tinctorius bHLH Transcription Factors. Int. J. Mol. Sci. 2019, 20, 3044. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.; Wang, S.; Xu, D.; Liu, X.; Li, X.; Xiao, W.; Cao, J.; Jiang, H.; Min, X.; Wang, J.; et al. Identification and Analysis of the GASR Gene Family in Common Wheat (Triticum aestivum L.) and Characterization of TaGASR34, a Gene Associated with Seed Dormancy and Germination. Front. Genet. 2019, 10, 980. [Google Scholar] [CrossRef]
- Rehman, O.U.; Uzair, M.; Chao, H.; Fiaz, S.; Khan, M.R.; Chen, M. Role of the type-B authentic response regulator gene family in fragrant rice under alkaline salt stress. Physiol. Plant 2022, 174, e13696. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Gui, S.; Wang, S.; Ding, Y. Molecular evolution and functional characterisation of an ancient phenylalanine ammonia-lyase gene (NnPAL1) from Nelumbo nucifera: Novel insight into the evolution of the PAL family in angiosperms. BMC Evol. Biol. 2014, 14, 100. [Google Scholar] [CrossRef]
- Liu, X.; Cao, X.; Shi, S.; Zhao, N.; Li, D.; Fang, P.; Chen, X.; Qi, W.; Zhang, Z. Comparative RNA-Seq analysis reveals a critical role for brassinosteroids in rose (Rosa hybrida) petal defense against Botrytis cinerea infection. BMC Genet. 2018, 19, 62. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Xiang, L.; Hong, J.; Xie, Z.; Li, B. Genome-wide analysis of bHLH transcription factor family reveals their involvement in biotic and abiotic stress responses in wheat (Triticum aestivum L.). 3 Biotech 2019, 9, 236. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Tian, F.; Yang, D.-C.; Meng, Y.-Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2016, 45, D1040–D1045. [Google Scholar] [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Clements, J.; Eddy, S.R. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 2011, 39, W29–W37. [Google Scholar] [CrossRef] [PubMed]
- Gilchrist, C.L.M.; Chooi, Y.-H. Synthaser: A CD-Search enabled Python toolkit for analysing domain architecture of fungal secondary metabolite megasynth(et)ases. Fungal Biol. Biotechnol. 2021, 8, 13. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef] [PubMed]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef] [PubMed]
- Pierleoni, A.; Martelli, P.L.; Fariselli, P.; Casadio, R. BaCelLo: A Balanced Subcellular Localization predictor. Bioinformatics 2006, 22, e408–e416. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.-J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef] [PubMed]
- Guo, A.-Y.; Zhu, Q.-H.; Chen, X.; Luo, J.-C. GSDS: A gene structure display server. Yi Chuan Hered. 2007, 29, 1023–1026. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, w202–w208. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Quidde, T.; Osbourn, A.; Tudzynski, P. Detoxification of α-tomatine byBotrytis cinerea. Physiol. Mol. Plant Pathol. 1998, 52, 151–165. [Google Scholar] [CrossRef]
- Meng, Y.; Li, N.; Tian, J.; Gao, J.; Zhang, C. Identification and validation of reference genes for gene expression studies in postharvest rose flower (Rosa hybrida). Sci. Hortic. 2013, 158, 16–21. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).