Effects of Intramolecular Distance between Amyloidogenic Domains on Amyloid Aggregation
Abstract
:1. Introduction
2. Results and Discussion
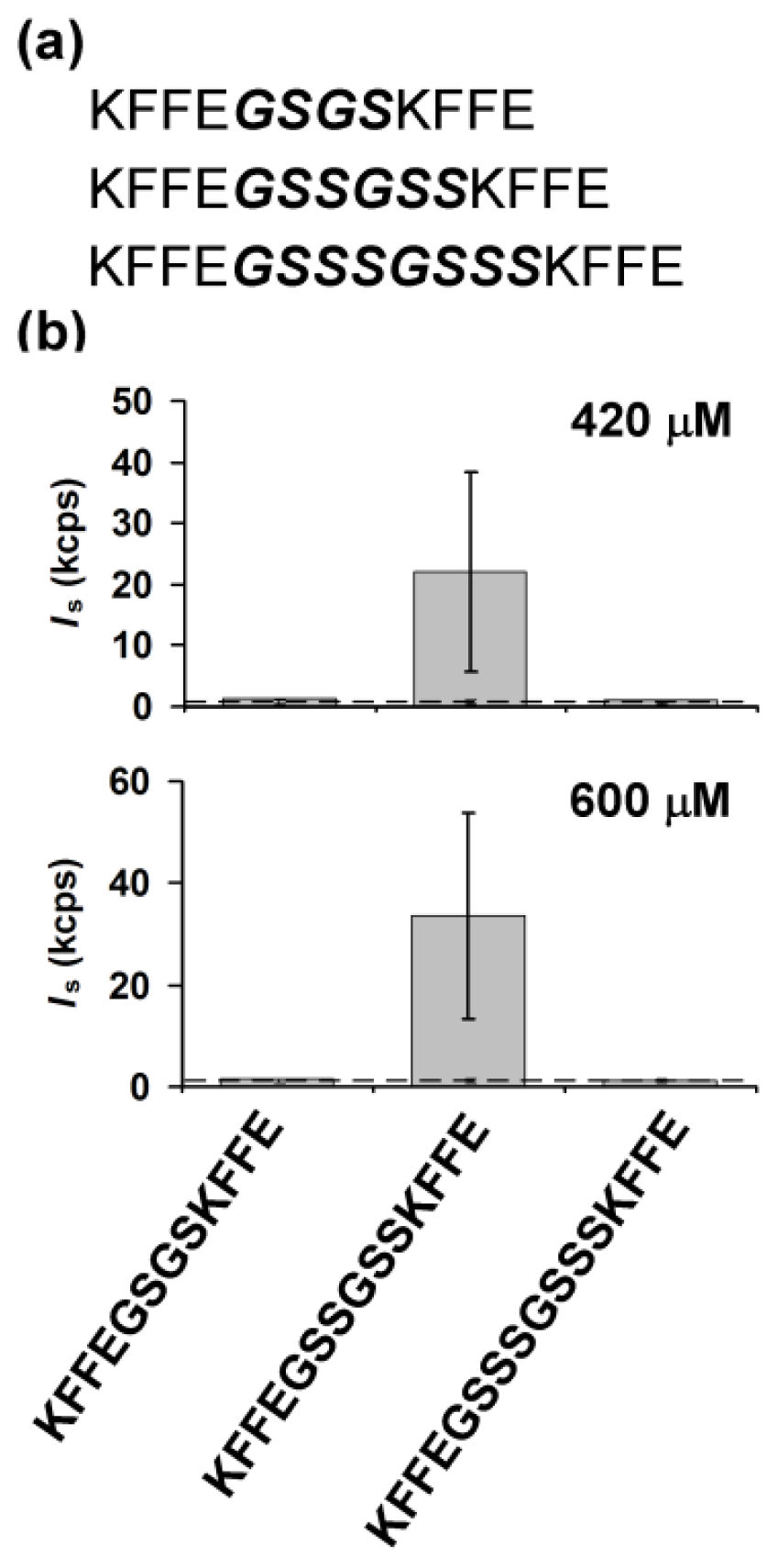
2.1. Design of Linker Sequences Connecting Two KFFE Domains
2.2. Effects of a Linker Length on Early Stage Aggregation of Peptides
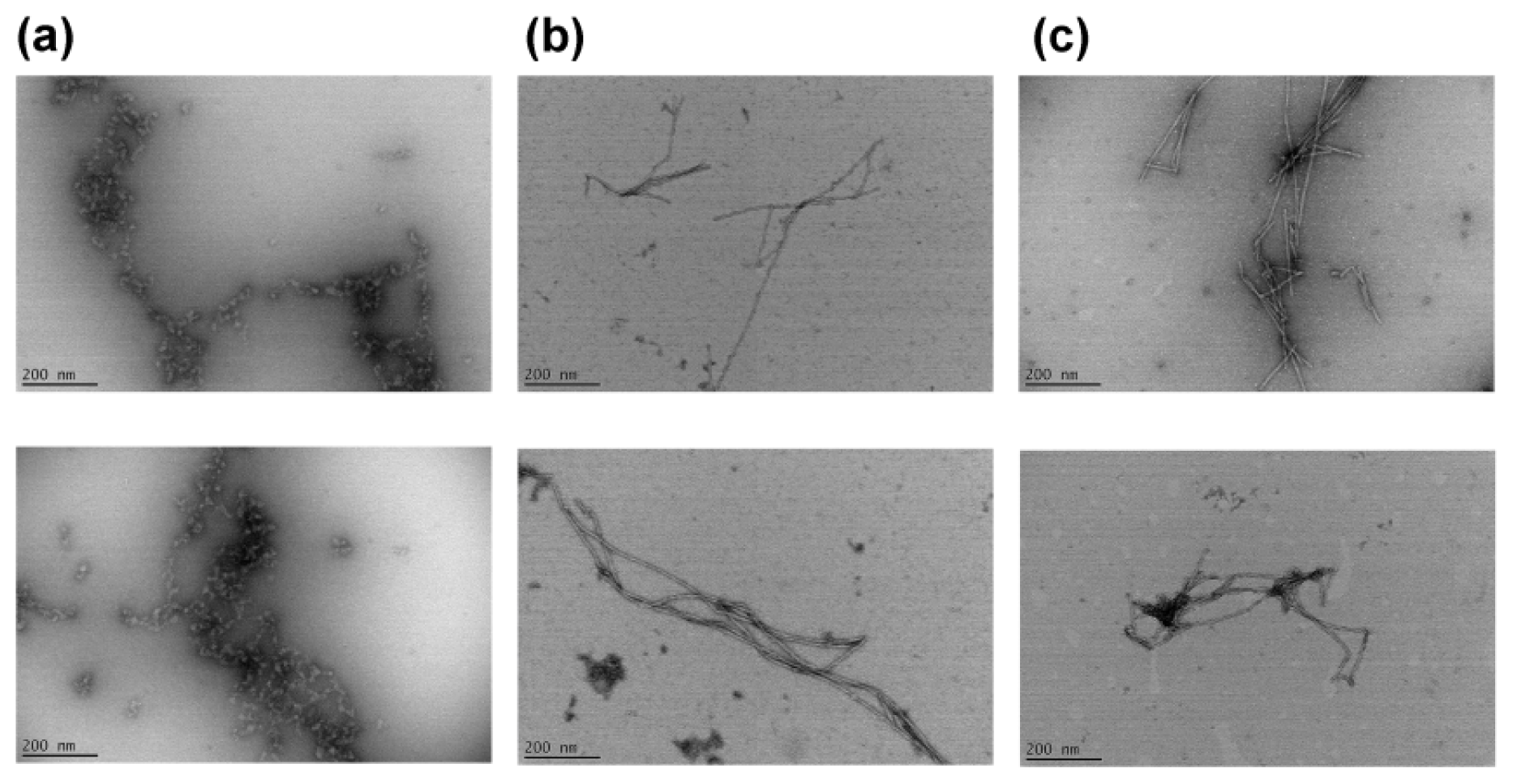
2.3. Effects of a Linker Length on the Morphology of Peptide Aggregates
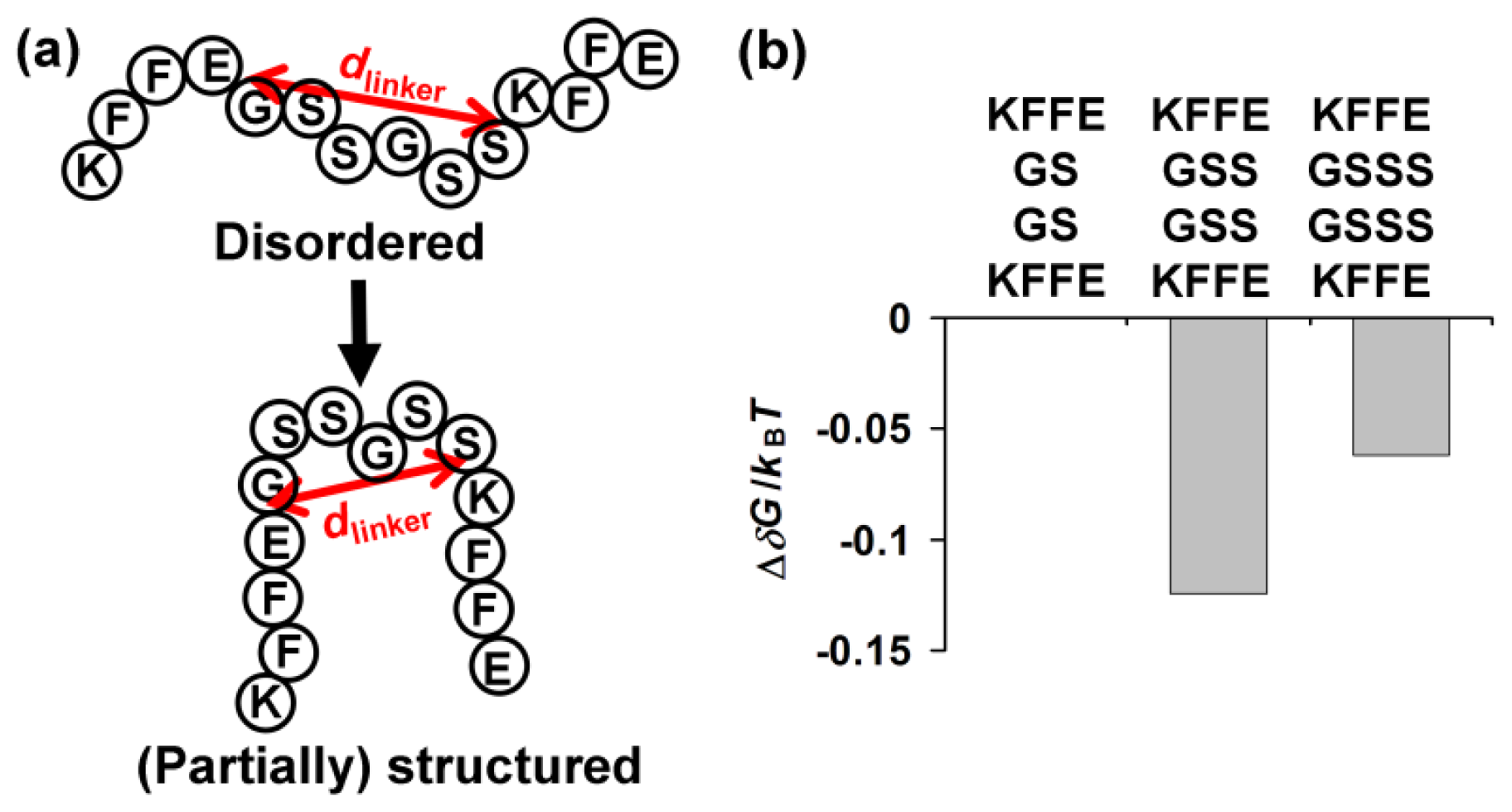
2.4. Entropic Contribution of a Linker Length toward Formation of (Partially) Structured Intermediates
3. Experimental Section
3.1. Materials
3.2. Sample Preparation
3.3. Laser Light Scattering
3.4. Transmission Electron Microscopy (TEM)
3.5. Circular Dichroism (CD) Spectroscopy
3.6. Thioflavin T (ThT) Fluorescence
4. Conclusions
Supplementary Information
Theoretical analysis on the free energy of formation of (partially) structured intermediates as a function of linker length
Acknowledgments
References
- Aguzzi, A.; O’Connor, T. Protein aggregation diseases: Pathogenicity and therapeutic perspectives. Nat. Rev. Drug Discov 2010, 9, 237–248. [Google Scholar]
- Goldsbury, C.; Frey, P.; Olivieri, V.; Aebi, U.; Muller, S.A. Multiple assembly pathways underlie amyloid-beta fibril polymorphisms. J. Mol. Biol 2005, 352, 282–298. [Google Scholar]
- Kodali, R.; Wetzel, R. Polymorphism in the intermediates and products of amyloid assembly. Curr. Opin. Struct. Biol 2007, 17, 48–57. [Google Scholar]
- Tartaglia, G.G.; Vendruscolo, M. The Zyggregator method for predicting protein aggregation propensities. Chem. Soc. Rev 2008, 37, 1395–1401. [Google Scholar]
- Chiti, F.; Stefani, M.; Taddei, N.; Ramponi, G.; Dobson, C.M. Rationalization of the effects of mutations on peptide and protein aggregation rates. Nature 2003, 424, 805–808. [Google Scholar]
- Fernandez-Escamilla, A.M.; Rousseau, F.; Schymkowitz, J.; Serrano, L. Prediction of sequence-dependent and mutational effects on the aggregation of peptides and proteins. Nat. Biotechnol 2004, 22, 1302–1306. [Google Scholar]
- Hu, Y.; Zheng, H.; Su, B.; Hernandez, M.; Kim, J.R. Modulation of β-amyloid aggregation by engineering the sequence connecting beta-strand forming domains. Biochim. Biophys. Acta 2012, 1824, 1069–1079. [Google Scholar]
- Hernandez, M.; Golbert, S.; Zhang, L.G.; Kim, J.R. Creation of aggregation-defective α-synuclein variants by engineering the sequence connecting β-strand-forming domains. Chembiochem 2011, 12, 2630–2639. [Google Scholar]
- Dobson, C.M. Protein misfolding, evolution and disease. Trends Biochem. Sci 1999, 24, 329–332. [Google Scholar]
- Dobson, C.M. Protein folding and misfolding. Nature 2003, 426, 884–890. [Google Scholar]
- Pawar, A.P.; Dubay, K.F.; Zurdo, J.; Chiti, F.; Vendruscolo, M.; Dobson, C.M. Prediction of “aggregation-prone” and “aggregation-susceptible” regions in proteins associated with neurodegenerative diseases. J. Mol. Biol 2005, 350, 379–392. [Google Scholar]
- Lopez de la Paz, M.; Serrano, L. Sequence determinants of amyloid fibril formation. Proc. Natl. Acad. Sci. USA 2004, 101, 87–92. [Google Scholar]
- Caflisch, A. Computational models for the prediction of polypeptide aggregation propensity. Curr. Opin. Chem. Biol 2006, 10, 437–444. [Google Scholar]
- Cohen, S.I.; Vendruscolo, M.; Dobson, C.M.; Knowles, T.P. From macroscopic measurements to microscopic mechanisms of protein aggregation. J. Mol. Biol 2012, 421, 160–171. [Google Scholar]
- Miller, Y.; Ma, B.; Nussinov, R. Polymorphism in Alzheimer Abeta amyloid organization reflects conformational selection in a rugged energy landscape. Chem. Rev 2010, 110, 4820–4838. [Google Scholar]
- Murphy, R.M. Peptide aggregation in neurodegenerative disease. Annu. Rev. Biomed. Eng 2002, 4, 155–174. [Google Scholar]
- Tjernberg, L.; Hosia, W.; Bark, N.; Thyberg, J.; Johansson, J. Charge attraction and beta propensity are necessary for amyloid fibril formation from tetrapeptides. J. Biol. Chem 2002, 277, 43243–43246. [Google Scholar]
- Melquiond, A.; Mousseau, N.; Derreumaux, P. Structures of soluble amyloid oligomers from computer simulations. Proteins 2006, 65, 180–191. [Google Scholar]
- Hosia, W.; Bark, N.; Liepinsh, E.; Tjernberg, A.; Persson, B.; Hallen, D.; Thyberg, J.; Johansson, J.; Tjernberg, L. Folding into a β-hairpin can prevent amyloid fibril formation. Biochemistry 2004, 43, 4655–4661. [Google Scholar]
- Argos, P. An investigation of oligopeptides linking domains in protein tertiary structures and possible candidates for general gene fusion. J. Mol. Biol 1990, 211, 943–958. [Google Scholar]
- Wriggers, W.; Chakravarty, S.; Jennings, P.A. Control of protein functional dynamics by peptide linkers. Biopolymers 2005, 80, 736–746. [Google Scholar]
- Edwards, W.R.; Busse, K.; Allemann, R.K.; Jones, D.D. Linking the functions of unrelated proteins using a novel directed evolution domain insertion method. Nucleic Acids Res 2008, 36, e78. [Google Scholar]
- Chou, P.Y.; Fasman, G.D. Empirical predictions of protein conformation. Annu. Rev. Biochem 1978, 47, 251–276. [Google Scholar]
- Kyte, J.; Doolittle, R.F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol 1982, 157, 105–132. [Google Scholar]
- George, R.A.; Heringa, J. An analysis of protein domain linkers: Their classification and role in protein folding. Protein Eng 2002, 15, 871–879. [Google Scholar]
- Zhou, H.X. Effect of backbone cyclization on protein folding stability: Chain entropies of both the unfolded and the folded states are restricted. J. Mol. Biol 2003, 332, 257–264. [Google Scholar]
- Baumketner, A.; Shea, J.E. Free energy landscapes for amyloidogenic tetrapeptides dimerization. Biophys. J 2005, 89, 1493–1503. [Google Scholar]
- Pallitto, M.M.; Murphy, R.M. A mathematical model of the kinetics of beta-amyloid fibril growth from the denatured state. Biophys. J 2001, 81, 1805–1822. [Google Scholar]
- Rucker, A.L.; Creamer, T.P. Polyproline II helical structure in protein unfolded states: Lysine peptides revisited. Protein Sci 2002, 11, 980–985. [Google Scholar]
- Gokce, I.; Woody, R.W.; Anderluh, G.; Lakey, J.H. Single peptide bonds exhibit poly(pro)II (“random coil”) circular dichroism spectra. J. Am. Chem. Soc 2005, 127, 9700–9701. [Google Scholar]
- Eker, F.; Griebenow, K.; Schweitzer-Stenner, R. Stable conformations of tripeptides in aqueous solution studied by UV circular dichroism spectroscopy. J. Am. Chem. Soc 2003, 125, 8178–8185. [Google Scholar]
- Tiffany, M.L.; Krimm, S. Effect of temperature on the circular dichroism spectra of polypeptides in the extended state. Biopolymers 1972, 11, 2309–2316. [Google Scholar]
- LeVine, H., III. Quantification of β-sheet amyloid fibril structures with thioflavin T. Methods Enzymol. 1999, 309, 274–284. [Google Scholar]
- Lazo, N.D.; Grant, M.A.; Condron, M.C.; Rigby, A.C.; Teplow, D.B. On the nucleation of amyloid β-protein monomer folding. Protein Sci 2005, 14, 1581–1596. [Google Scholar]
- Qin, Z.; Hu, D.; Zhu, M.; Fink, A.L. Structural characterization of the partially folded intermediates of an immunoglobulin light chain leading to amyloid fibrillation and amorphous aggregation. Biochemistry 2007, 46, 3521–3531. [Google Scholar]
- Munishkina, L.A.; Ahmad, A.; Fink, A.L.; Uversky, V.N. Guiding protein aggregation with macromolecular crowding. Biochemistry 2008, 47, 8993–9006. [Google Scholar]
- Zhou, H.X. Loops, linkages, rings, catenanes, cages, and crowders: Entropy-based strategies for stabilizing proteins. Acc. Chem. Res 2004, 37, 123–130. [Google Scholar]
- Zhou, H.X. Loops in proteins can be modeled as wormlike chains. J. Phys. Chem. B 2001, 105, 6763–6766. [Google Scholar]
- Serpell, L.C.; Berriman, J.; Jakes, R.; Goedert, M.; Crowther, R.A. Fiber diffraction of synthetic α-synuclein filaments shows amyloid-like cross-β conformation. Proc. Natl. Acad. Sci. USA 2000, 97, 4897–4902. [Google Scholar]
- Petkova, A.T.; Ishii, Y.; Balbach, J.J.; Antzutkin, O.N.; Leapman, R.D.; Delaglio, F.; Tycko, R. A structural model for Alzheimer’s β-amyloid fibrils based on experimental constraints from solid state NMR. Proc. Natl. Acad. Sci. USA 2002, 99, 16742–16747. [Google Scholar]
- Tartaglia, G.G.; Cavalli, A.; Pellarin, R.; Caflisch, A. Prediction of aggregation rate and aggregation-prone segments in polypeptide sequences. Protein Sci 2005, 14, 2723–2734. [Google Scholar]
- Tofoleanu, F.; Buchete, N.V. Molecular interactions of Alzheimer’s Abeta protofilaments with lipid membranes. J. Mol. Biol 2012, 421, 572–586. [Google Scholar]
- Tofoleanu, F.; Buchete, N.V. Alzheimer’s Abeta peptide interactions with lipid membranes: Fibrils, oligomers and polymorphic amyloid channels. Prion 2012, 6, 1–7. [Google Scholar]
- Gobush, W.; Stockmayer, W.H.; Yamakawa, H.; Magee, W.S. Statistical mechanics of wormlike chains. 1. Asymptotic Behavior. J. Chem. Phys 1972, 57, 2839–2843. [Google Scholar]
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ko, A.; Kim, J.R. Effects of Intramolecular Distance between Amyloidogenic Domains on Amyloid Aggregation. Int. J. Mol. Sci. 2012, 13, 12169-12181. https://doi.org/10.3390/ijms131012169
Ko A, Kim JR. Effects of Intramolecular Distance between Amyloidogenic Domains on Amyloid Aggregation. International Journal of Molecular Sciences. 2012; 13(10):12169-12181. https://doi.org/10.3390/ijms131012169
Chicago/Turabian StyleKo, Ahra, and Jin Ryoun Kim. 2012. "Effects of Intramolecular Distance between Amyloidogenic Domains on Amyloid Aggregation" International Journal of Molecular Sciences 13, no. 10: 12169-12181. https://doi.org/10.3390/ijms131012169



