Cis- and Trans-Encoded Small Regulatory RNAs in Bacillus subtilis
Abstract
:1. Introduction
2. Base-Pairing sRNAs
2.1. Cis-Encoded sRNAs (Bona Fide Antisense RNAs) in B. subtilis
2.1.1. Antitoxins in Type I Toxin–Antitoxin Systems
2.1.2. Other Cis-Encoded sRNAs in B. subtilis
2.2. Trans-Encoded sRNAs in B. subtilis
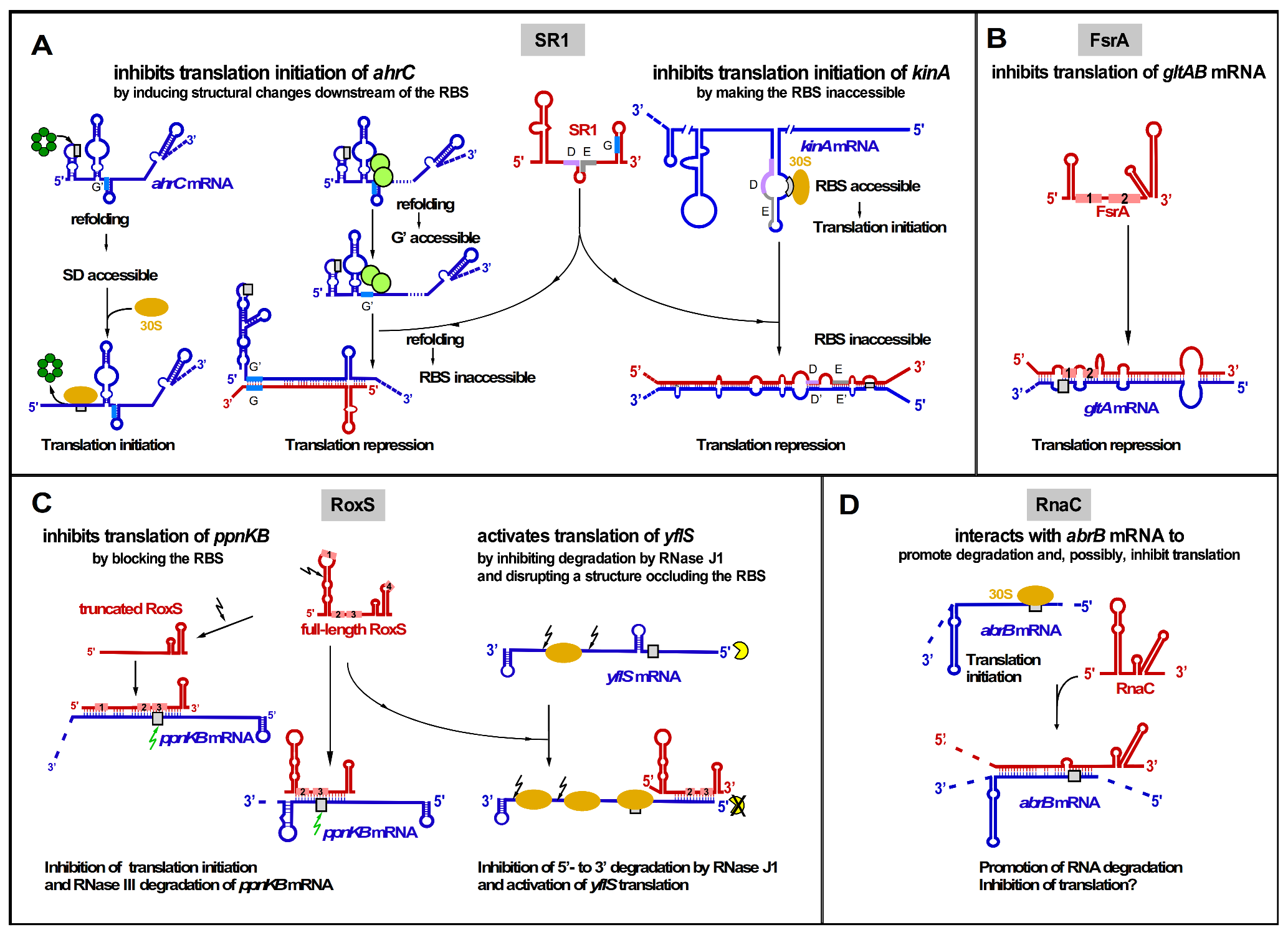
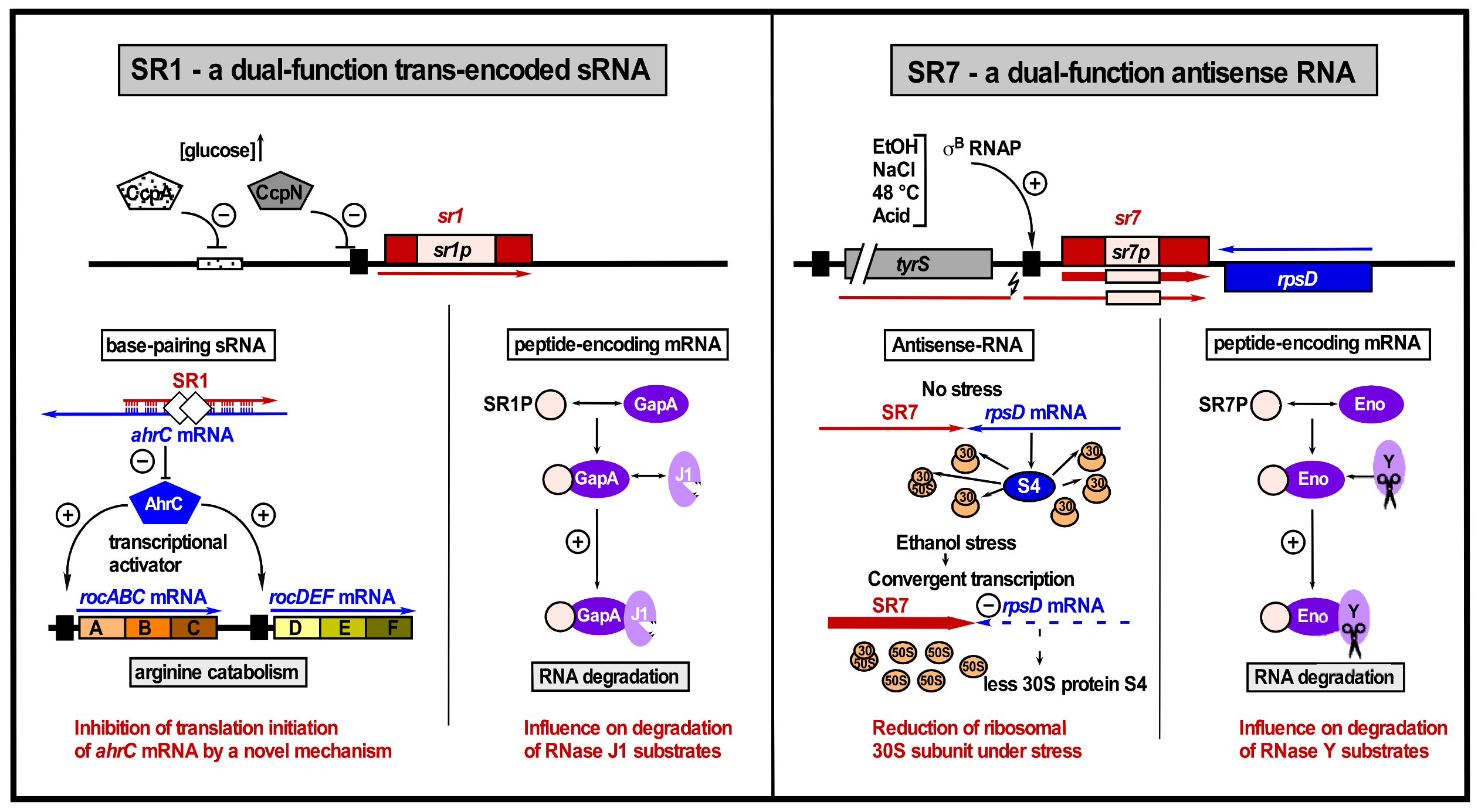
2.2.1. SR1
2.2.2. FsrA
2.2.3. RoxS
2.2.4. RosA
2.2.5. RnaC
3. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Brantl, S. Regulatory mechanisms employed by cis-encoded antisense RNAs. Curr. Opin. Microbiol. 2007, 10, 102–109. [Google Scholar] [CrossRef]
- Brantl, S. Acting antisense: Plasmid- and chromosome-encoded sRNAs from Gram-positive bacteria. Future Microbiol. 2012, 7, 853–871. [Google Scholar] [CrossRef]
- Wagner, E.G.; Romby, P. Small RNAs in bacteria and archaea: Who they are, what they do, and how they do it. Adv. Genet. 2015, 90, 133–208. [Google Scholar] [PubMed]
- Ul Haq, I.; Müller, P.; Brantl, S. Intermolecular communication in Bacillus subtilis: RNA-RNA, RNA-protein and small protein-protein interactions. Front. Mol. Biosci. 2020, 7, 178. [Google Scholar] [CrossRef] [PubMed]
- Pourciau, C.; Lai, Y.J.; Gorelik, M.; Babitzke, P.; Romeo, T. Diverse mechanisms and circuitry for global regulation by the RNA binding protein CsrA. Front. Microbiol. 2020, 11, 601352. [Google Scholar] [CrossRef] [PubMed]
- Chant, E.L.; Summers, D.K. Indole signalling contributes to the stable maintenance of Escherichia coli multicopy plasmids. Mol. Microbiol. 2007, 63, 35–43. [Google Scholar] [CrossRef]
- Brantl, S. Plasmid-replication control by antisense RNAs. Microbiol. Spec. 2014, 2, PLAS-0001-2013. [Google Scholar] [CrossRef] [Green Version]
- Brantl, S. Antisense RNAs in plasmids: Control of replication and maintenance. Plasmid 2002, 48, 165–173. [Google Scholar] [CrossRef]
- Argaman, L.; Herhsberg, R.; Vogel, J.; Bejerano, G.; Wagner, E.G.; Margalit, H.; Altuvia, S. Novel small RNA-encoding genes in the intergenic regions of Escherichia coli. Curr. Biol. 2001, 11, 941–950. [Google Scholar] [CrossRef] [Green Version]
- Wassarman, K.M.; Repoila, F.; Rosenow, C.; Storz, G.; Gottesman, S. Identification of novel small RNAs using comparative genomics and microarrays. Genes Dev. 2001, 15, 1637–1651. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adams, P.; Storz, G. Prevalence of small base-pairing RNAs derived from diverse genomic loci. Biochim. Biophys. Acta Gene Regul. Mech. 2020, 1863, 194524. [Google Scholar] [CrossRef]
- Georg, J.; Lalaouna, G.J.; Hou, S.; Lott, S.C.; Caldelari, I.; Marzi, S.; Hess, W.R.; Romby, P. The power of cooperation. Experimental and computational approaches in the functional characterization of bacterial sRNAs. Mol. Microbiol. 2020, 113, 603–612. [Google Scholar] [CrossRef]
- Rasmussen, S.; Nielsen, H.B.; Jarmer, H. The transcriptionally active regions in the genome of Bacillus subtilis. Mol. Microbiol. 2009, 73, 1043–1057. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Irnov, I.; Sharma, C.M.; Vogel, J.; Winkler, W.C. Identification of regulatory RNAs in Bacillus subtilis. Nucleic Acids Res. 2010, 38, 6637–6651. [Google Scholar] [CrossRef]
- Hör, J.; Matera, G.; Vogel, J.; Gottesman, S.; Storz, G. Trans-acting small RNAs and their effects on gene expression in Escherichia coli and Salmonella enterica. EcoSal Plus 2020, 9, 1–35. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brantl, S.; Müller, P. Toxin-antitoxin systems in Bacillus subtilis. Toxins 2019, 11, 262. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Durand, S.; Jahn, N.; Condon, C.; Brantl, S. Type I toxin-antitoxin systems in Bacillus subtilis. RNA Biol. 2012, 9, 1491–1497. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Silvaggi, J.M.; Perkins, J.B.; Losick, R. Small untranslated RNA antitoxin in Bacillus subtilis. J. Bacteriol. 2005, 187, 6641–6650. [Google Scholar] [CrossRef] [Green Version]
- Durand, S.; Gillet, L.; Condon, C. The essential function of B. subtilis RNase III is to silence foreign toxin genes. PLoS Genet. 2012, 8, e1003181. [Google Scholar] [CrossRef] [Green Version]
- Bloom-Ackermann, Z.; Steinberg, N.; Rosenberg, G.; Oppenheimer-Shaanan, Y.; Pollack, D.; Ely, S.; Storzi, N.; Levy, A.; Kolodkin-Gal, I. Toxin-Antitoxin systems eliminate defective cells and preserve symmetry in Bacillus subtilis biofilms. Environ. Microbiol. 2016, 18, 5032–5047. [Google Scholar] [CrossRef] [Green Version]
- Jahn, N.; Preis, H.; Wiedemann, C.; Brantl, S. BsrG/SR4 from Bacillus subtilis - the first temperature-dependent type I toxin-antitoxin system. Mol. Microbiol. 2012, 83, 579–598. [Google Scholar] [CrossRef]
- Jahn, N.; Brantl, S.; Strahl, H. Against the mainstream: The membrane-associated type I toxin BsrG from Bacillus subtilis interferes with cell envelope biosynthesis without increasing membrane permeability. Mol. Microbiol. 2015, 98, 651–666. [Google Scholar] [CrossRef] [PubMed]
- Jahn, N.; Brantl, S. One antitoxin—Two functions: SR4 controls toxin mRNA decay and translation. Nucleic Acids Res. 2013, 41, 9870–9880. [Google Scholar] [CrossRef] [PubMed]
- Heidrich, N.; Brantl, S. Antisense-RNA mediated transcriptional attenuation: Importance of a U-turn loop structure in the target RNA of plasmid pIP501 for efficient inhibition by the antisense RNA. J. Mol. Biol. 2003, 333, 917–929. [Google Scholar] [CrossRef]
- Jahn, N.; Brantl, S. Heat-shock-induced refolding entails rapid degradation of bsrG toxin mRNA by RNases Y and J1. Microbiology 2016, 162, 590–599. [Google Scholar] [CrossRef] [PubMed]
- Müller, P.; Jahn, N.; Ring, C.; Maiwald, C.; Neubert, R.; Meißner, C.; Brantl, S. A multistress responsive type I toxin-antitoxin system: bsrE/SR5 from the B. subtilis chromosome. RNA Biol. 2016, 13, 511–523. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meißner, C.; Jahn, N.; Brantl, S. In vitro characterization of the type I toxin-antitoxin system bsrE/SR5 from Bacillus subtilis. J. Biol. Chem. 2016, 291, 560–571. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reif, C.; Löser, C.; Brantl, S. Bacillus subtilis type I antitoxin SR6 promotes degradation of toxin yonT mRNA and is required to prevent toxic yoyJ overexpression. Toxins 2018, 10, 74. [Google Scholar] [CrossRef] [Green Version]
- Saito, S.; Kakeshita, H.; Nakamura, K. Novel small RNA-encoding genes in the intergenic regions of Bacillus subtilis. Gene 2009, 428, 2–8. [Google Scholar] [CrossRef]
- Eiamphungporn, W.; Helmann, J.D. Extracytoplasmic function sigma factors regulate expression of the Bacillus subtilis yabE gene via a cis-acting antisense RNA. J. Bacteriol. 2009, 191, 1101–1105. [Google Scholar] [CrossRef] [Green Version]
- Rao, F.; See, R.Y.; Zhang, D.; Toh, D.C.; Ji, Q.; Liang, Z.-X. YybT is a signaling protein that contains a cyclic dinucleotide phosphodiesterase domain and a GGDEF domain with ATPase activity. J. Biol. Chem. 2010, 285, 473–482. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, Y.; Helmann, J.D. A σD-dependent antisense-transcript modulates expression of the cyclic-di-AMP hydrolase GdpP in Bacillus subtilis. Microbiology 2012, 158, 2732–2741. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Noone, D.; Salzberg, L.I.; Botella, E.; Basell, K.; Becher, D.; Antelmann, H.; Devine, K.M. A highly unstable transcript makes CwlO D,L-endopeptidase expression responsive to growth conditions in Bacillus subtilis. J. Bacteriol. 2014, 196, 237–247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rath, H.; Reder, A.; Hoffmann, T.; Hammer, E.; Seubert, A.; Bremer, E.; Völker, U.; Mäder, U. Management of osmoprotectant uptake hierarchy in Bacillus subtilis via a SigB-dependent antisense RNA. Front. Microbiol. 2020, 11, 622. [Google Scholar] [CrossRef] [PubMed]
- Ul Haq, I.; Müller, P.; Brantl, S. SR7—A dual-function antisense RNA from Bacillus subtilis. RNA Biol. 2021, 18, 104–117. [Google Scholar] [CrossRef] [PubMed]
- Ul Haq, I.; Brantl, S. Moonlighting in Bacillus subtilis: The small proteins SR1P and SR7P regulate the moonlighting activity of glyceraldehyde 3-phosphate dehydrogenase A (GapA) and enolase in RNA degradation. Microorganisms 2021, 9, 1046. [Google Scholar] [CrossRef] [PubMed]
- Mars, R.A.; Mendonça, K.; Denham, E.L.; van Dijl, J.M. The reduction in small ribosomal subunit abundance in ethanol-stressed cells of Bacillus subtilis is mediated by a SigB-dependent antisense RNA. Biochim. Biophys. Acta 2015, 1853, 2553–2559. [Google Scholar] [CrossRef] [Green Version]
- Licht, A.; Preis, S.; Brantl, S. Implication of CcpN in the regulation of a novel untranslated RNA (SR1) in Bacillus subtilis. Mol. Microbiol. 2005, 58, 189–206. [Google Scholar] [CrossRef]
- Heidrich, N.; Chinali, A.; Gerth, U.; Brantl, S. The small untranslated RNA SR1 from the Bacillus subtilis genome is involved in the regulation of arginine catabolism. Mol. Microbiol. 2006, 62, 520–536. [Google Scholar] [CrossRef]
- Gaballa, A.; Antelmann, H.; Aguilar, C.; Khakh, S.K.; Song, K.B.; Smaldone, G.T.; Helmann, J.D. The Bacillus subtilis iron sparing response is mediated by a Fur regulated small RNA and three small, basic proteins. Proc. Natl. Acad. Sci. USA 2008, 105, 11927–11932. [Google Scholar] [CrossRef] [Green Version]
- Durand, S.; Braun, F.; Lioliou, E.; Romilly, C.; Helfer, A.C.; Kuhn, L.; Quittot, N.; Nicolas, P.; Romby, P.; Condon, C. A nitric oxide regulated small RNA controls expression of genes involved in redox homeostasis in Bacillus subtilis. PLoS Genet. 2015, 11, e1004957. [Google Scholar] [CrossRef] [Green Version]
- Durand, S.; Braun, F.; Helfer, A.C.; Romby, P.; Condon, C. sRNA-mediated activation of gene expression by inhibition of 5′-3′ exonucleolytic mRNA degradation. Elife 2017, 6, e23602. [Google Scholar] [CrossRef] [PubMed]
- Durand, S.; Callan-Sidat, A.; McKeown, J.; Li, S.; Kostova, G.; Hernandez-Fernaud, J.R.; Alam, M.T.; Millard, A.; Allouche, D.; Constantinidou, C.; et al. Identification of an RNA sponge that controls the RoxS riboregulatory of central metabolism in Bacillus subtilis. Nucleic Acids Res. 2021, 49, 6399–6419. [Google Scholar] [CrossRef]
- Mars, R.; Nicolas, P.; Ciccolini, M.; Rielman, E.; Reder, A.; Schaffer, M.; Mäder, U.; Völker, U.; van Dijl, J.M.; Denham, E.L. Small regulatory RNA-induced growth rate heterogeneity of Bacillus subtilis. PLoS Genet. 2015, 11, e1005046. [Google Scholar] [CrossRef] [Green Version]
- Heidrich, N.; Moll, I.; Brantl, S. In vitro analysis of the interaction between the small RNA SR1 and its primary target ahrC mRNA. Nucleic Acids Res. 2007, 35, 4331–4346. [Google Scholar] [CrossRef]
- Müller, P.; Gimpel, M.; Wildenhain, T.; Brantl, S. A new role for CsrA: Promotion of complex formation between an sRNA and its mRNA target in Bacillus subtilis. RNA Biol. 2019, 16, 972–987. [Google Scholar] [CrossRef] [Green Version]
- Licht, A.; Brantl, S. Transcriptional repressor CcpN from Bacillus subtilis compensates asymmetric contact distribution by cooperative binding. J. Mol. Biol. 2006, 364, 434–448. [Google Scholar] [CrossRef] [PubMed]
- Licht, A.; Golbik, R.; Brantl, S. Identification of ligands affecting the activity of the transcriptional repressor CcpN from Bacillus subtilis. J. Mol. Biol. 2008, 380, 17–30. [Google Scholar] [CrossRef]
- Licht, A.; Brantl, S. The transcriptional repressor CcpN from Bacillus subtilis uses different repression mechanisms at different promoters. J. Biol. Chem. 2009, 284, 30032–30038. [Google Scholar] [CrossRef] [Green Version]
- Czaplewski, L.G.; North, A.I.; Smith, M.C.; Baumberg, S.; Stockley, P.G. Purification and initial characterization of AhrC: The regulator of arginine metabolism genes in Bacillus subtilis. Mol. Microbiol. 1992, 6, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Gimpel, M.; Heidrich, N.; Mäder, U.; Krügel, H.; Brantl, S. A dual-function sRNA from B. subtilis: SR1 acts as a peptide encoding mRNA on the gapA operon. Mol. Microbiol. 2010, 76, 880–1009. [Google Scholar] [CrossRef] [Green Version]
- Gimpel, M.; Brantl, S. Dual-function sRNA encoded peptide SR1P modulates moonlighting activity of B. subtilis GapA. RNA Biol. 2016, 13, 916–926. [Google Scholar] [CrossRef] [Green Version]
- Gimpel, M.; Maiwald, C.; Wiedemann, C.; Görlach, M.; Brantl, S. Characterization of the interaction between the small RNA-encoded peptide SR1P and GapA from Bacillus subtilis. Microbiology 2017, 163, 1248–1259. [Google Scholar] [CrossRef]
- Gimpel, M.; Brantl, S. Dual-function small regulatory RNAs in bacteria. Mol. Microbiol. 2017, 103, 387–397. [Google Scholar] [CrossRef] [Green Version]
- Gimpel, M.; Preis, H.; Barth, E.; Gramzow, L.; Brantl, S. SR1—A small RNA with two remarkably conserved functions. Nucleic Acids Res. 2012, 40, 11659–11672. [Google Scholar] [CrossRef] [PubMed]
- Ul Haq, I.; Brantl, S.; Müller, P. A new role for SR1 from Bacillus subtilis - regulation of sporulation by inhibition of kinA translation. Nucleic Acids Res. 2021, in press. [Google Scholar] [CrossRef]
- Smaldone, G.T.; Revelles, O.; Gaballa, A.; Sauer, U.; Antelmann, H.; Helmann, J.D. A global investigation of the Bacillus subtilis iron-sparing response identifies major changes in metabolism. J. Bacteriol. 2012, 194, 2594–2605. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smaldone, G.T.; Antelmann, H.; Gaballa, A.; Helmann, J.D. The FsrA sRNA and FbpB protein mediate the iron-dependent induction of the Bacillus subtilis lutABC iron-sulfur-containing oxidases. J. Bacteriol. 2012, 194, 2586–2593. [Google Scholar] [CrossRef]
- Figueroa-Bossi, N.; Valentini, M.; Malleret, L.; Fiorini, F.; Bossi, L. Caught at its own game: Regulatory small RNA inactivated by an inducible transcript mimicking its target. Genes Dev. 2009, 23, 2004–2015. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miyakoshi, M.; Chao, Y.; Vogel, J. Cross talk between ABC transporter mRNAs via a target mRNA-derived sponge of the GcvB small RNA. EMBO J. 2015, 34, 1478–1492. [Google Scholar] [CrossRef]
- Bronesky, D.; Desgranges, E.; Corvaglia, A.; François, P.; Caballero, C.J.; Prado, L.; Toledo-Arana, A.; Lasa, I.; Moreau, K.; Vandenesch, F.; et al. A multifaceted small RNA modulates gene expression upon glucose limitation in Staphylococcus aureus. EMBO J. 2019, 38, e99363. [Google Scholar] [CrossRef] [PubMed]
- Li, G.W.; Oh, E.; Weissmann, J.S. The anti-Shine Dalgarno sequence drives translational pausing and codon choice in bacteria. Nature 2012, 484, 538–541. [Google Scholar] [CrossRef]
- Silvaggi, J.M.; Perkins, J.B.; Losick, R. Genes for small, noncoding RNAs under sporulation control in Bacillus subtilis. J. Bacteriol. 2006, 188, 532–541. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smirnov, A.; Schneider, C.; Hör, J.; Vogel, J. Discovery of new RNA classes and global RNA-binding proteins. Curr. Opin. Microbiol. 2017, 39, 152–160. [Google Scholar] [CrossRef]

| sRNA | sRNA Length | Mechanism of Action of sRNA | Target RNA | Target Gene Function | Regulation, Peculiarity |
|---|---|---|---|---|---|
| RatA | 222 nt | RD | txpA | Toxin | Glucose dependent |
| SR4 | 180 nt | RD + TI | bsrG | Toxin | Temperature dependent |
| SR5 | 163 nt | RD | bsrE | Toxin | Multistress responsive |
| SR6 | 100/215 nt * | RD TI | yonT yoyJ | Toxin Toxin | Multistress responsive |
| as-bsrH | 200 nt | RD | bsrH | Toxin | Multistress responsive |
| S25 | ≈1350 nt | unknown | yabE | Autolysin | sRNA under control of σM and σX |
| S1559 | 667 nt | unknown | gdpP | c-di-AMP PD | sRNA under σD control |
| S1326 | 700–2200 nt # | unknown | cwlO | Autolysin | sRNA under σB control |
| S1290 | 300–3800 nt # | T interference | opuB | Choline transporter | sRNA under σB control |
| SR7 | 185/259 nt | T interference | rpsD | Ribosomal protein S4 | sRNA under σB control; dual-function antisense RNA |
| sRNA | sRNA Length | Mechanism of Action of sRNA | Target RNA | Target Gene Function | Regulation |
|---|---|---|---|---|---|
| SR1 | 205 nt | TI TI | ahrC kinA | Arginine catabolism Sporulation initiation | CcpN, CcpA, sporulation |
| FsrA | 84 nt | TI TI TI TI TI TI | sdhCAB citB gltA lutABC dctP leuCD | Iron sparing response Aconitase Glutamate synthase Iron-sulfur oxidase Dicarboxylate permease Leucine biosynthesis | Fur, iron; Some targets need FbpA, B, or C |
| RnaC | 125 nt ? | RD + TI ? | abrB | Transition state regulation | Growth phase |
| RoxS | 115 nt | TI + RD TI + RD RD RS + TA | ppnKB sucC acsA yflS | Redox regulation, TCA cycle, Acetyl-CoA synthetase Malate transporter | ResD (NO), Rex (malate) |
| RosA | 92/128/ 193/225 nt * | Seq | RoxS FsrA | Trans-encoded sRNA Trans-encoded sRNA | CcpA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brantl, S.; Müller, P. Cis- and Trans-Encoded Small Regulatory RNAs in Bacillus subtilis. Microorganisms 2021, 9, 1865. https://doi.org/10.3390/microorganisms9091865
Brantl S, Müller P. Cis- and Trans-Encoded Small Regulatory RNAs in Bacillus subtilis. Microorganisms. 2021; 9(9):1865. https://doi.org/10.3390/microorganisms9091865
Chicago/Turabian StyleBrantl, Sabine, and Peter Müller. 2021. "Cis- and Trans-Encoded Small Regulatory RNAs in Bacillus subtilis" Microorganisms 9, no. 9: 1865. https://doi.org/10.3390/microorganisms9091865





