Comparative Transcriptomics Uncovers Upstream Factors Regulating BnFAD3 Expression and Affecting Linolenic Acid Biosynthesis in Yellow-Seeded Rapeseed (Brassica napus L.)
Abstract
:1. Introduction
2. Results
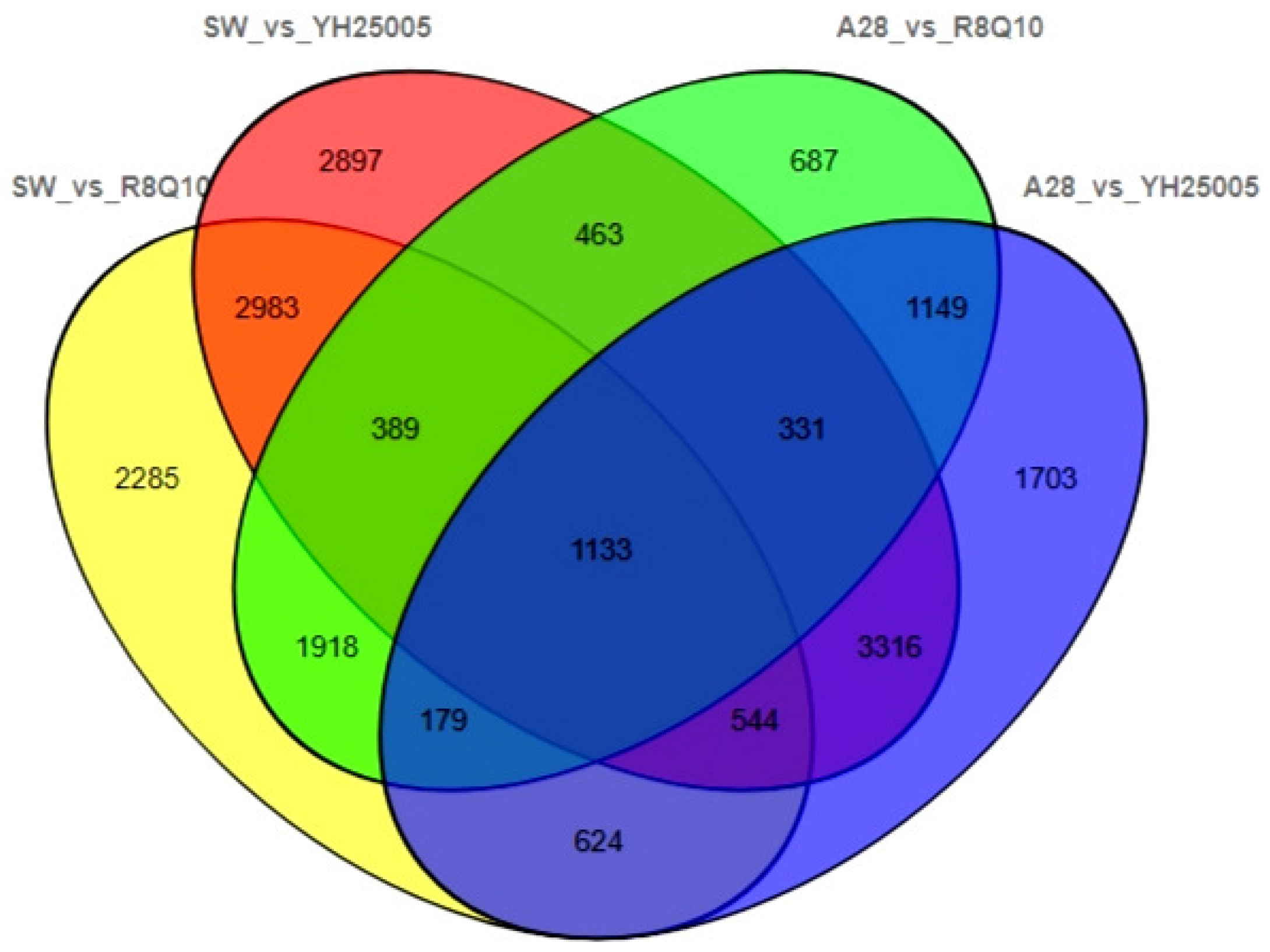
2.1. Screening Key Differentially Expressed Genes during Seed Development
2.2. KEGG Pathway Enrichment Analysis
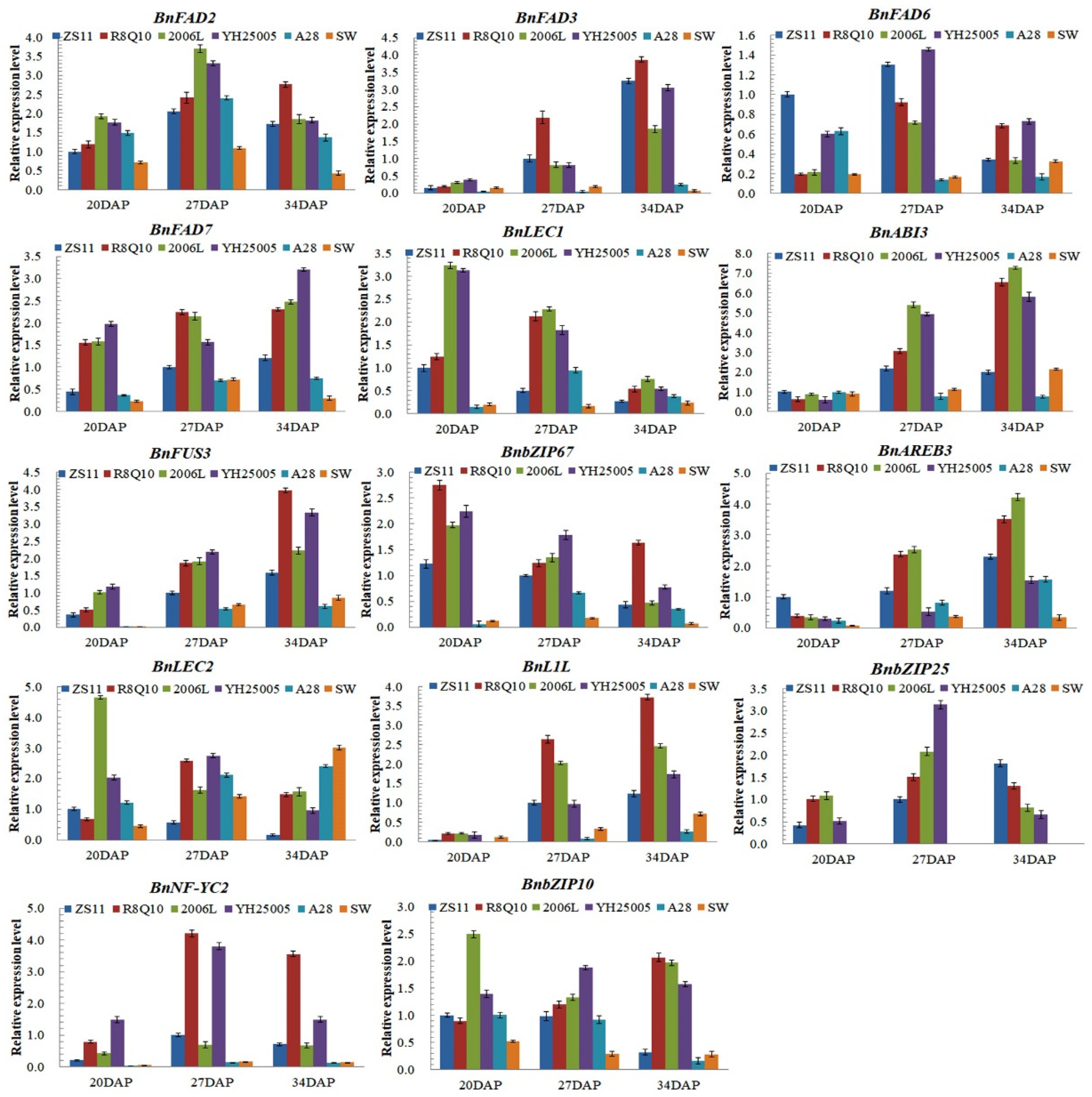
2.3. Expression Level of Rapeseed FAD Genes in Different Seed Development Stages
2.4. Differential Expression of TFs Related to ALA Synthesis
2.5. Expression Levels of Rapeseed Genes Related to Flavonoid Synthesis
3. Discussion
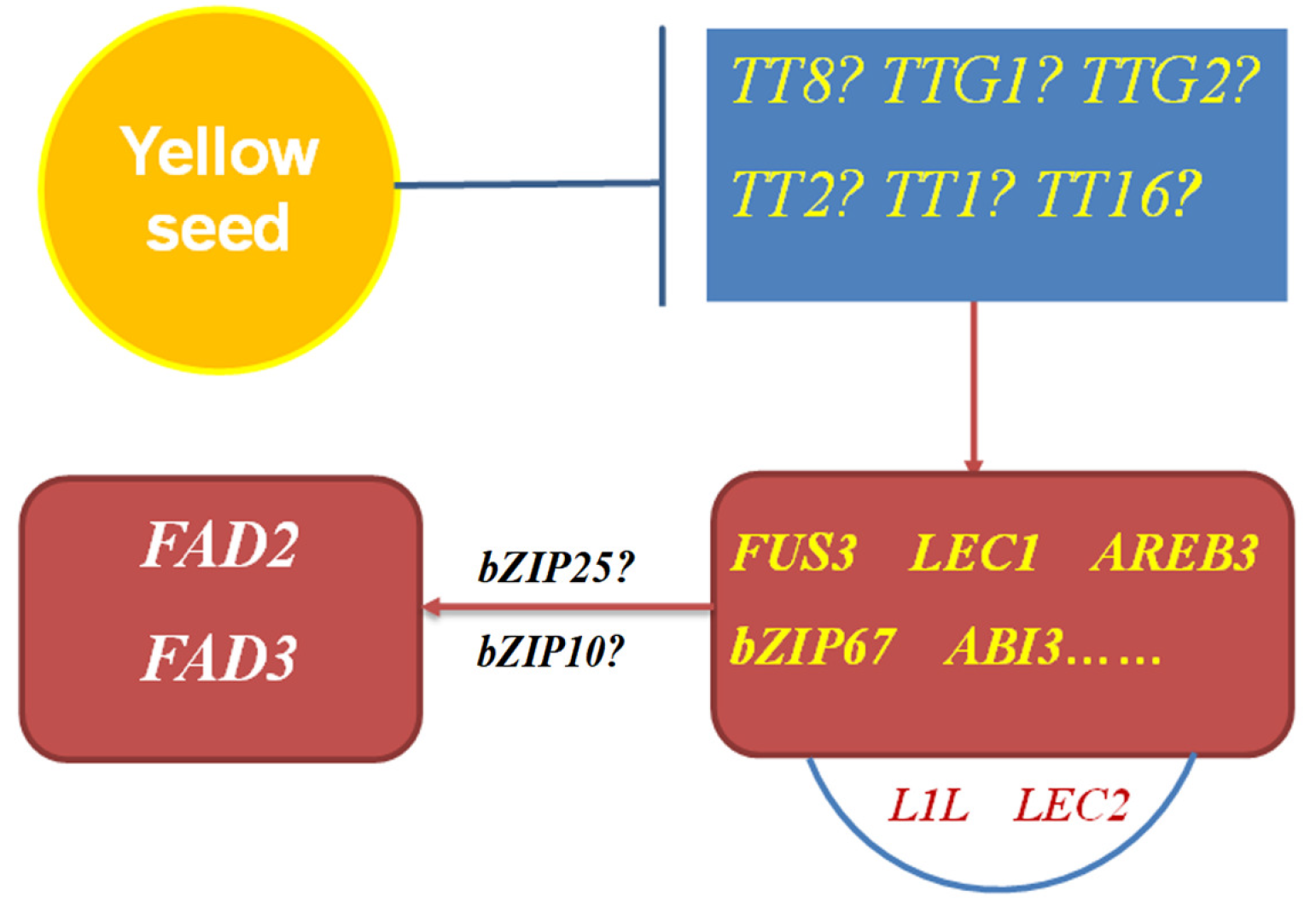
3.1. Upstream TFs Affect the Expression Levels of FAD Genes
3.2. Some tt or Yellow-Seededness Genes Can Regulate LAFL or LAZA TFs
3.3. Relationship between Yellow-Seededness and High-ALA Traits
4. Materials and Methods
4.1. Materials
4.2. Methods
4.2.1. Transcriptomic Comparison among R8Q10, YH25005, A28, and SW Seeds
4.2.2. Detection of Gene Expression Levels by Real-Time Quantitative PCR (RT-qPCR)
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Hu, X.Y.; Sullivan-Gilbert, M.; Gupta, M.; Thompson, S.A. G-to-A mutation at the 5′ splice site of fad3c caused impaired splicing in a low linolenic mutant of canola (Brassica napus L.). Plant Biotechnol. 2007, 24, 397–400. [Google Scholar] [CrossRef]
- Yang, Q.; Fan, C.; Guo, Z.; Qin, J.; Wu, J.; Li, Q.; Fu, T.; Zhou, Y. Identification of FAD2 and FAD3 genes in Brassica napus genome and development of allele-specific markers for high oleic and low-ALA contents. Theor. Appl. Genet. 2012, 125, 715–729. [Google Scholar] [CrossRef] [PubMed]
- Li, D.R.; Zhang, Y.W.; Zhang, W.X. A major breakthrough in the breeding of Brassica napus with high linolenic acid content, which linolenic acid content of germplasm resources exceeding 21% and of hybrids about 15%. J. Plant Sci. 2019, 3, 165–166. [Google Scholar]
- Zhang, X.H.; Lian, J.L.; Dai, C.Y.; Wang, X.L.; Zhang, M.Z.; Su, X.; Cheng, Y.H.; Yu, C.Y. Genetic segregation analysis of unsaturated fatty acids content in the filial generations of high-linolenic-acid rapeseed (Brassica napus). Oil Crop Sci. 2021, 6, 169–174. [Google Scholar] [CrossRef]
- Chen, Z.Z.; Zheng, H.G. High linolenic Acid Producing Brassica Plants. U.S. Patent US20160010096A1, 27 February 2020. [Google Scholar]
- Yu, C.Y. Molecular mechanism of manipulating seed coat coloration in oilseed Brassica species. J. Appl. Genet. 2013, 54, 135–145. [Google Scholar] [CrossRef]
- Zhao, Q.; Wu, J.; Cai, G.; Yang, Q.Y.; Shahid, M.; Fan, C.C.; Zhang, C.Y.; Zhou, Y.M. A novel quantitative trait locus on chromosome A9 controlling oleic acid content in Brassica napus. Plant Biotechnol. J. 2019, 7, 2313–2324. [Google Scholar] [CrossRef] [PubMed]
- Jo, L.; Pelletier, J.M.; Hsu, S.W.; Baden, R.; Goldberg, R.B.; Harada, J.J. Combinatorial interactions of the LEC1 transcription factor specify diverse developmental programs during soybean seed development. Proc. Natl. Acad. Sci. USA 2020, 117, 1223–1232. [Google Scholar] [CrossRef]
- Deng, W.; Chen, G.; Peng, F.; Truksa, M.; Snyder, C.L.; Weselake, R.J. Transparent testa16 plays multiple roles in plant development and is involved in lipid synthesis and embryo development in canola. Plant Physiol. 2012, 160, 978–989. [Google Scholar] [CrossRef]
- Wang, Z.; Chen, M.; Chen, T.; Xuan, L.; Li, Z.; Du, X.; Zhou, L.; Zhang, G.; Jiang, L. TRANSPARENT TESTA2 regulates embryonic fatty acid biosynthesis by targeting FUSCA3 during the early developmental stage of Arabidopsis seeds. Plant J. 2014, 77, 757–769. [Google Scholar] [CrossRef]
- Chen, M.X.; Xuan, L.J.; Wang, Z.; Zhou, L.H.; Li, Z.L.; Du, X.; Ali, E.; Zhang, G.P.; Jiang, L.X. TRANSPARENT TESTA8 inhibits seed fatty acid accumulation by targeting several seed development regulators in Arabidopsis. Plant Physiol. 2014, 165, 905–916. [Google Scholar] [CrossRef]
- Chen, M.X.; Zhang, B.; Li, C.X.; Kulaveerasingam, H.; Chew, F.T.; Yu, H. TRANSPARENT TESTA GLABRA1 regulates the accumulation of seed storage reserves in Arabidopsis. Plant Physiol. 2015, 169, 391–402. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.X.; Zhang, X.H.; Chen, X.Y.; Li, K.; Huang, Z.; Xu, A.X.; Dong, J.G.; Yu, C.Y. Landscape of sequence variations in homologous copies of FAD2 and FAD3 in rapeseed (Brassica napus L.) germplasm with high/low linolenic acid trait. Phyton-Int. J. Exp. Bot. 2024; accepted. [Google Scholar]
- Lian, J.; Lu, X.; Yin, N.; Ma, L.; Lu, J.; Liu, X.; Li, J.; Lu, J.; Lei, B.; Wang, R.; et al. Silencing of BnTT1 family genes affects seed flavonoid biosynthesis and alters seed fatty acid composition in Brassica napus. Plant Sci. 2017, 254, 32–47. [Google Scholar] [CrossRef]
- Xie, T.; Chen, X.; Guo, T.; Rong, H.; Chen, Z.; Sun, Q.; Batley, J.; Jiang, J.; Wang, Y. Targeted knockout of BnTT2 homologs for yellow-seeded Brassica napus with reduced flavonoids and improved fatty acid composition. J. Agric. Food Chem. 2020, 68, 5676–5690. [Google Scholar] [CrossRef]
- Mendes, A.; Kelly, A.A.; van Erp, H.; Shaw, E.; Powers, S.J.; Kurup, S.; Eastmond, P.J. bZIP67 regulates the omega-3 fatty acid content of Arabidopsis seed oil by activating fatty acid desaturase3. Plant Cell. 2013, 25, 3104–3116. [Google Scholar] [CrossRef]
- Elahi, N.; Duncan, R.W.; Stasolla, C. Decreased seed oil production in FUSCA3 Brassica napus mutant plants. Plant Physiol. Biochem. 2015, 96, 222–230. [Google Scholar] [CrossRef]
- Kim, M.J.; Kim, J.; Shin, J.S.; Suh, M.C. The SebHLH transcription factor mediates trans-activation of the SeFAD2 gene promoter through binding to E- and G-box elements. Plant Mol. Biol. 2007, 64, 453–466. [Google Scholar] [CrossRef] [PubMed]
- Tian, R.; Wang, F.; Zheng, Q.; Niza, V.M.A.G.E.; Downie, A.B.; Perry, S.E. Direct and indirect targets of the Arabidopsis seed transcription factor ABSCISIC ACID INSENSITIVE3. Plant J. 2020, 103, 1679–1694. [Google Scholar] [CrossRef]
- Hong, M.; Hu, K.; Tian, T.; Li, X.; Chen, L.; Zhang, Y.; Yi, B.; Wen, J.; Ma, C.; Shen, J.; et al. Transcriptomic analysis of seed coats in yellow-seeded Brassica napus reveals novel genes that influence proanthocyanidin biosynthesis. Front. Plant Sci. 2017, 8, 1674. [Google Scholar] [CrossRef]
- Liu, X.; Lu, Y.; Yan, M.; Sun, D.; Hu, X.; Liu, S.; Chen, S.; Guan, C.; Liu, Z. Genome-wide identification, localization, and expression analysis of proanthocyanidin-associated genes in Brassica. Front. Plant Sci. 2016, 7, 1831. [Google Scholar] [CrossRef] [PubMed]
- Rong, H.; Yang, W.; Xie, T.; Wang, Y.; Wang, X.; Jiang, J.; Wang, Y. Transcriptional profiling between yellow- and black-seeded Brassica napus reveals molecular modulations on flavonoid and fatty acid content. J. Integr. Agric. 2022, 21, 2211–2226. [Google Scholar] [CrossRef]
- Zhai, Y.; Yu, K.; Cai, S.; Hu, L.; Amoo, O.; Xu, L.; Yang, Y.; Ma, B.; Jiao, Y.; Zhang, C.; et al. Targeted mutagenesis of BnTT8 homologs controls yellow seed coat development for effective oil production in Brassica napus L. Plant Biotechnol. J. 2020, 18, 1153–1168. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.N.; Li, X.R.; Zhang, J.W.; Yu, C.Y. Genetic classification and diversity of yellow-seeded rapeseed (Brassica napus L.) accessions. Chin. Agric. Sci. Bull. 2015, 31, 156–164. [Google Scholar] [CrossRef]
- Xue, Y.F.; Alain, I.T.; Yin, N.; Jiang, J.; Zhao, Y.; Lu, K.; Li, J.; Ding, Y.; Zhang, S.; Chai, Y. Biotechnology of α-linolenic acid in oilseed rape (Brassica napus) using FAD2 and FAD3 from chia (Salvia hispanica). J. Integr. Agric. 2023, 22, 3810–3815. [Google Scholar] [CrossRef]
- Xue, Y.F.; Fu, C.; Chai, C.; Liao, F.; Chen, B.; Wei, S.; Wang, R.; Gao, H.; Fan, T.; Chai, Y. Engineering the staple oil crop Brassica napus enriched with α-linolenic acid using the perilla FAD2-FAD3 fusion gene. J. Agric. Food Chem. 2023, 71, 7324–7333. [Google Scholar] [CrossRef]
- Zhang, J.; Lu, Y.; Yuan, Y.; Zhang, X.; Geng, J.; Chen, Y.; Cloutier, S.; McVetty, P.B.; Li, G. Map-based cloning and characterization of a gene controlling hairiness and seed coat color traits in Brassica rapa. Plant Mol. Biol. 2009, 69, 553–563. [Google Scholar] [CrossRef]
- Li, X.; Chen, L.; Hong, M.; Zhang, Y.; Zu, F.; Wen, J.; Yi, B.; Ma, C.; Shen, J.; Tu, J.; et al. A large insertion in bHLH transcription factor BrTT8 resulting in yellow seed coat in Brassica rapa. PLoS ONE 2012, 7, e44145. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xiao, L.; Dun, X.; Liu, K.; Du, D. Characterization of the BrTT1 gene responsible for seed coat color formation in Dahuang (Brassica rapa L. landrace). Mol. Breed. 2017, 37, 137. [Google Scholar] [CrossRef]
- Zhang, Y.; Qin, Y.; Li, D.; Wang, W.; Gao, X.; Hao, C.; Feng, H.; Wang, Y.; Li, T. Fine mapping and cloning of a novel BrSCC1 gene for seed coat color in Brassica rapa. Theor. Appl. Genet. 2023, 136, 11. [Google Scholar] [CrossRef]
- Qu, C.; Zhu, M.; Hu, R.; Niu, Y.; Chen, S.; Zhao, H.; Li, C.; Wang, Z.; Yin, N.; Sun, F.; et al. Comparative genomic analyses reveal the genetic basis of the yellow-seed trait in Brassica napus. Nat. Commun. 2023, 14, 5194. [Google Scholar] [CrossRef]



| Comparison | DEGs | Upregulated Genes | Downregulated Genes | GO Database | KEGG Database | NR Database |
|---|---|---|---|---|---|---|
| A28 vs. R8Q10 | 6249 | 3165 | 3084 | 5136 | 2414 | 6113 |
| A28 vs. YH25005 | 8979 | 5438 | 3541 | 7535 | 3310 | 8780 |
| SW vs. R8Q10 | 10,055 | 4581 | 5474 | 8457 | 3971 | 9817 |
| SW vs. YH25005 | 12,056 | 6192 | 5864 | 10,154 | 4588 | 11,775 |
| R8Q10 vs. YH25005 | 11,587 | 4909 | 6678 | 10,010 | 4675 | 11,341 |
| R8Q10 Compared to SW | Log2(Foldchange) | R8Q10 Compared to A28 | Log2(Foldchange) |
|---|---|---|---|
| LEC1 NF-YB9 BnaAnng04140D | 1.467 | NF-YC3 | 1.19 |
| bZIP38, AREB4 ABI5-like protein 7 | Inf | bZIP25 BnaA06g16690D | Inf |
| AREB3 bZIP66 ABI5-like protein 2 | Inf | bZIP10 BnaC09g00090D | 3.16 |
| AREB3 bZIP66 ABI5-like protein 2 | 3.403 | bZIP46 BnaCnng20400D | 3.10 |
| bZIP67 ABI5-like protein 1, DPBF2 | 2.9 | BnaA.TT10b | 2.23 |
| bZIP25 | 2.358 | LEC1 BnaAnng04140D | −1.43 |
| Laccase-15-like precursor | 3.89 | TRANSPARENT TESTA 12 | −1.84 |
| Laccase type BnaA.TT10b | 3.083 | TRANSPARENT TESTA 12-like | −2.76 |
| bZIP46 BnaAnng26550D | −1.65 | TRANSPARENT TESTA 12-like DTX 27 | −Inf |
| TT8-like | −1.39 | BnaC.TT8 | −1.80 |
| BnaC.TT8 | −1.67 | TT8-like | −1.88 |
| TRANSPARENT TESTA 12 | −1.81 | AHA10 | −1.79 |
| YH25005 compared to SW | Log2(Foldchange) | YH25005 compared to A28 | Log2(Foldchange) |
| bZIP38, AREB4, ABI5-like protein 7 | Inf | ABI3, BnaC03g44820D | 1.60 |
| bZIP BnaCnng04010D | 3.648 | FUS3, BnaA02g28280D | 1.29 |
| bZIP25, BnaAnng30260D | 3.131 | FUS3, BnaC02g36350D | 1.36 |
| ABI5-like protein 3, bZIP12, DPBF4 | 2.655 | FUS3 | 1.64 |
| bZIP10, BnaA09g00980D | 1.962 | ABI3, BnaC03g44820D | 1.60 |
| bZIP35, AREB1 ABI5-like protein 4 | 1.808 | L1L, NF-YB6, BnaC02g33430D | 3.73 |
| ABI3, BnaC03g44820D | 1.033 | NF-YA9, BnaA05g19990D | 1.56 |
| TRANSPARENT TESTA 12 | −2.68 | bZIP12, DPBF4, ABI5-like protein 3 | 2.67 |
| TRANSPARENT TESTA 9 | −1.63 | bZIP25, BnaAnng30260D | 2.00 |
| TRANSPARENT TESTA 16 | −1.77 | TRANSPARENT TESTA 12 | −2.63 |
| TRANSPARENT TESTA 1 | −2.95 | TRANSPARENT TESTA 16 | −1.63 |
| Parameters | Seed Color | Average | Standard Error | p-Value of T-Test |
|---|---|---|---|---|
| Oil content % | Yellow | 45.32 | 3.263 | 0.005 |
| Brown | 43.41 | 3.756 | ||
| Oleic acid content % | Yellow | 64.54 | 5.87 | 0.000 |
| Brown | 66.86 | 6.112 | ||
| Linoleic acid content % | Yellow | 19.03 | 2.352 | 0.000 |
| Brown | 17.63 | 2.004 | ||
| Linolenic acid content % | Yellow | 13.24 | 2.271 | 0.000 |
| Brown | 10.98 | 1.514 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, X.-Y.; Wu, H.-X.; Zhang, X.-H.; Guo, R.-H.; Li, K.; Fu, Y.-L.; Huang, Z.; Xu, A.-X.; Dong, J.-G.; Yu, C.-Y. Comparative Transcriptomics Uncovers Upstream Factors Regulating BnFAD3 Expression and Affecting Linolenic Acid Biosynthesis in Yellow-Seeded Rapeseed (Brassica napus L.). Plants 2024, 13, 760. https://doi.org/10.3390/plants13060760
Chen X-Y, Wu H-X, Zhang X-H, Guo R-H, Li K, Fu Y-L, Huang Z, Xu A-X, Dong J-G, Yu C-Y. Comparative Transcriptomics Uncovers Upstream Factors Regulating BnFAD3 Expression and Affecting Linolenic Acid Biosynthesis in Yellow-Seeded Rapeseed (Brassica napus L.). Plants. 2024; 13(6):760. https://doi.org/10.3390/plants13060760
Chicago/Turabian StyleChen, Xiao-Yu, Hao-Xue Wu, Xiao-Han Zhang, Rong-Hao Guo, Kang Li, Yong-Li Fu, Zhen Huang, Ai-Xia Xu, Jun-Gang Dong, and Cheng-Yu Yu. 2024. "Comparative Transcriptomics Uncovers Upstream Factors Regulating BnFAD3 Expression and Affecting Linolenic Acid Biosynthesis in Yellow-Seeded Rapeseed (Brassica napus L.)" Plants 13, no. 6: 760. https://doi.org/10.3390/plants13060760






