Journal Description
Microorganisms
Microorganisms
is a scientific, peer-reviewed, open access journal of microbiology, published monthly online by MDPI. The Hellenic Society Mikrobiokosmos (MBK), the Spanish Society for Nitrogen Fixation (SEFIN) and the Society for Microbial Ecology and Disease (SOMED) are affiliated with Microorganisms, and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q1 (Microbiology (medical))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 20 days after submission; acceptance to publication is undertaken in 2.9 days (median values for papers published in this journal in the second half of 2025).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about Microorganisms.
- Companion journal for Microorganisms include: Applied Microbiology and Bacteria.
- Journal Cluster of Microbiology: Acta Microbiologica Hellenica, Applied Microbiology, Bacteria, Journal of Fungi, Microorganisms, Microbiology Research, Pathogens and Viruses.
Impact Factor:
4.2 (2024);
5-Year Impact Factor:
4.6 (2024)
Latest Articles
Isolation, Identification, Biological Characteristics, and In Vitro and In Vivo Antibacterial Effects of a Bovine-Derived Escherichia coli Bacteriophage XJA18
Microorganisms 2026, 14(5), 1118; https://doi.org/10.3390/microorganisms14051118 (registering DOI) - 14 May 2026
Abstract
To prevent the spread of antibiotic resistance, bacteriophages have gradually become the most promising alternative to antibiotics for treating bacterial infectious diseases. In this study, using E. coli DC1 as the host strain, we isolated a bacteriophage named Escherichia coli phage XJA18 from
[...] Read more.
To prevent the spread of antibiotic resistance, bacteriophages have gradually become the most promising alternative to antibiotics for treating bacterial infectious diseases. In this study, using E. coli DC1 as the host strain, we isolated a bacteriophage named Escherichia coli phage XJA18 from farm sewage. We conducted morphological identification, host range determination, biological characteristic analysis, genomic feature analysis, and evaluation of in vitro and in vivo antibacterial effects. Electron microscopy revealed that phage XJA18 belongs to the class Caudoviricetes, with an icosahedral head and a non-contractile long tail. Whole-genome sequencing revealed that the phage has dsDNA with a length of 50,572 bp, with a GC content of 45.33%. The genome does not contain any antibiotic resistance genes or virulence genes, indicating good safety. XJA18 showed lytic activity against 24% of clinically isolated E. coli strains. The optimal multiplicity of infection (MOI) was 0.001, with a latent period of 10 min, a burst period of 30 min, and a burst size of 2.22 × 102 PFU/cell. It remained stable at 4–50 °C and pH 4–12. In vitro antibacterial results revealed that XJA18 had the most pronounced initial bacterial growth suppression at MOI = 0.001 during the first 4 h. In vivo experiments demonstrated that both prophylactic and therapeutic administration of XJA18 could protect against E. coli infection, significantly reducing inflammatory cytokine levels and bacterial loads in the livers and spleens of mice (p < 0.001), significantly increasing body weight (p < 0.05), and reducing histopathological damage to the colon, liver, and lungs. In summary, phage XJA18 can effectively inhibit E. coli and is safe and stable. These characteristics indicate that phage XJA18 has great potential as a novel biological agent to replace antibiotics for treating bacterial infectious diarrhea in calves.
Full article
(This article belongs to the Section Antimicrobial Agents and Resistance)
►
Show Figures
Open AccessArticle
Epidemiological Study of the Relationship Between Antimicrobial Resistance Genes and Biofilm-Forming Capacity in Pathogens Causing Chronic Wound Infections
by
Silvia Ioana Musuroi, Adela Voinescu, Corina Musuroi, Delia Muntean, Florin George Horhat, Luminita Mirela Baditoiu, Oana Izmendi, Andrei Cosnita, Valentin Ordodi, Zorin Crainiceanu, Edward Seclaman and Monica Licker
Microorganisms 2026, 14(5), 1117; https://doi.org/10.3390/microorganisms14051117 (registering DOI) - 14 May 2026
Abstract
Chronic wounds represent a major complication of underlying conditions such as diabetes mellitus, arterial ischemia, surgical wound and burns. This study aimed at the phenotypic and molecular characterization of antimicrobial resistance for a selection of bacterial isolates, originating from wounds harvested from patients
[...] Read more.
Chronic wounds represent a major complication of underlying conditions such as diabetes mellitus, arterial ischemia, surgical wound and burns. This study aimed at the phenotypic and molecular characterization of antimicrobial resistance for a selection of bacterial isolates, originating from wounds harvested from patients hospitalized in the Vascular Surgery and Plastic Surgery wards. The microbiological diagnosis of wound infections was established according to the laboratory’s working protocol. PCR screening of antibiotic resistance genes was performed using a real-time PCR, while the microtiter plate assay was used to determine the biofilm-forming capacity. Testing of biofilm susceptibility to meropenem and amikacin was performed on Calgary biofilm device. Of the 88 bacterial isolates studied, 78.40% were Gram-negative bacilli (GNB)—Klebsiella pneumoniae (K.P), Pseudomonas aeruginosa (P.A), Proteus mirabilis (P.M), Acinetobacter baumannii (A.B), while the remaining 21.60% were Gram-positive cocci (GPC)—Staphylococcus aureus (S.A). All A.B isolates and 92.59% of K.P were carriers of β-lactamase- and carbapenemase-encoding genes, while 57.89% of S. aureus isolates were carriers of mecA (methicillin-resistant). Strong biofilm-forming isolates (B+++) were more frequent in P.A than in K.P (p = 0.002) and P.M (p = 0.02), with a frequency comparable to that of A.B strains (p = 0.212). When analyzing the biofilm reaction to meropenem, a significantly lower susceptibility was detected in the biofilm for K.P isolates, compared to the planktonic ones. Most GNB have been extensively multidrug-resistant, particularly K.P and A.B. Isolates from chronic wounds are major biofilm-formers. A strong and statistically significant association has been identified in the case of K.P and P.M between the presence of resistance genes and the biofilm-forming capacity. These findings highlight the need for a customized therapeutic approach for each chronic wound, considering the mechanisms underlying treatment resistance. These include bacterial virulence factors and the wound microenvironment colonized by the biofilm and the relative contribution of each to the overall resistance profile.
Full article
(This article belongs to the Special Issue Bacterial Pathogens: Biofilm Formation and Eradication)
►▼
Show Figures

Figure 1
Open AccessArticle
Senescent Eimeria acervulina Oocysts Maintain Transcriptional Activity During Extended Refrigerated Storage and Differentially Express Characteristic Genes
by
Matthew S. Tucker, Doaa Naguib, Celia N. O’Brien, Christina Yeager, Benjamin M. Rosenthal, Mark C. Jenkins and Asis Khan
Microorganisms 2026, 14(5), 1116; https://doi.org/10.3390/microorganisms14051116 (registering DOI) - 14 May 2026
Abstract
Enteric coccidian parasites harm agriculture and human health. Infectious, sporulated parasites eventually senesce. Here, we examined transcriptional changes in sporulated oocysts of Eimeria acervulina stored for 4–30 months at 4 °C. Precipitous decline in RNA abundance and transcription followed an interval of stability.
[...] Read more.
Enteric coccidian parasites harm agriculture and human health. Infectious, sporulated parasites eventually senesce. Here, we examined transcriptional changes in sporulated oocysts of Eimeria acervulina stored for 4–30 months at 4 °C. Precipitous decline in RNA abundance and transcription followed an interval of stability. Sixty constitutively expressed genes each contributed > 1000 transcripts per million (TPM) throughout, including a serine protease inhibitor, surface antigen genes, a cation-transporting ATPase, an oocyst wall protein, a zinc finger DHHC domain-containing protein, and highly expressed hypothetical proteins with no known function. Strikingly, ~82% of 6867 annotated genes underwent differential expression when comparing freshly sporulated parasites to those held for 30 months; nearly one-third of these underwent significant expression change. In freshly sporulated oocysts, 86 significantly DEGs exceeded 1000 TPM; these encoded heat shock proteins, lactate dehydrogenase, glucose-6 isomerase, and various hypothetical proteins. The oldest parasites expressed 66 DEGs, including many ribosomal subunits, a haloacid dehalogenase-like hydrolase domain-containing protein, and various hypothetical proteins. Taken together, these findings helped us to identify markers of mature parasites that remain relatively abundant in the transcript pool as oocysts age and identify other transcripts (e.g., ribosomal RNA) that increase in their relative abundance even as RNA abundance declines in senescent parasites.
Full article
(This article belongs to the Section Molecular Microbiology and Immunology)
►▼
Show Figures

Figure 1
Open AccessArticle
Assessment of the Impacts of Common Morel (Morchella sextelata) Cultivation on Soil Physicochemical Properties and Microbial Communities in Different Environments
by
Zhongyan Tang, Chen Chen, Li Dong, Liuyuan Bao, Chengcui Yang, Xiaodan Wang, Xiaoling Chen, Xiaokun Li, Fajun Xiang and Shunqiang Yang
Microorganisms 2026, 14(5), 1115; https://doi.org/10.3390/microorganisms14051115 (registering DOI) - 14 May 2026
Abstract
Morchella sextelata a species of high nutritional and economic value, is widely cultivated. To investigate how different cultivation environments affect the soil physicochemical properties and microbial communities associated with common morel, this study established cultivation plots under three distinct settings: apple orchard canopies,
[...] Read more.
Morchella sextelata a species of high nutritional and economic value, is widely cultivated. To investigate how different cultivation environments affect the soil physicochemical properties and microbial communities associated with common morel, this study established cultivation plots under three distinct settings: apple orchard canopies, dry upland fields, and paddy fields. The objective was to compare the differential impacts of common morel cultivation on soil environmental conditions across these habitats. The results indicate that cultivating common morel effectively enhances soil fertility. Across all environments, soil hydrolyzable nitrogen (HN), available potassium (AK), and organic matter content were higher than in the control. In apple orchard and dryland soils, total phosphorus (TP), total potassium (TK), available phosphorus (AP), and pH values were also elevated compared to the control, with most differences reaching significant levels. Solid Sucrase (S-SC) activity increased in all environments compared to the control, with values of 17.52 mg/d/g in PG, 17.39 mg/d/g in HD, and 21.68 mg/d/g in DT soils. Soil Amylase (S-AL) activity was higher in PG (451.28 μg/h/g) and HD (475.38 μg/h/g) soils. In contrast, Soil-acid phosphatase (S-ACP) activity was significantly elevated in DT soil (2922.08 nmol/h/g). PG soil exhibited significantly higher activities of Solid-Catalase (S-CAT), Solid polyphenol oxidase (S-PPO), and Solid Urease (S-UE), with S-CAT reaching 952.5 μmol/h/g. Following common morel cultivation, bacterial richness and diversity decreased across all conditions, while fungal richness increased but diversity declined. At the phylum level, Proteobacteria remained the dominant bacterial group, accounting for 26.78% in PG, 28.27% in HD, and 20.05% in DT soils. Ascomycota was the predominant fungal phylum, comprising 68.03% in PG, 72.16% in HD, and 68.94% in DT soils. Predicted bacterial functional pathways were primarily associated with metabolism, genetic information processing, environmental information processing, and cellular processes. Key metabolic pathways included carbohydrate metabolism, amino acid metabolism, and metabolism of cofactors and vitamins. fungal functional guilds were mainly classified as pathotrophic, pathotrophic–saprotrophic, pathotrophic–saprotrophic–symbiotrophic, and saprotrophic. Among these, saprotrophic and pathotrophic guilds showed higher abundance compared to the control. This shift is characterized by a reduction in both the diversity and abundance of beneficial microorganisms, alongside an increase in the richness of harmful microbial taxa. The combined effect of these factors disrupts the soil microbial equilibrium. The findings of this study provide a theoretical foundation for the cultivation of common morel and the management of associated soils.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures

Figure 1
Open AccessArticle
Green-Synthesized vs. Chemical Silver Nanoparticles: A Comparative Study on S. aureus Adaptability and Cross-Activity
by
Akamu Ewunkem, Josiah Dixon, Jordan Queenie, Uchenna Iloghalu, Franklin Ezeanowai and Sada Boyd
Microorganisms 2026, 14(5), 1114; https://doi.org/10.3390/microorganisms14051114 - 14 May 2026
Abstract
Rising antibiotic resistance necessitates alternatives such as silver nanoparticles (AgNPs), which exhibit bactericidal activity via multi-target mechanisms (e.g., membrane disruption, ROS production). While resistance to chemically synthesized AgNPs exists, the potential for resistance to green-synthesized AgNPs, such as those from reishi mushroom, is
[...] Read more.
Rising antibiotic resistance necessitates alternatives such as silver nanoparticles (AgNPs), which exhibit bactericidal activity via multi-target mechanisms (e.g., membrane disruption, ROS production). While resistance to chemically synthesized AgNPs exists, the potential for resistance to green-synthesized AgNPs, such as those from reishi mushroom, is unknown. This study compared S. aureus resistance development against both AgNP types using experimental evolution by analyzing genomic and morphological changes. Additionally, this work evaluated potential cross-resistance responses to ionic silver and investigated how adaptation to green-synthesized AgNPs affects sensitivity to chemically synthesized AgNPs (and vice versa). Rapid resistance, along with cross-resistance to silver ions, emerged in bacteria following 14 days of sublethal exposure to silver nanoparticles, regardless of whether they were chemically or biologically synthesized. While green-synthesized AgNPs demonstrated a substantial resistance to chemical variants (p < 0.05), the reverse effect was not as strong, and resistant populations showed distinct morphological adaptations. Genomic analysis highlighted convergent hard selective sweeps, identifying common mutations across both chemical and green AgNP-treated populations, with limited unique mutations found for either. These findings enhance our understanding of bacterial resistance mechanisms to nanomaterials, contributing to the development of safer, eco-friendly, and high-efficacy treatments against multidrug-resistant infection.
Full article
(This article belongs to the Special Issue Advances in Microbial Adaptation and Evolution)
►▼
Show Figures

Figure 1
Open AccessArticle
Debaryomyces hansenii Reshapes the Fungal Community of Iberian Cured Pork Loin: An ITS1 Metabarcoding Approach
by
Helena Chacón-Navarrete, Marina Barbudo-Lunar, Francisco Javier Ruiz-Castilla and José Ramos
Microorganisms 2026, 14(5), 1113; https://doi.org/10.3390/microorganisms14051113 - 14 May 2026
Abstract
Increasing consumer demand for natural and safe food products has led to the exploration of biocontrol alternatives to chemical preservatives, especially in the cured meat industry. The yeast Debaryomyces hansenii has emerged as a promising biocontrol candidate due to its antagonistic properties against
[...] Read more.
Increasing consumer demand for natural and safe food products has led to the exploration of biocontrol alternatives to chemical preservatives, especially in the cured meat industry. The yeast Debaryomyces hansenii has emerged as a promising biocontrol candidate due to its antagonistic properties against spoilage fungi. This study assessed the impact of D. hansenii inoculation on the fungal community structure of Iberian cured pork loin using high-throughput sequencing of the ITS1 region. Ion Torrent ITS1 amplicon sequencing, QIIME2/DADA2 pipeline, and ALDEx2 differential abundance analysis were applied to this study. Pork loin samples inoculated with D. hansenii were compared to non-inoculated controls to evaluate changes in the fungal microbiome. Inoculation resulted in a marked decrease in fungal diversity and evenness, indicating strong competition by D. hansenii against native fungal populations. This effect was reflected in a significant reduction in alpha diversity in inoculated samples (Shannon, p = 0.0042; Pielou p = 0.0075; Gini–Simpson, p = 0.0081). Notably, genera associated with spoilage and mycotoxin production, particularly Aspergillus and Penicillium, were significantly reduced in inoculated samples. Simultaneously, D. hansenii became dominant, reducing other yeasts and filamentous fungi. These findings highlight the powerful competitive and biocontrol potential of D. hansenii, demonstrating its ability to improve microbial safety by potentially reducing mycotoxin-associated risks through the suppression of toxigenic genera. This is the first study to characterise the fungal community of Iberian pork loin using metabarcoding under D. hansenii inoculation. The findings confirm that the inoculation of D. hansenii can substantially reduce fungal contamination risks. Overall, the results contribute valuable insights into microbial interactions during meat curing and underscore the practical benefits of targeted starter cultures for enhancing food safety and quality.
Full article
(This article belongs to the Special Issue Microbial Biotechnology in the Food Industry: Innovations, Applications, and Future Perspectives)
►▼
Show Figures

Figure 1
Open AccessReview
Next-Generation Vaccines Against Neglected Diseases: New Promises from Genetically Modified Live-Attenuated Parasites and RNA Vaccines
by
Marina Ferreira Batista-Zauli, Maria Eduarda Carvalho Guimarães Brasil, Carlos Roberto de Almeida-Júnior, Bárbara Germana Soares de Abreu, Nailma Silva Aprigio dos Santos, Mayra Fernanda Ricci and Santuza Maria Ribeiro Teixeira
Microorganisms 2026, 14(5), 1112; https://doi.org/10.3390/microorganisms14051112 - 14 May 2026
Abstract
Different protozoan parasites are the causative agents of tropical diseases, including malaria, toxoplasmosis, leishmaniasis, and Chagas disease (CD), which, altogether, affect over 300 million people throughout the world. Except for two recently approved malaria vaccines, individuals affected by or at risk of contracting
[...] Read more.
Different protozoan parasites are the causative agents of tropical diseases, including malaria, toxoplasmosis, leishmaniasis, and Chagas disease (CD), which, altogether, affect over 300 million people throughout the world. Except for two recently approved malaria vaccines, individuals affected by or at risk of contracting any of these four diseases still experience a lack of effective treatments and vaccines. Many vaccine studies, including those that have reached clinical trials, are based on inactivated parasites, adjuvanted recombinant proteins, or viral vector vaccines. Here, we review the current advances towards the development of vaccines based on genetically modified live-attenuated parasites (GMLAP) as well as RNA formulations encoding parasite antigens. Because these are diseases caused by intracellular pathogens that depend on efficient T-cell responses for parasite control, these two new vaccine platforms have generated great expectations, since they are known to induce a robust cellular immune response. Although preclinical studies aimed at developing new malaria, toxoplasmosis, and leishmaniasis vaccines have led to significant progress that may soon result in clinical trials, advances in next-generation vaccines against CD are lagging behind. Increased collaborative efforts between research groups, governments, and the pharmaceutical industry, particularly in Africa, Asia, and Latin American countries, are urgently needed to accelerate the development of vaccines for all neglected and less-studied diseases.
Full article
(This article belongs to the Special Issue Emerging Trends in Next-Generation Vaccines to Combat Infectious Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
Antifungal Resistance Patterns of Oral and Intestinal Candida Isolates Among People Living with HIV in a Tertiary Hospital in Gabon: A Cross-Sectional Study
by
Geril Sekangue Obili, Bridy Chelsy Moutombi Ditombi, Charlene Manomba Boulingui, Roger Hadry Sibi Matotou, Joyce Coëlla Mihindou, Dimitri Mabicka Moussavou, Denise Patricia Mawili Mboumba and Marielle Karine Bouyou-Akotet
Microorganisms 2026, 14(5), 1111; https://doi.org/10.3390/microorganisms14051111 (registering DOI) - 14 May 2026
Abstract
Digestive candidiasis is a major opportunistic infection among people living with HIV (PLHIV). In Gabon, data on antifungal resistance remain limited. This study aimed to characterise Candida colonisation and antifungal resistance according to anatomical site and species in Libreville. In this cross-sectional study,
[...] Read more.
Digestive candidiasis is a major opportunistic infection among people living with HIV (PLHIV). In Gabon, data on antifungal resistance remain limited. This study aimed to characterise Candida colonisation and antifungal resistance according to anatomical site and species in Libreville. In this cross-sectional study, 108 PLHIV provided paired oral and stool samples. Candida spp. was identified using conventional phenotypic methods. Antifungal susceptibility to azoles and polyenes was assessed by disc diffusion following CLSI guidelines. Resistance burden was classified by drug class and by cumulative number of antifungal agents involved. Digestive colonisation was detected in 97 (89.8%) participants. Oral and intestinal colonisation rates were 78.7% and 66.7%, respectively, with dual-site involvement in 55.6%. Among resistant isolates, Candida albicans accounted for 55.2% (oral) and 48.9% (intestinal), while non-albicans Candida represented 29.8% and 44.4%, respectively. Multidrug resistance was significantly higher in intestinal than oral isolates (36.2% vs. 11.8%; OR = 4.99; 95% CI: 2.04–12.16; p < 0.01). Resistance was predominantly azole-driven, with complex cumulative resistance profiles in intestinal isolates. The intestinal tract showed resistance profiles consistent with a preferential accumulation of MDR Candida populations in PLHIV. Site-specific resistance patterns underscore the importance of targeted sampling and antifungal stewardship strategies in resource-limited settings.
Full article
(This article belongs to the Section Antimicrobial Agents and Resistance)
►▼
Show Figures

Figure 1
Open AccessReview
Infectious Spondylodiscitis of Bacterial Causes in Adults: Epidemiology, Pathophysiology, Diagnostic and Treatment Challenges
by
Bogdan Sendrea, Argyrios Periferakis, Aristodemos-Theodoros Periferakis, Ioannis Xefteris, Lamprini Troumpata, Konstantinos Periferakis, Andreea-Elena Scheau, Emi Marinela Preda, Dana-Georgiana Nedelea, Diana-Elena Vulpe, Rares-Mircea Birlutiu, Cristian Scheau and Romica Cergan
Microorganisms 2026, 14(5), 1110; https://doi.org/10.3390/microorganisms14051110 - 13 May 2026
Abstract
Spinal infections in general, and infectious spondylodiscitis in particular, are increasingly diagnosed in the Western world, in recent decades. This rise in incidence is associated with an ageing population and with an increased availability of accurate diagnostic modalities. Even so, due to the
[...] Read more.
Spinal infections in general, and infectious spondylodiscitis in particular, are increasingly diagnosed in the Western world, in recent decades. This rise in incidence is associated with an ageing population and with an increased availability of accurate diagnostic modalities. Even so, due to the non-specific nature of clinical manifestations, and of the implicated blood and serum markers, there is a risk of underdiagnosis or misdiagnosis of the disease in its initial stages. Ionizing radiation methods, such as plain radiography (X-ray) and computed tomography (CT), are also not reliable in the early stages of the diseases, and the golden standard of imagistic diagnosis, magnetic resonance imaging (MRI), is not always available or requested. Still, MRI remains the most reliable method in most cases where there is a need for differential diagnosis with other pathologies, namely Andersson lesions, destructive spondyloarthropathy, erosive osteochondritis, micro-crystalline spondylitis, Modic 1 lesion, Charcot spinal arthropathy, osteoporotic fractures, SAPHO syndrome with spinal involvement, and Schmorl’s nodes. Infectious spondylodiscitis is caused by bacteria, and, less frequently, by fungi. Rare cases of parasitic causes have also been reported in the literature. Infectious spondylodiscitis of bacterial causes may be pyogenic, more frequently caused by Staphylococcus spp. or Streptococcus spp., or granulomatous, usually caused by Mycobacterium tuberculosis complex (MTBC) or from classical brucellosis. In all these cases, therapy may be conservative, with antibiotics, or surgical, when the former fails or in patients with significant spinal instability or other neurological manifestations. There are various surgical approaches, each with its own drawbacks, and usually used according to the preference of the attending physician. Even in cases of surgical treatment, antibiotic administration is prolonged, and it is important for a proper scheme to be selected based on antimicrobial susceptibility testing. However, given that in many cases, the causative agent cannot be identified, empirical treatment must be initiated. Finally, newer approaches, including the incorporation of antimicrobial substances, may offer better solutions for improving treatment and rehabilitation outcomes.
Full article
(This article belongs to the Special Issue The Epidemiology, Clinical Aspects, Treatment and Preventive Strategies for Infectious Diseases in the Era of Antimicrobial Resistance (AMR))
Open AccessReview
The Silent Spillover Threat: Nipah Virus Epidemiology, Pathogenesis, Clinical Manifestations, and Advances in Therapeutics and Vaccine Development
by
Elli-Panagiota Magklara, Maria Kkirgia, Andreas G. Tsantes, Petros Ioannou, Alexandra Mpakosi, Vasiliki Mougiou, Zoi Iliodromiti, Theodora Boutsikou, Nicoletta Iacovidou and Rozeta Sokou
Microorganisms 2026, 14(5), 1109; https://doi.org/10.3390/microorganisms14051109 - 13 May 2026
Abstract
Nipah virus (NiV) is an animal-borne RNA virus of the genus Henipavirus that poses a significant global health threat. This threat is driven by the virus’s high mortality rate, its capacity to cause epidemics, and the lack of licensed therapeutic interventions or vaccines.
[...] Read more.
Nipah virus (NiV) is an animal-borne RNA virus of the genus Henipavirus that poses a significant global health threat. This threat is driven by the virus’s high mortality rate, its capacity to cause epidemics, and the lack of licensed therapeutic interventions or vaccines. Since its initial identification during the 1998–1999 outbreak in Malaysia and Singapore, recurrent episodes have occurred primarily in Bangladesh and India, with mortality rates frequently exceeding 70%. Fruit bats of the genus Pteropus serve as the biological host for the virus. Transmission to humans occurs via contact with infected wildlife, consumption of contaminated products, such as freshly harvested date palm sap, or direct person-to-person exposure. Other modes of transmission, such as transplacentally or via breast milk, are still under investigation. The clinical presentation of NiV infection varies widely, from mild flu-like symptoms to life-threatening respiratory disease and acute encephalitis. It frequently attacks the nervous system, which can lead to coma, permanent neurological damage, or relapsing encephalitis. The virus enters host cells via ephrin-B2/B3 receptors, enabling systemic dissemination and infiltration of the central nervous system. Diagnosis relies primarily on RT-PCR and serological assays, and virus isolation requires high-containment laboratories. Management remains largely supportive, as no approved antiviral therapy exists. Experimental agents, such as remdesivir, favipiravir, and monoclonal antibodies such as m102.4, have shown promise in preclinical studies. Multiple vaccine platforms—including subunit, viral vector, mRNA, and nanoparticle-based approaches—are under development, though none is yet licensed for human use. Strengthened surveillance, infection control measures, and continued research are essential to mitigate the threat posed by this emerging pathogen. This review summarizes current knowledge on NiV, including its virology, epidemiology, pathogenesis, transmission, and recent progress in therapeutic and vaccine development.
Full article
(This article belongs to the Section Virology)
Open AccessArticle
Fluoroquinolone-Induced Metabolic Dysregulation and Oxidative Stress Orchestrate Bacterial Demise
by
Caiyuan Zhou, Jing Sun, Yihan Luo, Fang Wang, Luqi Li, Tong Wu, Peng Xie, Chenxi Liu, Yibin Hu, Leilei Sun and Chengbao Wang
Microorganisms 2026, 14(5), 1108; https://doi.org/10.3390/microorganisms14051108 - 13 May 2026
Abstract
The bactericidal mechanisms of fluoroquinolones extend beyond their canonical inhibition of DNA topoisomerases, yet the associated metabolic perturbations remain incompletely understood. In this study, we systematically investigated the metabolic responses of Escherichia coli to three representative FQs—ofloxacin, enrofloxacin, and ciprofloxacin—using untargeted UPLC–Q Exactive
[...] Read more.
The bactericidal mechanisms of fluoroquinolones extend beyond their canonical inhibition of DNA topoisomerases, yet the associated metabolic perturbations remain incompletely understood. In this study, we systematically investigated the metabolic responses of Escherichia coli to three representative FQs—ofloxacin, enrofloxacin, and ciprofloxacin—using untargeted UPLC–Q Exactive Orbitrap–MS-based metabolomics. Bacterial cells were exposed to bactericidal concentrations (2 × MIC) for a single-time point (1 h), followed by comprehensive metabolomic profiling with six biological replicates per group. Our findings demonstrate that FQ-induced metabolic reprogramming serves as a primary driver of oxidative stress and nucleic acid damage, rather than a mere secondary effect. All three FQs induced substantial metabolic reprogramming characterized by disruptions in nucleotide biosynthesis, central carbon metabolism, and redox-related pathways, with notable drug-specific differences. Ciprofloxacin exhibited the most pronounced suppression of energy metabolism and antioxidant systems, whereas ofloxacin and enrofloxacin showed partial compensatory metabolic responses. Consistently, intracellular ROS levels were significantly elevated in all treatment groups, and this effect was attenuated by antioxidant supplementation. Furthermore, increased accumulation of 8-hydroxydeoxyguanosine and 8-hydroxyguanosine confirmed the occurrence of oxidative DNA and RNA damage. Collectively, these findings indicate that FQs induce distinct metabolic perturbations that are closely associated with oxidative stress and nucleic acid damage, providing a metabolic perspective on their bactericidal activity and suggesting potential targets for metabolic adjuvant strategies.
Full article
(This article belongs to the Section Molecular Microbiology and Immunology)
►▼
Show Figures
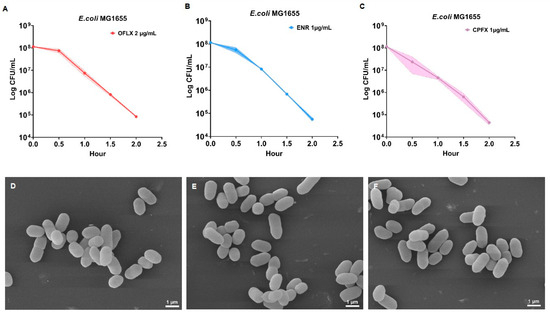
Figure 1
Open AccessArticle
Mechanism and Control of Black Spot Deterioration on Lacquered Architectural Components of Dajue Temple
by
Sifan Ai, Yu Wang, Jiao Pan, Gang Hu and Ruiting Zhao
Microorganisms 2026, 14(5), 1107; https://doi.org/10.3390/microorganisms14051107 - 13 May 2026
Abstract
Dajue Temple, a representative ancient architectural heritage in North China, houses numerous lacquered wooden components of exceptional historical and artistic value. Prolonged environmental exposure causes severe dark discoloration and black spotting on lacquer surfaces, threatening their structural integrity. This first investigation into the
[...] Read more.
Dajue Temple, a representative ancient architectural heritage in North China, houses numerous lacquered wooden components of exceptional historical and artistic value. Prolonged environmental exposure causes severe dark discoloration and black spotting on lacquer surfaces, threatening their structural integrity. This first investigation into the damage identifies the spots as microbial in origin, with Cladosporium spp. as the primary agent driving deterioration and possessing wood-degrading capabilities. Antifungal tests show that thymol, clove essential oil, and nano-silver gel are all effective inhibitors. We proposed targeted, relic-friendly microbial control strategies tailored for ancient lacquered wooden components. These findings provided scientific guidance for the sustainable conservation and restoration of lacquered architectural elements in historic temples and comparable cultural heritage sites. In future work, environmental monitoring and the biocides’ compatibility should be involved, which will help to clarify microbe–environment interactions, enable early warning of biodeterioration risks and explore the wood-friendly biocides.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures
Figure 1
Open AccessArticle
Scalp Microbiota Dysbiosis in Seborrheic Alopecia and Restoration Following Herbal Extract Shampoo Intervention
by
Jing Feng, Jiancong Huang, Shaolu Zhou, Xianghai Chen, Gang Zhou, Xudong Wang, Xia Wen, Qingshan Shi, Pianjuan Guo, Qiongfei Li and Xiaobao Xie
Microorganisms 2026, 14(5), 1106; https://doi.org/10.3390/microorganisms14051106 - 13 May 2026
Abstract
Seborrheic alopecia (SA) is one of the most common forms of hair loss with a complex pathogenesis involving multiple etiological factors. Although the scalp microbiome has been implicated in various scalp disorders, its specific role in the development and progression of SA remains
[...] Read more.
Seborrheic alopecia (SA) is one of the most common forms of hair loss with a complex pathogenesis involving multiple etiological factors. Although the scalp microbiome has been implicated in various scalp disorders, its specific role in the development and progression of SA remains incompletely understood. To characterize the scalp microbiome in SA, we performed high-throughput sequencing of the 16S rRNA gene and ITS region on scalp samples from 41 Chinese SA participants before and after a 12-week intervention with a shampoo containing herbal extracts (ginger root, Polygonum multiflorum, and Platycladus orientalis leaf) and 29 healthy controls. The untreated SA group exhibited significant microbial dysbiosis compared to the healthy controls, characterized by reduced bacterial and fungal alpha diversity and increased relative abundances of Staphylococcus, Cutibacterium, and Malassezia. LEfSe analysis confirmed the significant enrichment of these three genera. Correlation network analysis revealed a substantial restructuring of microbial interactions in the untreated SA group: Staphylococcus and Malassezia lost all positive correlations with other genera, whereas Cutibacterium displayed relatively stable topological relationships. Following the 12-week intervention, the treated SA group showed significant clinical improvement (reduced hair loss and scalp sebum content), along with a restoration of microbial diversity to levels comparable to the healthy group and a normalization of the abundances of Staphylococcus and Malassezia. Our study confirms the critical role of scalp microecological dysbiosis in SA pathogenesis and identifies Staphylococcus and Malassezia as key taxa strongly associated with this dysbiosis. These findings provide a theoretical foundation for developing microbiome-targeted strategies for SA treatment and support the use of multi-targeted, plant-based interventions to restore microbial homeostasis and promote hair growth.
Full article
(This article belongs to the Section Microbiomes)
►▼
Show Figures
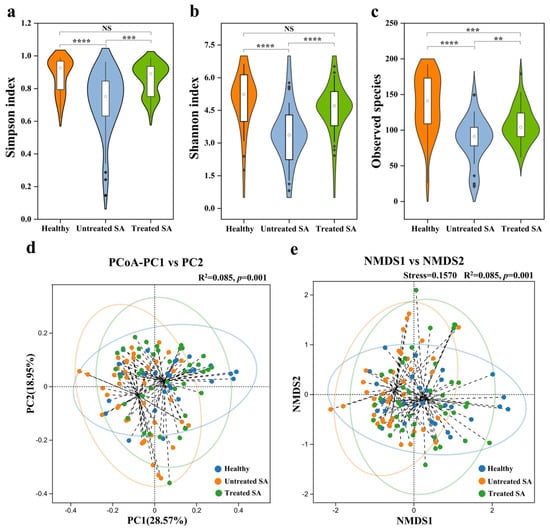
Figure 1
Open AccessArticle
Early-Life Iron Exposure Influences Long-Term Gut Microbiota Recovery After Intestinal Dysbiosis
by
Thibault Maumy, Claire McCartney, Ayodeji Samuel Ajayi, Claire Gerkins, Gabriela Fragoso, Annie Calvé and Manuela M. Santos
Microorganisms 2026, 14(5), 1105; https://doi.org/10.3390/microorganisms14051105 - 13 May 2026
Abstract
Host–microbiota interactions during the neonatal window of opportunity have gained significant interest as factors influencing long-term health. Factors such as nutrient availability may shape the microbial community, and iron is no exception to this rule. Although the use of iron supplementation is widespread
[...] Read more.
Host–microbiota interactions during the neonatal window of opportunity have gained significant interest as factors influencing long-term health. Factors such as nutrient availability may shape the microbial community, and iron is no exception to this rule. Although the use of iron supplementation is widespread during infancy, this micronutrient is known to have profound effects on gut microbiota. This study aimed to determine how early-life iron supplementation shapes gut microbiota composition and whether it influences recovery from gut dysbiosis later in life. Three-week-old female C57BL/6 mice were fed an iron-excess diet for five weeks during the critical period of microbiota establishment. After a two-week washout period to normalize luminal iron content, dysbiosis was induced using either dextran sulfate sodium-induced acute colitis or antibiotic treatment. Mice were then allowed an 8-week recovery period. Gut microbiota composition was longitudinally analyzed by 16S rRNA gene sequencing of fecal samples. Early-life iron supplementation induced durable alterations in gut microbiota composition, with differences persisting even after luminal iron normalization (β-diversity, PERMANOVA p < 0.01). At the endpoint, mice exposed to an iron-sufficient diet remained significantly more distant from their baseline compared to the excess iron group in both the dextran sulfate sodium (33% greater distance) and antibiotic (41% greater distance) models (both p < 0.05). Notably, this pattern was not observed when supplementation occurred in adulthood. In the dextran sulfate sodium model, mice that received an iron-sufficient diet during early life showed an expansion of the probiotic Ligilactobacillus murinus, which positively correlated with fecal succinate levels. Conversely, in the antibiotic-induced model, early-life exposure to an iron-sufficient diet was associated with a more pronounced dysbiosis characterized by a nearly two-fold-greater loss in α-diversity compared to 500 ppm mice (∆Shannon: 1.98 ± 0.22 vs. 1.02 ± 0.25, p < 0.01). These findings suggest that early-life iron supplementation influences long-term host–microbiota interactions and recovery from gut dysbiosis.
Full article
(This article belongs to the Special Issue Effects of Diet and Nutrition on Gut Microbiota)
►▼
Show Figures

Figure 1
Open AccessReview
Tobacco-Induced Oral Dysbiosis and Microbial Shifts: A Narrative Review of Their Role in Systemic Inflammation and Disease
by
Glenda M. Davison, Tandi Matsha, Shanel Raghubeer, Stanton Hector, Saarah Davids and Yvonne Prince
Microorganisms 2026, 14(5), 1104; https://doi.org/10.3390/microorganisms14051104 - 13 May 2026
Abstract
The oral cavity is home to a diverse community of microbiota comprising bacteria, viruses, protozoa, and fungi. These microorganisms inhabit several oral niches and play a significant role in supporting both oral and systemic health. The fine balance between the microbial communities can
[...] Read more.
The oral cavity is home to a diverse community of microbiota comprising bacteria, viruses, protozoa, and fungi. These microorganisms inhabit several oral niches and play a significant role in supporting both oral and systemic health. The fine balance between the microbial communities can be influenced by genetics and environmental factors, potentially leading to dysbiosis. Alterations in the oral microbiota have been implicated in periodontitis, chronic inflammation, and systemic disease. Tobacco has been identified as a major player in altering the oral microenvironment and disturbing the balance between potentially pathogenic and beneficial commensals. The resulting dysbiosis promotes inflammation and assists in the passage of pathogenic microorganisms into the blood system. This narrative review examines current evidence linking the use of tobacco with the dominance of pathogenic oral bacteria and a dysfunctional immune response. We explore how the chemicals and toxins in cigarettes promote a reduction in oxygen and cause changes in the abundance of anaerobic bacteria. After discussing the mechanistic pathways leading to periodontitis and the entry of microorganisms into the circulation, the review will interrogate previous studies and identify opportunities and priorities for future research.
Full article
(This article belongs to the Special Issue Microbiomes in Human Health and Diseases)
►▼
Show Figures
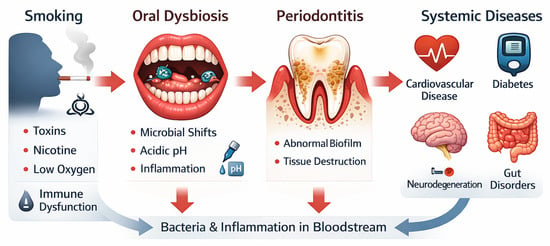
Graphical abstract
Open AccessArticle
Isolation, Identification, and Functional Characterization of a Rhizosphere Bacterium Promoting the Growth of Alsophila spinulosa
by
Jiya Wu, Weicheng Yang, Xiaona Zhang, Xianyu Li, Bibo Zhou, Tianyu Liang and Fen Liu
Microorganisms 2026, 14(5), 1103; https://doi.org/10.3390/microorganisms14051103 - 13 May 2026
Abstract
Alsophila spinulosa is a tree fern designated as a second-class nationally protected species in China and valued for its medicinal and ornamental properties. Its slow growth and susceptibility to environmental stresses pose challenges to its cultivation. Plant-growth-promoting rhizobacteria (PGPR) can enhance plant development
[...] Read more.
Alsophila spinulosa is a tree fern designated as a second-class nationally protected species in China and valued for its medicinal and ornamental properties. Its slow growth and susceptibility to environmental stresses pose challenges to its cultivation. Plant-growth-promoting rhizobacteria (PGPR) can enhance plant development by producing phytohormones, such as indole-3-acetic acid (IAA). In this study, 39 IAA-producing strains were isolated from the rhizosphere of A. spinulosa. Morphological and molecular analyses identified the highest IAA-producing strain, R74, as Burkholderia pyrrocinia. Its optimal inoculum age was determined to be 12–20 h, and its optimal culture conditions for IAA production were 24 h of incubation, 32 °C and pH 7.0. Whole-genome sequencing revealed that the genome of strain R74 is 8,347,169 bp in size with a GC content of 67%, comprising 7543 genetic elements. Further genomic analysis showed that IAA biosynthesis in R74 involves the tryptophan side-chain oxidase (TSO) pathway and the tryptophan-independent pathway. Pot experiments revealed that inoculation with R74 increased the height, root length, stem diameter, and biomass of A. spinulosa seedlings. It also increased antioxidant enzyme activities, elevated soluble protein and chlorophyll contents, and reduced malondialdehyde levels. This study provides an empirical basis for the development of Burkholderia-based biofertilizers to promote A. spinulosa growth.
Full article
(This article belongs to the Section Plant Microbe Interactions)
►▼
Show Figures
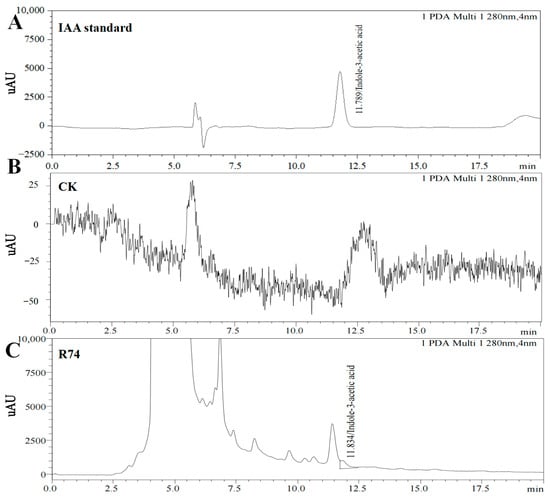
Figure 1
Open AccessArticle
Mutation-Tolerant Inhibition of HIV-1 Integrase Strand Transfer by Secondary Metabolites from the Endophytic Fungus Alternaria alternata PO4PR2
by
Ndzalo Mashabela, Darian Naidu, Ernest Oduro-Kwateng and Nompumelelo P. Mkhwanazi
Microorganisms 2026, 14(5), 1102; https://doi.org/10.3390/microorganisms14051102 - 13 May 2026
Abstract
Endophytic fungi are promising sources of novel antiviral compounds, and the crude extract from Alternaria alternata PO4PR2 has previously shown anti-HIV-1 activity. This study evaluated its efficacy against integrase strand-transfer inhibitor (INSTI)-resistant HIV-1 and its mechanism of action. Key resistance mutations (Y143H, G118R,
[...] Read more.
Endophytic fungi are promising sources of novel antiviral compounds, and the crude extract from Alternaria alternata PO4PR2 has previously shown anti-HIV-1 activity. This study evaluated its efficacy against integrase strand-transfer inhibitor (INSTI)-resistant HIV-1 and its mechanism of action. Key resistance mutations (Y143H, G118R, N155H, and R263K) were introduced into the HIV-1 pNL4.3 clone via site-directed mutagenesis and confirmed through Sanger sequencing. Viral infectivity was assessed in TZM-bl cells, while cytotoxicity was measured using an MTT assay. Antiviral activity was determined through a luciferase-based assay, and integration inhibition was evaluated using integrase activity assays and Alu-gag nested PCR. The extract demonstrated potent inhibition of resistant mutants, with low IC50 values (0.02971–0.1652 μg/mL), and showed minimal cytotoxicity (CC50 = 300 μg/mL), maintaining over 80% cell viability. It inhibited integrase activity by 67%, specifically targeting the strand-transfer step, and significantly reduced integrated viral DNA. Molecular docking of 14 compounds identified coumarin derivatives as key bioactive metabolites, exhibiting mutation-tolerant binding within the integrase catalytic pocket. Overall, these findings highlight PO4PR2 as a promising source of compounds for developing new therapies targeting drug-resistant HIV-1 integrase.
Full article
(This article belongs to the Section Virology)
►▼
Show Figures
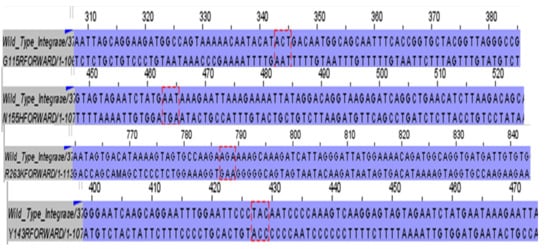
Figure 1
Open AccessArticle
From the Carp Gut to Plastic Solutions: Hafnia Strain from Cyprinus carpio Demonstrates Robust Degradation of Synthetic Polymers
by
Mina Popovic, Boris Rajcic and Neveka Rajic
Microorganisms 2026, 14(5), 1101; https://doi.org/10.3390/microorganisms14051101 (registering DOI) - 13 May 2026
Abstract
The accumulation of polyethylene (PE) in aquatic ecosystems represents a significant environmental challenge due to the polymer’s high molecular weight and chemical stability. This study investigates the biodegradation potential of Hafnia paralvei UUNT_MP29, a bacterial strain isolated from the gut of common carp
[...] Read more.
The accumulation of polyethylene (PE) in aquatic ecosystems represents a significant environmental challenge due to the polymer’s high molecular weight and chemical stability. This study investigates the biodegradation potential of Hafnia paralvei UUNT_MP29, a bacterial strain isolated from the gut of common carp (Cyprinus carpio), for low-density polyethylene (LDPE). Initial screening on LDPE-emulsified agar confirmed extracellular enzymatic activity through the formation of distinct clear zones. Quantitative analysis showed a cumulative mass loss of 24.10% by Day 16, with the most intensive degradation occurring between Days 4 and 8, which closely correlated with maximum bacterial count (CFU/mL). Kinetic modeling indicated that the degradation followed a first-order rate law (R2 = 0.9269), with a rate constant (k) of 0.2991 days−1 and a remarkably short half-life (t1/2) of 2.32 days. Structural characterization via FTIR spectroscopy demonstrated oxidative transformation, evidenced by a reduction in sp3 C-H stretching and the emergence of C-O/C-O-C functional groups. SEM micrographs further confirmed extensive bio-deterioration, including surface pitting and macroscale erosion. Thermal analysis (TGA/DTG) supported these findings, showing a significant 10.95 °C decrease in the maximum degradation temperature (Tmax), indicating a reduction in polymer chain length. These results suggest that H. paralvei UUNT_MP29 is a highly efficient agent for the rapid breakdown of polyethylene and highlight the potential of aquatic gut microbiota as reservoirs for plastic-degrading biotechnologies.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures

Figure 1
Open AccessEditorial
Editorial for the Special Issue Research on Biology of Dinoflagellates
by
Tania Islas-Flores, Estefanía Morales-Ruiz and Marco A. Villanueva
Microorganisms 2026, 14(5), 1100; https://doi.org/10.3390/microorganisms14051100 - 13 May 2026
Abstract
Protists arose about 1 [...]
Full article
(This article belongs to the Special Issue Research on Biology of Dinoflagellates)
Open AccessArticle
Effect of Aeration Rate Redistribution on Nitrogen Removal Performance of a Novel Multi-Compartment Fixed-Biofilm Cyclic Activated Sludge System
by
Zichun Yan, Shuichao Fan, Wankai Yan, Haopeng Ma and Tianhao Zhao
Microorganisms 2026, 14(5), 1099; https://doi.org/10.3390/microorganisms14051099 - 13 May 2026
Abstract
To address the problems of short-circuit flow and dead zones, complicated operation and control caused by intermittent influent, and the mismatch between aeration rate and oxygen demand in the Cyclic Activated Sludge System (CASS), a novel Multi-Compartment Fixed-Biofilm Cyclic Activated Sludge System (MCFCASS)
[...] Read more.
To address the problems of short-circuit flow and dead zones, complicated operation and control caused by intermittent influent, and the mismatch between aeration rate and oxygen demand in the Cyclic Activated Sludge System (CASS), a novel Multi-Compartment Fixed-Biofilm Cyclic Activated Sludge System (MCFCASS) was developed. This system operated in continuous-flow mode, and the aeration rate of each compartment was redistributed using a mathematical model. The results show that the plug flow ratio of the MCFCASS reactor increased from 18.75% to 31.25% compared with the CASS reactor. After aeration rate redistribution, the average total nitrogen (TN) removal efficiency of the MCFCASS reactor rose from 83.34% to 86.80%, and the effluent TN concentration consistently met the Grade I-A limit (15 mg/L) specified in the Discharge Standard of Pollutants for Municipal Wastewater Treatment Plant (GB 18918-2002). The average removal efficiencies of chemical oxygen demand (COD) and ammonium nitrogen (NH4+-N) increased from 91.58% and 93.39% to 92.98% and 94.57%, respectively. Microbial community analysis revealed that after aeration rate redistribution, the relative abundances of Pseudomonadota, Bacteroidota, and Bacillota in the pre-reaction zone of MCFCASS were 39.17%, 17.78%, and 10.33%, respectively. In addition, the abundances of some functional bacterial groups in the first and fourth compartments of the main reaction zone shifted adaptively in response to the aeration rate redistribution, consistent with the trends in pollutant removal contributions in these compartments. Hierarchical clustering and principal coordinate analysis (PCoA) further indicated that aeration rate redistribution influenced the microbial community structure. The above laboratory-scale optimization results may provide a preliminary reference for aeration control and improvement of denitrification performance in similar processes.
Full article
(This article belongs to the Collection Feature Papers in Environmental Microbiology)
►▼
Show Figures
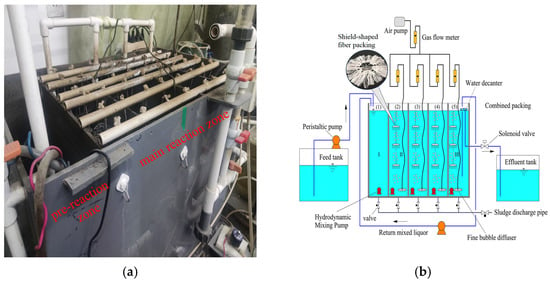
Figure 1

Journal Menu
► ▼ Journal Menu-
- Microorganisms Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Applied Biosciences, Applied Microbiology, Fermentation, Marine Drugs, Microorganisms
Microbial Cell Factories for Natural Products
Topic Editors: Carlos Barreiro, Ana Ibáñez, José L. BarredoDeadline: 31 May 2026
Topic in
Energies, Geosciences, Hydrology, Plants, Microorganisms, Sustainability, Agriculture
Transport, Transformation and Cycling of Elements in Water and Soil and Their Response to Human Activities
Topic Editors: Jiutan Liu, Junfu DongDeadline: 30 June 2026
Topic in
Dairy, Foods, Microorganisms, Nutrients, Metabolites
The Efficacy of Probiotics and Their Functional Metabolites in Fermented Foods
Topic Editors: Rina Wu, Wenjun Liu, Zhen Wu, Feiyu AnDeadline: 31 July 2026
Topic in
Microorganisms, JCM, Biomedicines, Oral, Clinics and Practice
Periodontal and Peri-Implant Disease: Contemporary Perspectives on Pathogenesis and Precision Management
Topic Editors: Giuseppe Troiano, Vito Carlo Alberto Caponio, Mario DioguardiDeadline: 31 August 2026

Conferences
Special Issues
Special Issue in
Microorganisms
Pathobiology, Infection Biology and Control of Protozoan Parasites—the ONE HEALTH Approach
Guest Editors: Cláudia Marques, Maria João Gouveia, Susana R. SousaDeadline: 15 May 2026
Special Issue in
Microorganisms
Role of Soil Microorganisms in Agricultural Production and Environmental Biotechnologies
Guest Editor: Gianluigi CardinaliDeadline: 20 May 2026
Special Issue in
Microorganisms
Borreliosis and Other Tick-Borne Diseases in the Northern and Southern Hemispheres, Second Edition
Guest Editor: Giusto TrevisanDeadline: 30 May 2026
Special Issue in
Microorganisms
Phytoplankton Faced with Emergent Pollutants
Guest Editors: Katia Comte, Stéphanie FayolleDeadline: 31 May 2026
Topical Collections
Topical Collection in
Microorganisms
Microbial Life in Extreme Environments
Collection Editor: Ricardo Amils
Topical Collection in
Microorganisms
Trends in Yeast Biochemistry and Biotechnology
Collection Editors: Seiji Shibasaki, Mitsuyoshi Ueda
Topical Collection in
Microorganisms
Epidemiology and Pathogenicity of Animal-Adapted Streptococci
Collection Editors: Peter Valentin-Weigand, Marcus Fulde










