Transcriptomic Analysis Reveals Transcription Factors Related to Leaf Anthocyanin Biosynthesis in Paeonia qiui
Abstract
:1. Introduction
2. Results
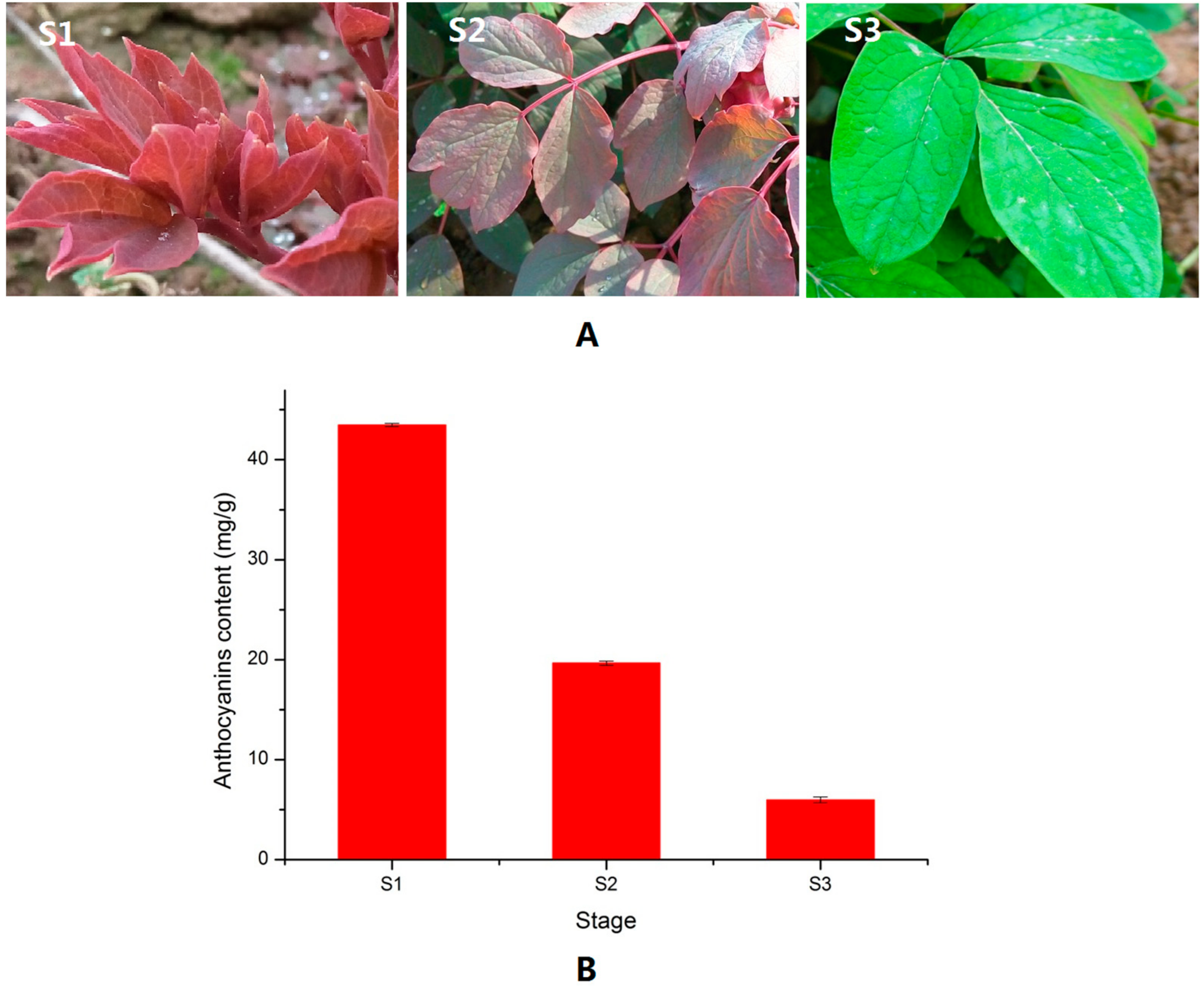
2.1. Anthocyanin Level in the Different Leaf Stages of P. qiui
2.2. Library Construction, De Novo Assembly, and Gene Annotation
2.3. Identification of DEGs Involved in Flavonoid/Anthocyanin Biosynthesis
2.4. Identification of MYBs Involved in Anthocyanin Biosynthesis
2.5. Identification bHLHs Involved in Anthocyanin Biosynthesis
2.6. Identification WD40 Involved in Anthocyanin Biosynthesis
2.7. Quantitative RT-PCR Analysis of the DEGs Involved in Anthocythanin Biosynthesis
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Anthocyanin Level Measurement
4.3. RNA Extraction, Library Construction, De Novo Assembly, and Gene Annotation
4.4. Identification and Analysis of DEGs
4.5. Phylogentic Tree and Sequence Alignment
4.6. Quantitative RT-PCR (q-PCR) Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ji, L.; Wang, Q.; Teixeira da Silva, J.A.; Yu, X.N. The genetic diversity of Paeonia L. Sci. Hortic. 2012, 143, 62–74. [Google Scholar] [CrossRef]
- Grotewold, E. The genetics and biochemistry of floral pigments. Annu. Rev. Plant Biol. 2006, 57, 761–780. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Sasaki, N.; Ohmiya, A. Biosynthesis of plant pigments: Anthocyanin, betalains and carotenoid. Plant J. 2008, 54, 733–749. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.Q.; Wei, M.R.; Liu, D.; Tao, J. Anatomical and biochemical analysis reveal the role of anthocyanin in flower coloration of herbaceous peony. Plant Physiol. Biochem. 2016, 102, 97–106. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Chiu, L.W.; Li, L. Transcriptional regulation of anthocyanin biosynthesis in red cabbage. Planta 2009, 230, 1141–1153. [Google Scholar] [CrossRef] [PubMed]
- Petroni, K.; Tonelli, C. Recent advances on the regulation of anthocyanin synthesis in reproductive organs. Plant Sci. 2011, 181, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Jaakola, L. New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant Sci. 2013, 18, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.S.; Shiraishi, A.; Hashimoto, F.; Aoki, N.; Shimizu, K.; Sakata, Y. Analysis of petal anthocyanin to investigate flower coloration of Zhongyuan (Chinese) and Daikon Island (Japanese) tree peony cultivars. J. Plant Res. 2001, 114, 33–43. [Google Scholar] [CrossRef]
- Wang, L.S.; Hashimoto, F.; Shiraishi, A.; Aoki, N.; Li, J.J.; Sakata, Y. Chemical taxonomy of the Xibei tree peony from China by floral pigmentation. J. Plant Res. 2004, 117, 47–55. [Google Scholar] [PubMed]
- Zhang, J.; Wang, L.; Shu, Q.; Liu, Z.A.; Li, C.; Zhang, J.; Wei, X.; Tian, D. Comparison of anthocyanin in non-blotches and blotches of the petals of Xibei tree peony. Sci. Hortic. 2007, 114, 104–111. [Google Scholar] [CrossRef]
- Zhang, C.; Wang, W.; Wang, Y.; Gao, S.; Du, D.; Fu, J.; Dong, L. Anthocyanin biosynthesis and accumulation in developing flowers of tree peony (Paeonia suffruticosa) ‘Luoyang Hong’. Postharvest Biol. Technol. 2014, 97, 11–22. [Google Scholar] [CrossRef]
- Zhang, Y.; Cheng, Y.; Ya, H.; Xu, S.; Han, J. Transcriptome sequencing of purple petal spot region in tree peony reveals differentially expressed anthocyanin structural genes. Front. Plant Sci. 2015, 6, 964. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Wang, Y.; Ren, L.; Shi, Q.; Zheng, B.; Miao, K.; Guo, X. Overexpression of Ps-CHI1, a homologue of the chalcone isomerase gene from tree peony (Paeonia suffruticosa), reduces the intensity of flower pigmentation in transgenic tobacco. Plant Cell Tissue Organ Cult. 2013, 116, 285–295. [Google Scholar] [CrossRef]
- Shi, Q.; Zhou, L.; Wang, Y.; Li, K.; Zheng, B.; Miao, K. Transcriptomic analysis of paeonia delavayi wild population flowers to identify differentially expressed genes involved in purple-red and yellow petal pigmentation. PLoS ONE 2015, 10, e0135038. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.D.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Erika, C.; José, T.M.; Laura, F.; Sara, Z.; Rodrigo, L.; Flavia, G.; Rudolf, S.; Agnès, A.; Patricio, A.J.; Giovanni, B.T. The Phenylpropanoid Pathway Is Controlled at Different Branches by a Set of R2R3-MYB C2 Repressors in Grapevine. Plant Physiol. 2015, 167, 1448–1470. [Google Scholar]
- Wang, L.K.; Bolitho, K.; Grafton, K.; Kortstee, A.; Karunairetnam, A.; McGhie, T.K.; Espley, R.V.; Hellens, R.P.; Allan, A.C. An R2R3 MYB transcription factor associated with regulation of the anthocyanin biosynthetic pathway in Rosaceae. BMC Plant Biol. 2010, 10, 50. [Google Scholar]
- Zimmermann, I.M.; Heim, M.A.; Weisshaar, B.; Uhrig, J.F. Comprehensive identification of Arabidopsis thaliana MYB transcription factors interacting with R/B-like BHLH proteins. Plant J. 2004, 40, 22–34. [Google Scholar] [CrossRef] [PubMed]
- Kagale, S.; Rozwadowski, K. EAR motif-mediated transcriptional repression in plants: An underlying mechanism for epigenetic regulation of gene expression. Epigenetics 2011, 6, 141–146. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.R.; Shi, Q.Q.; Niu, L.X.; Zhang, Y.L. Transcriptomic analysis of leaf in tree peony reveals differentially expressed pigments genes. Molecules 2017, 22, 324. [Google Scholar] [CrossRef] [PubMed]
- Heim, M.A.; Jakoby, M.; Werber, M.; Martin, C.; Weisshaar, B.; Bailey, P.C. The basic helix–loop–helix transcription factor family in plants: A genome-wide study of protein structure and functional diversity. Mol. Biol. Evol. 2003, 20, 735–747. [Google Scholar] [CrossRef] [PubMed]
- Shi, Q.Q.; Li, L.; Zhang, X.X.; Luo, J.R.; Li, X.; Zhai, L.J.; He, L.X.; Zhang, Y.L. Biochemical and comparative transcriptomic analyses identify candidate genes related to variegation formation in Paeonia rockii. Molecules 2017, 22, 1364. [Google Scholar] [CrossRef] [PubMed]
- Hichri, I.; Barrieu, F.; Bogs, J.; Kappel, C.; Delrot, S.; Lauvergeat, V. Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J. Exp. Bot. 2011, 62, 2465–2483. [Google Scholar] [CrossRef] [PubMed]
- Schwinn, K.; Venail, J.; Shang, Y.; Mackay, S.; Alm, V.; Butelli, E.; Oyama, R.; Bailey, P.; Davies, K.; Martin, C. A small family of MYB-regulatory genes controls floral pigmentation intensity and patterning in the genus Antirrhinum. Plant Cell 2006, 18, 831–851. [Google Scholar] [CrossRef] [PubMed]
- Chiu, L.W.; Zhou, X.; Burke, S.; Wu, X.; Prior, R.L.; Li, L. The purple cauliflower arises from activation of a MYB transcription factor. Plant Physiol. 2010, 154, 1470–1480. [Google Scholar] [CrossRef] [PubMed]
- Albert, N.W.; Lewis, D.H.; Zhang, H.; Schwinn, K.E.; Jameson, P.E.; Davies, K.M. Members of an R2R3-MYB transcription factor family in Petunia are developmentally and environmentally regulated to control complex floral and vegetative pigmentation patterning. Plant J. 2011, 65, 771–784. [Google Scholar] [CrossRef] [PubMed]
- Yamagishi, M.; Toda, S.; Tasaki, K. The novel allele of the LhMYB12 gene is involved in splatter-type spot formation on the flower tepals of Asiatic hybrid lilies (Lilium spp.). New Phytol. 2014, 201, 1009–1020. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.W.; Sagawa, J.M.; Frost, L.; Vela, J.P.; Bradshaw, H.D., Jr. Transcriptional control of floral anthocyanin pigmentation in monkey flowers (Mimulus). New Phytol. 2014, 204, 1013–1027. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.C.; Chen, Y.Y.; Tsai, W.C.; Chen, W.H.; Chen, H.H. Three R2R3-MYB transcription factors regulate distinct floral pigmentation patterning in Phalaenopsis spp. Plant Physiol. 2015, 168, 175–191. [Google Scholar] [CrossRef] [PubMed]
- Takos, A.M.; Jaffe, F.W.; Jacob, S.R.; Bogs, J.; Robinson, S.P.; Walker, A.R. Light-induced expression of a MYB gene regulates anthocyanin biosynthesis in red apples. Plant Physiol. 2006, 142, 1216–1232. [Google Scholar] [CrossRef] [PubMed]
- Dubos, C.; Gourrierec, J.; Baudry, A.; Huep, G.; Lanet, E.; Debeaujon, I.; Routaboul, J.M.; Alboresi, A.; Weisshaar, B.; Lepiniec, L. MYBL2 is a new regulator of flavonoid biosynthesis in Arabidopsis thaliana. Plant J. 2008, 55, 940–953. [Google Scholar] [CrossRef] [PubMed]
- Schaart, J.G.; Dubos, C.; Romero, D.L.F.I.; Houwelingen, A.M.; de Vos, R.C.; Jonker, H.H.; Xu, W.; Routaboul, J.M.; Lepiniec, L.; Bovy, A.G. Identification and characterization of MYB-bHLH-WD40 regulatory complexes controlling proanthocyanidin biosynthesis in strawberry (Fragaria ananassa) fruits. New Phytol. 2013, 197, 454–467. [Google Scholar] [CrossRef] [PubMed]
- Albert, N.W.; Davies, K.M.; Lewis, D.H.; Zhang, H.; Montefiori, M.; Brendolise, C.; Boase, M.R.; Ngo, H.; Jameson, P.E.; Schwinn, K.E. A conserved network of transcriptional activators and repressors regulates anthocyanin pigmentation in eudicots. Plant Cell 2014, 26, 962–980. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.K.; Micheletti, D.; Palmer, J.; Volz, R.; Lozano, L.; Espley, R.; Hellens, R.P.; Chagnè, D.; Rowan, D.D.; Troggio, M.; et al. High temperature reduces apple fruit colour via modulation of the anthocyanin regulatory complex. Plant Cell Environ. 2011, 34, 1176–1190. [Google Scholar] [CrossRef] [PubMed]
- Salvatierra, A.; Pimentel, P.; Moya-León, M.A.; Herrera, R. Increased accumulation of anthocyanins in Fragaria chiloensis fruits by transient suppression of FcMYB1 gene. Phytochemistry 2013, 90, 25–36. [Google Scholar] [CrossRef] [PubMed]
- Mortazavi, A.; Williams, B.A.; McCue, K.; Schaeffer, L.; Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are available from the authors. |








| Gene ID | log2 Ratio (S3/S1) | FDR | Annotation |
|---|---|---|---|
| Unigene0055753 | −10.26229064 | 1.81 × 10−25 | Chalcone synthase (Petunia x hybrida) |
| Unigene0017118 | −1.778044372 | 4.66 × 10−40 | Chalcone synthase (Paeonia suffruticosa) |
| Unigene0053168 | 10.49914828 | 2.90 × 10−11 | Chalcone isomerase (Paeonia lactiflora) |
| Unigene0016797 | −1.255285418 | 5.88 × 10−28 | Chalcone isomerase (Paeonia suffruticosa) |
| Unigene0052849 | −2.566247123 | 4.90 × 10−3 | Flavonol synthase (Paeonia lactiflora) |
| Unigene0054556 | −2.6979596 | 8.02 × 10−4 | Flavonol synthase (Paeonia lactiflora) |
| Unigene0041186 | 1.051737457 | 4.40 × 10−2 | Flavanone 3′-hydroxylase (Paeonia suffruticosa) |
| Unigene0041187 | 1.616420273 | 2.53 × 10−35 | Flavanone 3′-hydroxylase (Paeonia suffruticosa) |
| Unigene0030889 | −4.77004356 | 9.17 × 10−206 | Dihydroflavonol-4-reductase (Paeonia suffruticosa) |
| Unigene0012475 | −3.959416654 | 2.24 × 10−147 | Anthocyanidin synthase (Paeonia suffruticosa) |
| Unigene0014577 | 1.167626811 | 1.70 × 10−4 | Leucoanthocyanidin reductase (Cephalotus follicularis) |
| Unigene0040486 | 9.329871068 | 5.28 × 10−5 | Anthocyanidin reductase (Malus domestica) |
| Unigene0049627 | −4.379465382 | 2.57 × 10−6 | Caffeoyl-CoA O-methyltransferase |
| Unigene0010574 | −2.325935914 | 4.03 × 10−27 | Caffeoyl-CoA O-methyltransferase |
| Unigene0034757 | 2.750132254 | 2.90 × 10−62 | Caffeoyl-CoA O-methyltransferase |
| Unigene0034758 | 1.774744649 | 1.62 × 10−4 | Caffeoyl-CoA O-methyltransferase |
| Unigene0057244 | −2.050989001 | 8.69 × 10−43 | Caffeoyl-CoA O-methyltransferase |
| Unigene0034759 | 2.495157744 | 3.70 × 10−76 | Caffeoyl-CoA O-methyltransferase |
| Unigene0031564 | −2.848573896 | 4.97 × 10−6 | Coumaroylquinate 3-monooxygenase |
| Unigene0054125 | 5.051629256 | 1.30 × 10−8 | Coumaroylquinate 4′-monooxygenase |
| Unigene0035320 | 2.392910853 | 3.76 × 10−69 | Coumaroylquinate 5′-monooxygenase |
| Unigene0045080 | 3.096711889 | 3.19 × 10−59 | Shikimate O-hydroxycinnamoyltransferase |
| Unigene0058586 | 9.136649403 | 6.65 × 10−3 | Shikimate O-hydroxycinnamoyltransferase |
| Unigene0029038 | 2.210473634 | 7.78 × 10−67 | Shikimate O-hydroxycinnamoyltransferase |
| Unigene0052531 | −1.54109082 | 1.15 × 10−13 | Shikimate O-hydroxycinnamoyltransferase |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, J.; Duan, J.; Huo, D.; Shi, Q.; Niu, L.; Zhang, Y. Transcriptomic Analysis Reveals Transcription Factors Related to Leaf Anthocyanin Biosynthesis in Paeonia qiui. Molecules 2017, 22, 2186. https://doi.org/10.3390/molecules22122186
Luo J, Duan J, Huo D, Shi Q, Niu L, Zhang Y. Transcriptomic Analysis Reveals Transcription Factors Related to Leaf Anthocyanin Biosynthesis in Paeonia qiui. Molecules. 2017; 22(12):2186. https://doi.org/10.3390/molecules22122186
Chicago/Turabian StyleLuo, Jianrang, Jingjing Duan, Dan Huo, Qianqian Shi, Lixin Niu, and Yanlong Zhang. 2017. "Transcriptomic Analysis Reveals Transcription Factors Related to Leaf Anthocyanin Biosynthesis in Paeonia qiui" Molecules 22, no. 12: 2186. https://doi.org/10.3390/molecules22122186





