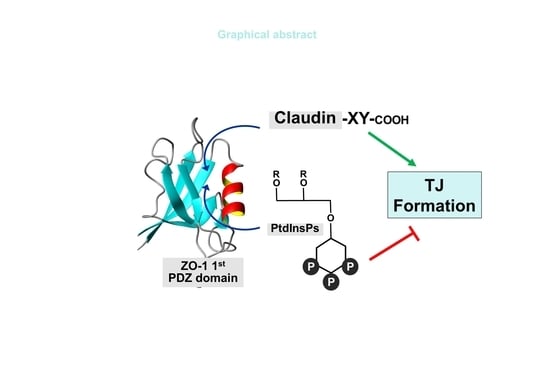
Spatial Overlap of Claudin- and Phosphatidylinositol Phosphate-Binding Sites on the First PDZ Domain of Zonula Occludens 1 Studied by NMR
Abstract
:1. Introduction
2. Results
2.1. ZO-1(PDZ1) as a Phosphoinositide-Binding Domain
2.2. Solution Structure of ZO-1(PDZ1)
2.3. Determination of the CLD- and PIP-Binding Sites by NMR Titration Experiments
2.4. Spatial Overlap of the PIP-Binding Site and CLD C-Terminal Peptide-Binding Site on ZO-1(PDZ1) Causes Competitive Binding of Phytic Acid to the ZO-1(PDZ1)-CLD Interaction
2.5. Identification of Key Residues of ZO-1(PDZ1) That Recognize CLD
3. Discussion
4. Materials and Methods
4.1. Protein Expression and Purification
4.2. PIP Strip Assays
4.3. NMR Spectroscopy
4.4. NMR Titration Experiments
4.5. Surface Plasmon Resonance (SPR) Analysis
4.6. Mutation Studies and CLD-3 Binding Assays
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Furuse, M.; Hirase, T.; Itoh, M.; Nagafuchi, A.; Yonemura, S.; Tsukita, S. Occludin: A novel integral membrane protein localizing at tight junctions. J. Cell Biol. 1993, 123, 1777–1788. [Google Scholar] [CrossRef] [PubMed]
- Ikenouchi, J.; Umeda, K.; Tsukita, S.; Furuse, M.; Tsukita, S. Requirement of ZO-1 for the formation of belt-like adherens junctions during epithelial cell polarization. J. Cell Biol. 2007, 176, 779–786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsukita, S.; Furuse, M.; Itoh, M. Multifunctional strands in tight junctions. Nat. Rev. Mol. Cell Biol. 2001, 2, 285–293. [Google Scholar] [CrossRef] [PubMed]
- Ikenouchi, J.; Furuse, M.; Furuse, K.; Sasaki, H.; Tsukita, S.; Tsukita, S. Tricellulin constitutes a novel barrier at tricellular contacts of epithelial cells. J. Cell Biol. 2005, 171, 939–945. [Google Scholar] [CrossRef] [PubMed]
- Martìn-Padura, I.; Lostaglio, S.; Schneemann, M.; Williams, L.; Romano, M.; Fruscella, P.; Panzeri, C.; Stoppacciaro, A.; Ruco, L.; Villa, A.; et al. Junctional adhesion molecule, a novel member of the immunoglobulin superfamily that distributes at intercellular junctions and modulates monocyte transmigration. J. Cell Biol. 1998, 142, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Furuse, M.; Fujita, K.; Hiiragi, T.; Fujimoto, K.; Tsukita, S. Claudin-1 and -2: novel integral membrane proteins localizing at tight junctions with no sequence similarity to occludin. J. Cell Biol. 1998, 141, 1539–1550. [Google Scholar] [CrossRef] [PubMed]
- Fanning, A.S.; Mitic, L.L.; Anderson, J.M. Transmembrane proteins in the tight junction barrier. J. Am. Soc. Nephrol. 1999, 10, 1337–1345. [Google Scholar] [PubMed]
- González-Mariscal, L.; Betanzos, A.; Nava, P.; Jaramillo, B.E. Tight junction proteins. Prog. Biophys. Mol. Biol. 2003, 81, 1–44. [Google Scholar] [CrossRef]
- González-Mariscal, L.; Betanzos, A.; Avila-Flores, A. MAGUK proteins: Structure and role in the tight junction. Semin. Cell Dev. Biol. 2000, 11, 315–324. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, S.; Iwamoto, N.; Sasaki, H.; Ohashi, M.; Oda, Y.; Tsukita, S.; Furuse, M. The E3 ubiquitin ligase LNX1p80 promotes the removal of claudins from tight junctions in MDCK cells. J. Cell Sci. 2009, 122, 985–994. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef] [PubMed]
- Harris, B.Z.; Lim, W.A. Mechanism and role of PDZ domains in signaling complex assembly. J. Cell Sci. 2001, 114, 3219–3231. [Google Scholar] [PubMed]
- Sheng, M. Molecular organization of the postsynaptic specialization. Proc. Natl. Acad. Sci. U. S. A. 2001, 98, 7058–7061. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Itoh, M.; Furuse, M.; Morita, K.; Kubota, K.; Saitou, M.; Tsukita, S. Direct binding of three tight junction-associated MAGUKs, ZO-1, ZO-2, and ZO-3, with the COOH termini of claudins. J. Cell Biol. 1999, 147, 1351–1363. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Feng, W.; Chen, J.; Chan, L.-N.; Huang, S.; Zhang, M. PDZ domains of Par-3 as potential phosphoinositide signaling integrators. Mol. Cell 2007, 28, 886–898. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, P. The prevalence and significance of PDZ domain-phosphoinositide interactions. Biochim. Biophys. Acta 2006, 1761, 947–956. [Google Scholar] [CrossRef] [PubMed]
- Kojima, T.; Fukuda, M.; Watanabe, Y.; Hamazato, F.; Mikoshiba, K. Characterization of the pleckstrin homology domain of Btk as an inositol polyphosphate and phosphoinositide binding domain. Biochem. Biophys. Res. Commun. 1997, 236, 333–339. [Google Scholar] [CrossRef] [PubMed]
- Umetsu, Y.; Goda, N.; Taniguchi, R.; Satomura, K.; Ikegami, T.; Furuse, M.; Hiroaki, H. 1H, 13C, and 15N resonance assignment of the first PDZ domain of mouse ZO-1. Biomol. NMR Assign. 2011, 5, 207–210. [Google Scholar] [CrossRef] [PubMed]
- Güntert, P. Automated NMR structure calculation with CYANA. Methods Mol. Biol. 2004, 278, 353–378. [Google Scholar] [PubMed]
- Brunger, A.T. Version 1.2 of the Crystallography and NMR system. Nat. Protoc. 2007, 2, 2728–2733. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Appleton, B.A.; Zhang, Y.; Wu, P.; Yin, J.P.; Hunziker, W.; Skelton, N.J.; Sidhu, S.S.; Wiesmann, C. Comparative structural analysis of the Erbin PDZ domain and the first PDZ domain of ZO-1. Insights into determinants of PDZ domain specificity. J. Biol. Chem. 2006, 281, 22312–22320. [Google Scholar] [CrossRef] [PubMed]
- Eswar, N.; Eramian, D.; Webb, B.; Shen, M.-Y.; Sali, A. Protein structure modeling with MODELLER. Methods Mol. Biol. 2008, 426, 145–159. [Google Scholar] [PubMed]
- Rizo, J.; Südhof, T.C. C2-domains, structure and function of a universal Ca2+-binding domain. J. Biol. Chem. 1998, 273, 15879–15882. [Google Scholar] [CrossRef] [PubMed]
- Iwaya, N.; Takasu, H.; Goda, N.; Shirakawa, M.; Tanaka, T.; Hamada, D.; Hiroaki, H. MIT domain of Vps4 is a Ca2+-dependent phosphoinositide-binding domain. J. Biochem. 2013, 153, 473–481. [Google Scholar] [CrossRef] [PubMed]
- Meerschaert, K.; Tun, M.P.; Remue, E.; De Ganck, A.; Boucherie, C.; Vanloo, B.; Degeest, G.; Vandekerckhove, J.; Zimmermann, P.; Bhardwaj, N.; et al. The PDZ2 domain of zonula occludens-1 and -2 is a phosphoinositide binding domain. Cell. Mol. Life Sci. 2009, 66, 3951. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, P.; Meerschaert, K.; Reekmans, G.; Leenaerts, I.; Small, J.V.; Vandekerckhove, J.; David, G.; Gettemans, J. PIP(2)-PDZ domain binding controls the association of syntenin with the plasma membrane. Mol. Cell 2002, 9, 1215–1225. [Google Scholar] [CrossRef]
- Zimmermann, P.; Zhang, Z.; Degeest, G.; Mortier, E.; Leenaerts, I.; Coomans, C.; Schulz, J.; N’Kuli, F.; Courtoy, P.J.; David, G. Syndecan recycling [corrected] is controlled by syntenin-PIP2 interaction and Arf6. Dev. Cell 2005, 9, 377–388. [Google Scholar] [CrossRef] [PubMed]
- Mortier, E.; Wuytens, G.; Leenaerts, I.; Hannes, F.; Heung, M.Y.; Degeest, G.; David, G.; Zimmermann, P. Nuclear speckles and nucleoli targeting by PIP2-PDZ domain interactions. EMBO J. 2005, 24, 2556–2565. [Google Scholar] [CrossRef] [PubMed]
- Gallardo, R.; Ivarsson, Y.; Schymkowitz, J.; Rousseau, F.; Zimmermann, P. Structural diversity of PDZ-lipid interactions. ChemBioChem 2010, 11, 456–467. [Google Scholar] [CrossRef] [PubMed]
- Sugi, T.; Oyama, T.; Morikawa, K.; Jingami, H. Structural insights into the PIP2 recognition by syntenin-1 PDZ domain. Biochem. Biophys. Res. Commun. 2008, 366, 373–378. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.; Nishizawa, T.; Tani, K.; Yamazaki, Y.; Tamura, A.; Ishitani, R.; Dohmae, N.; Tsukita, S.; Nureki, O.; Fujiyoshi, Y. Crystal structure of a claudin provides insight into the architecture of tight junctions. Science 2014, 344, 304–307. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.; Tani, K.; Tamura, A.; Tsukita, S.; Fujiyoshi, Y. Model for the architecture of claudin-based paracellular ion channels through tight junctions. J. Mol. Biol. 2015, 427, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Shinoda, T.; Shinya, N.; Ito, K.; Ohsawa, N.; Terada, T.; Hirata, K.; Kawano, Y.; Yamamoto, M.; Kimura-Someya, T.; Yokoyama, S.; et al. Structural basis for disruption of claudin assembly in tight junctions by an enterotoxin. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Schuck, S.; Simons, K. Polarized sorting in epithelial cells: raft clustering and the biogenesis of the apical membrane. J. Cell Sci. 2004, 117, 5955–5964. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Balda, M.S.; González-Mariscal, L.; Contreras, R.G.; Macias-Silva, M.; Torres-Marquez, M.E.; García-Sáinz, J.A.; Cereijido, M. Assembly and sealing of tight junctions: Possible participation of G-proteins, phospholipase C, protein kinase C and calmodulin. J. Membr. Biol. 1991, 122, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Balda, M.S.; Gonzalez-Mariscal, L.; Matter, K.; Cereijido, M.; Anderson, J.M. Assembly of the tight junction: the role of diacylglycerol. J. Cell Biol. 1993, 123, 293–302. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ward, P.D.; Klein, R.R.; Troutman, M.D.; Desai, S.; Thakker, D.R. Phospholipase C-gamma modulates epithelial tight junction permeability through hyperphosphorylation of tight junction proteins. J. Biol. Chem. 2002, 277, 35760–35765. [Google Scholar] [CrossRef] [PubMed]
- Ward, P.D.; Ouyang, H.; Thakker, D.R. Role of phospholipase C-beta in the modulation of epithelial tight junction permeability. J. Pharmacol. Exp. Ther. 2003, 304, 689–698. [Google Scholar] [CrossRef] [PubMed]
- Jin, R.; Song, Z.; Yu, S.; Piazza, A.; Nanda, A.; Penninger, J.M.; Granger, D.N.; Li, G. Phosphatidylinositol-3-kinase gamma plays a central role in blood-brain barrier dysfunction in acute experimental stroke. Stroke 2011, 42, 2033–2044. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Li, B.; Zhao, W.-D.; Liu, Y.-J.; Shang, D.-S.; Fang, W.-G.; Chen, Y.-H. Perfluorooctane sulfonate triggers tight junction “opening” in brain endothelial cells via phosphatidylinositol 3-kinase. Biochem. Biophys. Res. Commun. 2011, 410, 258–263. [Google Scholar] [CrossRef] [PubMed]
- Bowen, K.A.; Doan, H.Q.; Zhou, B.P.; Wang, Q.; Zhou, Y.; Rychahou, P.G.; Evers, B.M. PTEN loss induces epithelial—Mesenchymal transition in human colon cancer cells. Anticancer Res. 2009, 29, 4439–4449. [Google Scholar] [PubMed]
- Wang, H.; Quah, S.Y.; Dong, J.M.; Manser, E.; Tang, J.P.; Zeng, Q. PRL-3 down-regulates PTEN expression and signals through PI3K to promote epithelial-mesenchymal transition. Cancer Res. 2007, 67, 2922–2926. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Li, J.; Liao, W.; Feng, Y.; Yu, C.; Hu, L.; Kong, Q.; Xu, L.; Zhang, X.; Liu, W.; et al. The polycomb group protein Bmi-1 represses the tumor suppressor PTEN and induces epithelial-mesenchymal transition in human nasopharyngeal epithelial cells. J. Clin. Investig. 2009, 119, 3626–3636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, M.; Liu, M.; Wang, Y.; Chen, X.; Xu, J.; Sun, Y.; Zhao, L.; Qu, H.; Fan, Y.; Wu, C. Antagonism of miR-21 reverses epithelial-mesenchymal transition and cancer stem cell phenotype through AKT/ERK1/2 inactivation by targeting PTEN. PLoS ONE 2012, 7, e39520. [Google Scholar] [CrossRef] [PubMed]
- Goda, N.; Tenno, T.; Takasu, H.; Hiroaki, H.; Shirakawa, M. The PRESAT-vector: Asymmetric T-vector for high-throughput screening of soluble protein domains for structural proteomics. Protein Sci. 2004, 13, 652–658. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tenno, T.; Goda, N.; Tateishi, Y.; Tochio, H.; Mishima, M.; Hayashi, H.; Shirakawa, M.; Hiroaki, H. High-throughput construction method for expression vector of peptides for NMR study suited for isotopic labeling. Protein Eng. Des. Sel. 2004, 17, 305–314. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamazaki, T.; Lee, W.; Arrowsmith, C.H.; Muhandiram, D.R.; Kay, L.E. A Suite of Triple Resonance NMR Experiments for the Backbone Assignment of 15N, 13C, 2H Labeled Proteins with High Sensitivity. J. Am. Chem. Soc. 1994, 116, 11655–11666. [Google Scholar] [CrossRef]
- Cavanagh, J.; Fairbrother, W.J.; Palmer, A.G., III; Skelton, N.J.; Rance, M. Protein NMR Spectroscopy: Principles and Practice, 2nd ed.; Elsevier Academic Press: Burlington, MA, USA, 2006. [Google Scholar]
- Delaglio, F.; Grzesiek, S.; Vuister, G.W.; Zhu, G.; Pfeifer, J.; Bax, A. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 1995, 6, 277–293. [Google Scholar] [CrossRef] [PubMed]
- Goddard, T.D.; Kneller, D.G. Sparky—NMR Assignment and Integration Software. Available online: https://www.cgl.ucsf.edu/home/sparky/ (accessed on 26 September 2018).
- Jung, Y.-S.S.; Zweckstetter, M. Mars - Robust automatic backbone assignment of proteins. J. Biomol. NMR 2004, 30, 11–23. [Google Scholar] [CrossRef] [PubMed]
- Cornilescu, G.; Delaglio, F.; Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 1999, 13, 289–302. [Google Scholar] [CrossRef] [PubMed]
- Koradi, R.; Billeter, M.; Wüthrich, K. MOLMOL: A program for display and analysis of macromolecular structures. J. Mol. Graph. 1996, 14, 29–32, 51–55. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Rullmannn, J.A.C.; MacArthur, M.W.; Kaptein, R.; Thornton, J.M. AQUA and PROCHECK-NMR: Programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 1996, 8, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Piotto, M.; Saudek, V.; Sklenár, V. Gradient-tailored excitation for single-quantum NMR spectroscopy of aqueous solutions. J. Biomol. NMR 1992, 2, 661–665. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Expression vectors of ZO-1(PDZ) and its mutants are available from the authors. |







© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hiroaki, H.; Satomura, K.; Goda, N.; Nakakura, Y.; Hiranuma, M.; Tenno, T.; Hamada, D.; Ikegami, T. Spatial Overlap of Claudin- and Phosphatidylinositol Phosphate-Binding Sites on the First PDZ Domain of Zonula Occludens 1 Studied by NMR. Molecules 2018, 23, 2465. https://doi.org/10.3390/molecules23102465
Hiroaki H, Satomura K, Goda N, Nakakura Y, Hiranuma M, Tenno T, Hamada D, Ikegami T. Spatial Overlap of Claudin- and Phosphatidylinositol Phosphate-Binding Sites on the First PDZ Domain of Zonula Occludens 1 Studied by NMR. Molecules. 2018; 23(10):2465. https://doi.org/10.3390/molecules23102465
Chicago/Turabian StyleHiroaki, Hidekazu, Kaori Satomura, Natsuko Goda, Yukako Nakakura, Minami Hiranuma, Takeshi Tenno, Daizo Hamada, and Takahisa Ikegami. 2018. "Spatial Overlap of Claudin- and Phosphatidylinositol Phosphate-Binding Sites on the First PDZ Domain of Zonula Occludens 1 Studied by NMR" Molecules 23, no. 10: 2465. https://doi.org/10.3390/molecules23102465