Authentication of Herbal Medicines Dipsacus asper and Phlomoides umbrosa Using DNA Barcodes, Chloroplast Genome, and Sequence Characterized Amplified Region (SCAR) Marker
Abstract
:1. Introduction
2. Results and Discussion
2.1. Authentication of D. asper, D. japonicus, and P. umbrosa Using DNA Barcodes
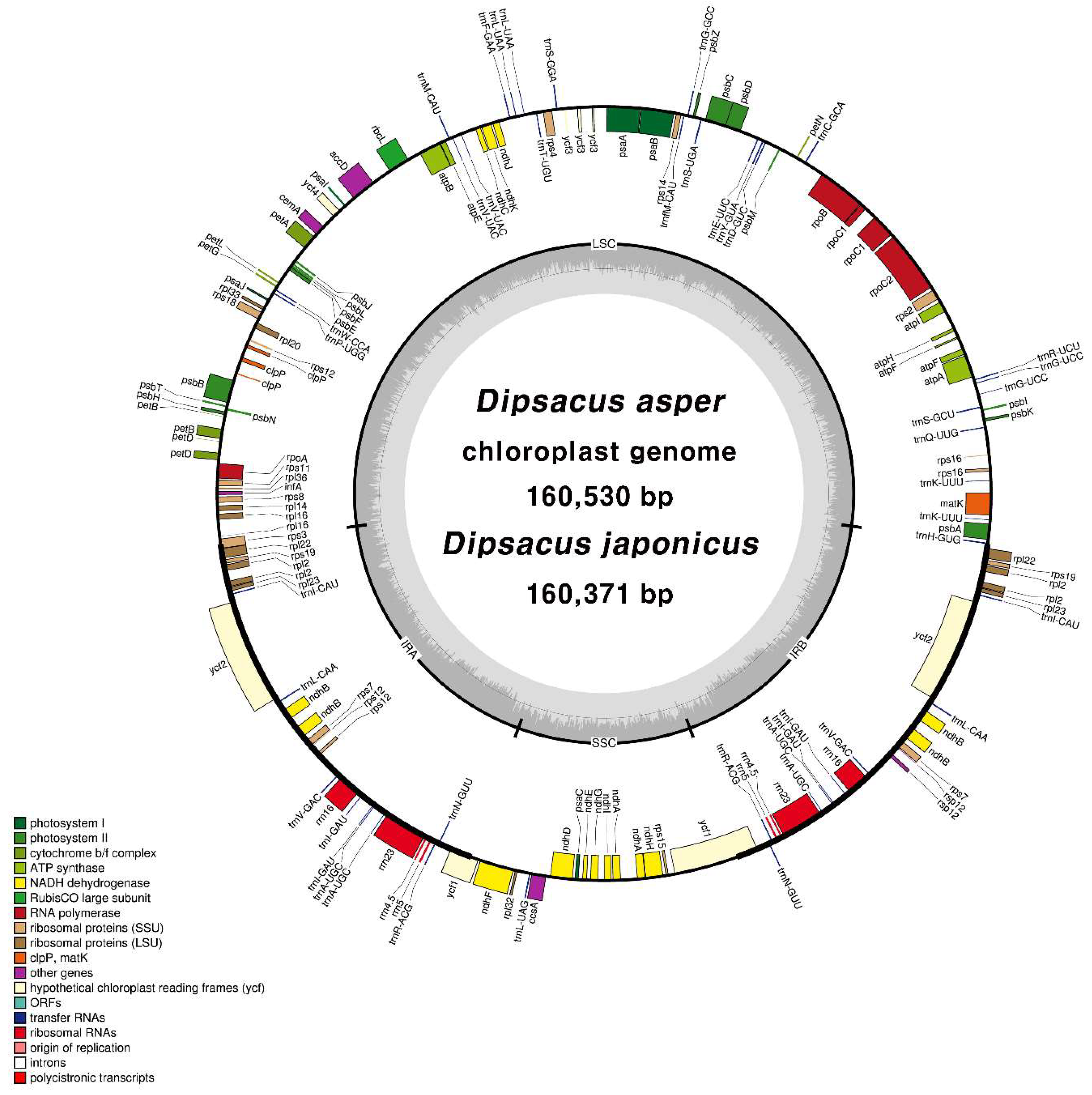
2.2. CP Genome Organisation of D. asper and D. japonicus
2.3. Analysis of Repeats in the CP Genomes of D. asper and D. japonicus
2.4. Comparative Analysis of CP Genomes within Caprifoliaceae
2.5. Authentication of D. asper, D. japonicus, and P. umbrosa Using SCAR Markers
3. Materials and Methods
3.1. Plant Materials
3.2. Universal DNA Barcode Analysis for D. asper, D. japonicus, and P. umbrosa
3.3. Genome Sequencing and Assembly
3.4. Genome Annotation and Comparative Analysis
3.5. Repeat Analysis
3.6. Phylogenetic Analysis
3.7. Development of SCAR Markers for D. asper, D. japonicus, and P. umbrosa
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ali, M.A.; Gyulai, G.; Al-Hermaid, F. Plant DNA Barcoding and Phyogenetics; LAP LAMBERT Academic Publishing: Saarbrücken, Germany, 2015; pp. 109–130. [Google Scholar]
- Li, X.; Yang, Y.; Henry, R.J.; Rossetto, M.; Wang, Y.; Chen, S. Plant DNA barcoding: From gene to genome. Biol. Rev. Camb. Philos. Soc. 2015, 90, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Kress, W.J.; Wurdack, K.J.; Zimmer, E.A.; Weigt, L.A.; Janzen, D.H. Use of DNA barcodes to identify flowering plants. Proc. Natl. Acad. Sci. USA 2005, 102, 8369–8374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Austerlitz, F.; David, O.; Schaeffer, B.; Bleakley, K.; Olteanu, M.; Leblois, R.; Veuille, M.; Laredo, C. DNA barcode analysis: A comparison of phylogenetic and statistical classification methods. BMC Bioinform. 2009, 10 (Suppl. 14), S10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mishra, P.; Kumar, A.; Nagireddy, A.; Mani, D.N.; Shukla, A.K.; Tiwari, R.; Sundaresan, V. DNA barcoding: An efficient tool to overcome authentication challenges in the herbal market. Plant Biotechnol. J. 2016, 14, 8–21. [Google Scholar] [CrossRef] [PubMed]
- Wicke, S.; Schneeweiss, G.M.; dePamphilis, C.W.; Muller, K.F.; Quandt, D. The evolution of the plastid chromosome in land plants: Gene content, gene order, gene function. Plant Mol. Biol. 2011, 76, 273–297. [Google Scholar] [CrossRef] [PubMed]
- Delannoy, E.; Fujii, S.; Colas des Francs-Small, C.; Brundrett, M.; Small, I. Rampant gene loss in the underground orchid Rhizanthella gardneri highlights evolutionary constraints on plastid genomes. Mol. Biol. Evol. 2011, 28, 2077–2086. [Google Scholar] [CrossRef] [PubMed]
- Stegemann, S.; Keuthe, M.; Greiner, S.; Bock, R. Horizontal transfer of chloroplast genomes between plant species. Proc. Natl. Acad. Sci. USA 2012, 109, 2434–2438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jansen, R.K.; Ruhlman, T.A. Plastid genomes of seed plants. In Genomics of Chloroplasts and Mitochondria; Springer: Dordrecht, The Netherlands, 2012; pp. 103–126. [Google Scholar]
- Daniell, H.; Lin, C.S.; Yu, M.; Chang, W.J. Chloroplast genomes: Diversity, evolution, and applications in genetic engineering. Genome Biol. 2016, 17, 134. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Lee, S.C.; Lee, J.; Lee, H.O.; Joh, H.J.; Kim, N.H.; Park, H.S.; Yang, T.J. Comprehensive survey of genetic diversity in chloroplast genomes and 45S nrDNAs within Panax ginseng species. PLoS ONE 2015, 10, e0117159. [Google Scholar] [CrossRef] [PubMed]
- Parks, M.; Cronn, R.; Liston, A. Increasing phylogenetic resolution at low taxonomic levels using massively parallel sequencing of chloroplast genomes. BMC Biol. 2009, 7, 84. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.S.; Yun, B.K.; Yoon, Y.H.; Hong, S.Y.; Mekapogu, M.; Kim, K.H.; Yang, T.J. Complete chloroplast genome sequence of tartary buckwheat (Fagopyrum tataricum) and comparative analysis with common buckwheat (F. esculentum). PLoS ONE 2015, 10, e0125332. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.Y.; Cheon, K.S.; Yoo, K.O.; Lee, H.O.; Cho, K.S.; Suh, J.T.; Kim, S.J.; Nam, J.H.; Sohn, H.B.; Kim, Y.H. Complete chloroplast genome sequences and comparative analysis of Chenopodium quinoa and C. album. Front. Plant Sci. 2017, 8, 1696. [Google Scholar] [CrossRef] [PubMed]
- Bhagyawant, S.S. Rapd-SCAR markers: An interface tool for authentication of traits. J. Biosci. Med. 2016, 4, 1–9. [Google Scholar] [CrossRef]
- Park, I.; Kim, W.J.; Yang, S.; Yeo, S.M.; Li, H.; Moon, B.C. The complete chloroplast genome sequence of Aconitum coreanum and Aconitum carmichaelii and comparative analysis with other Aconitum species. PLoS ONE 2017, 12, e0184257. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.J.; Moon, B.C.; Yang, S.; Han, K.S.; Choi, G.; Lee, A.Y. Rapid authentication of the herbal medicine plant species Aralia continentalis kitag. and Angelica biserrata C.Q. Yuan and R.H. Shan using ITS2 sequences and multiplex-SCAR markers. Molecules 2016, 21, 270. [Google Scholar] [CrossRef] [PubMed]
- Moon, B.C.; Kim, W.J.; Han, K.S.; Yang, S.; Kang, Y.; Park, I.; Piao, R. Differentiating authentic Adenophorae radix from its adulterants in commercially-processed samples using multiplexed its sequence-based SCAR markers. Appl. Sci. 2017, 7, 660. [Google Scholar] [CrossRef]
- Park, I.; Kim, J.; Lee, J.; Kim, S.; Cho, O.; Yang, K.; Ahn, J.; Nahm, S.; Kim, H. Development of SSR markers by next-generation sequencing of Korean landraces of chamoe (Cucumis melo var. Makuwa). Mol. Biol. Rep. 2013, 40, 6855–6862. [Google Scholar] [CrossRef] [PubMed]
- Zalapa, J.E.; Cuevas, H.; Zhu, H.; Steffan, S.; Senalik, D.; Zeldin, E.; McCown, B.; Harbut, R.; Simon, P. Using next-generation sequencing approaches to isolate simple sequence repeat (SSR) loci in the plant sciences. Am. J. Bot. 2012, 99, 193–208. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qian, J.; Song, J.; Gao, H.; Zhu, Y.; Xu, J.; Pang, X.; Yao, H.; Sun, C.; Li, X.; Li, C.; et al. The complete chloroplast genome sequence of the medicinal plant Salvia miltiorrhiza. PLoS ONE 2013, 8, e57607. [Google Scholar] [CrossRef] [PubMed]
- Park, I.; Kim, W.J.; Yeo, S.-M.; Choi, G.; Kang, Y.-M.; Piao, R.; Moon, B.C. The complete chloroplast genome sequences of Fritillaria ussuriensis Maxim. and Fritillaria cirrhosa D. Don, and comparative analysis with other Fritillaria species. Molecules 2017, 22, 982. [Google Scholar] [CrossRef] [PubMed]
- Korea Institute of Oriental Medicine (KIOM). Defining Dictionary for Medicinal Herbs. 2018. Available online: http://boncho.kiom.re.kr/codex/ (accessed on 2 May 2018).
- Moon, B.C.; Choi, G.; Yuan, Y. Origins of herbal medicines and adulterants in Korea and china. Korea Inst. Orient. Med. 2017, 1, 117. [Google Scholar]
- Ministry of Food and Drug safety (M.F.D). Korean Food Standard Codex 2010; Ministry of Food and Drug Safety: Cheongju, Korea, 2010.
- Wong, R.W.; Rabie, A.B.; Hagg, E.U. The effect of crude extract from radix Dipsaci on bone in mice. Phytother. Res. 2007, 21, 596–598. [Google Scholar] [CrossRef] [PubMed]
- Song, J.S.; Lim, K.M.; Kang, S.; Noh, J.Y.; Kim, K.; Bae, O.N.; Chung, J.H. Procoagulant and prothrombotic effects of the herbal medicine, Dipsacus asper and its active ingredient, dipsacus saponin C, on human platelets. J. Thromb. Haemost. 2012, 10, 895–906. [Google Scholar] [CrossRef] [PubMed]
- Shang, X.; Wang, J.; Li, M.; Miao, X.; Pan, H.; Yang, Y.; Wang, Y. Antinociceptive and anti-inflammatory activities of Phlomis umbrosa Turcz extract. Fitoterapia 2011, 82, 716–721. [Google Scholar] [CrossRef] [PubMed]
- Shin, T.Y.; Kim, S.H.; Kim, D.K.; Leem, K.H.; Park, J.S. Phlomis umbrosa root inhibits mast cell-dependent allergic reactions and inflammatory cytokine secretion. Phytother. Res. 2008, 22, 153–158. [Google Scholar] [CrossRef] [PubMed]
- Stevens, P.F.; Davis, H. Angiosperm Phylogeny Website. 2001. Available online: http://www.mobot.org/MOBOT/research/APweb/ (accessed on 18 May 2018).
- Deyuan, H.; Liming, M.; Fred, R.B. Dipsacus L. In Flora of China; Wu, Z.Y., Raven, P.H., Eds.; Science Press: Beijing, China; Missouri Botanical Garden Press: St. Louis, MO, USA, 2011; Volume 19, pp. 658–660. [Google Scholar]
- Dahui, L.; Zaigui, W.; Xueshi, L.; Yi, Y. Identification of the medicinal plant Dipsacus asperoides from three other species in genus Dipsacus (Dipsaceae) by internal transcribed spacer of ribosomal deoxyribonucleic acid (rDNA ITS). J. Med. Plant Res. 2012, 6, 289–295. [Google Scholar] [CrossRef]
- Sasaki, T.; Yukawa, Y.; Miyamoto, T.; Obokata, J.; Sugiura, M. Identification of RNA editing sites in chloroplast transcripts from the maternal and paternal progenitors of tobacco (Nicotiana tabacum): Comparative analysis shows the involvement of distinct trans-factors for ndhB editing. Mol. Biol. Evol. 2003, 20, 1028–1035. [Google Scholar] [CrossRef] [PubMed]
- Kahlau, S.; Aspinall, S.; Gray, J.C.; Bock, R. Sequence of the tomato chloroplast DNA and evolutionary comparison of Solanaceous plastid genomes. J. Mol. Evol. 2006, 63, 194–207. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Yi, X.; Yang, Y.X.; Su, Y.J.; Wang, T. Complete chloroplast genome sequence of a tree fern Alsophila spinulosa: Insights into evolutionary changes in fern chloroplast genomes. BMC Evol. Biol. 2009, 9, 130. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhan, D.F.; Jia, X.; Mei, W.L.; Dai, H.F.; Chen, X.T.; Peng, S.Q. Complete chloroplast genome sequence of Aquilaria sinensis (lour.) gilg and evolution analysis within the Malvales order. Front. Plant Sci. 2016, 7, 280. [Google Scholar] [PubMed]
- Ivanova, Z.; Sablok, G.; Daskalova, E.; Zahmanova, G.; Apostolova, E.; Yahubyan, G.; Baev, V. Chloroplast genome analysis of resurrection tertiary relict Haberlea rhodopensis highlights genes important for desiccation stress response. Front. Plant Sci. 2017, 8, 204. [Google Scholar] [CrossRef] [PubMed]
- Asaf, S.; Khan, A.L.; Khan, A.R.; Waqas, M.; Kang, S.M.; Khan, M.A.; Lee, S.M.; Lee, I.J. Complete chloroplast genome of Nicotiana otophora and its comparison with related species. Front. Plant Sci. 2016, 7, 843. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Cui, Y.; Chen, X.; Li, Y.; Xu, Z.; Duan, B.; Li, Y.; Song, J.; Yao, H. Complete chloroplast genomes of Papaver rhoeas and Papaver orientale: Molecular structures, comparative analysis, and phylogenetic analysis. Molecules 2018, 23, 437. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.Y.; Matzke, A.J.; Matzke, M. Complete sequence and comparative analysis of the chloroplast genome of coconut palm (Cocos nucifera). PLoS ONE 2013, 8, e74736. [Google Scholar] [CrossRef] [PubMed]
- Raubeson, L.A.; Peery, R.; Chumley, T.W.; Dziubek, C.; Fourcade, H.M.; Boore, J.L.; Jansen, R.K. Comparative chloroplast genomics: Analyses including new sequences from the angiosperms Nuphar advena and Ranunculus macranthus. BMC Genom. 2007, 8, 174. [Google Scholar] [CrossRef] [PubMed]
- Nie, X.; Lv, S.; Zhang, Y.; Du, X.; Wang, L.; Biradar, S.S.; Tan, X.; Wan, F.; Weining, S. Complete chloroplast genome sequence of a major invasive species, crofton weed (Ageratina adenophora). PLoS ONE 2012, 7, e36869. [Google Scholar] [CrossRef] [PubMed]
- Khakhlova, O.; Bock, R. Elimination of deleterious mutations in plastid genomes by gene conversion. Plant J. 2006, 46, 85–94. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kimura, M. The neutral theory of molecular evolution and the world view of the neutralists. Genome 1989, 31, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Raman, G.; Park, S. The complete chloroplast genome sequence of ampelopsis: Gene organization, comparative analysis, and phylogenetic relationships to other angiosperms. Front. Plant Sci. 2016, 7, 341. [Google Scholar] [CrossRef] [PubMed]
- Rockenbach, K.; Havird, J.C.; Monroe, J.G.; Triant, D.A.; Taylor, D.R.; Sloan, D.B. Positive selection in rapidly evolving plastid-nuclear enzyme complexes. Genetics 2016, 204, 1507–1522. [Google Scholar] [CrossRef] [PubMed]
- Redwan, R.M.; Saidin, A.; Kumar, S.V. Complete chloroplast genome sequence of MD-2 pineapple and its comparative analysis among nine other plants from the subclass Commelinidae. BMC Plant Biol. 2015, 15, 196. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Li, Q.; Hu, Z.; Li, X.; Chen, S. The complete Amomum kravanh chloroplast genome sequence and phylogenetic analysis of the Commelinids. Molecules 2017, 22, 1875. [Google Scholar]
- Park, I.; Yang, S.; Choi, G.; Kim, W.J.; Moon, B.C. The complete chloroplast genome sequences of Aconitum pseudolaeve and Aconitum longecassidatum, and development of molecular markers for distinguishing species in the Aconitum subgenus Lycoctonum. Molecules 2017, 22, 2012. [Google Scholar] [CrossRef] [PubMed]
- Jansen, R.K.; Cai, Z.; Raubeson, L.A.; Daniell, H.; Leebens-Mack, J.; Müller, K.F.; Guisinger-Bellian, M.; Haberle, R.C.; Hansen, A.K.; Chumley, T.W. Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc. Natl. Acad. Sci. USA 2007, 104, 19369–19374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moore, M.J.; Bell, C.D.; Soltis, P.S.; Soltis, D.E. Using plastid genome-scale data to resolve enigmatic relationships among basal angiosperms. Proc. Natl. Acad. Sci. USA 2007, 104, 19363–19368. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- The Angiosperm Phylogeny Group. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 2016, 181, 1–20. [Google Scholar] [CrossRef] [Green Version]
- Park, I.; Noh, P.; Kim, W.J.; Yang, S.; Moon, B.C. Discrimination of Dipsacus asperoides and D. japonicus based on the analysis of ITS2 DNA barcode sequences. Korean Herb. Med. Inf. 2017, 5, 21–29. [Google Scholar]
- Bell, C.D. Preliminary phylogeny of Valerianaceae (Dipsacales) inferred from nuclear and chloroplast DNA sequence data. Mol. Phylogenet. Evol. 2004, 31, 340–350. [Google Scholar] [CrossRef] [PubMed]
- Carlson, S.E.; Mayer, V.; Donoghue, M.J. Phylogenetic relationships, taxonomy, and morphological evolution in Dipsacaceae (Dipsacales) inferred by DNA sequence data. Taxon 2009, 58, 1075–1091. [Google Scholar]
- Yao, H.; Song, J.; Liu, C.; Luo, K.; Han, J.; Li, Y.; Pang, X.; Xu, H.; Zhu, Y.; Xiao, P. Use of ITS2 region as the universal DNA barcode for plants and animals. PLoS ONE 2010, 5, e13102. [Google Scholar] [CrossRef] [PubMed]
- Kato, H.; Oginuma, K.; GU, Z.; Hammel, B.; Tobe, H. Phylogenetic relationships of Betulaceae based on matK sequences with particular reference to the position of Ostryopsis. Acta Phytotaxon. Geobot. 1999, 49, 89–97. [Google Scholar]
- Käss, E.; Wink, M. Phylogenetic relationships in the Papilionoideae (Family leguminosae) based on nucleotide sequences of cpDNA (rbcL) and ncDNA (ITS 1 and 2). Mol. Phylogenet. Evol. 1997, 8, 65–88. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. Bioedit: A user-friendly biological sequence alignment editor and analysis program for windows 95/98/nt. Nucleic Acid Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Librado, P.; Rozas, J. Dnasp v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef] [PubMed]
- Delcher, A.L.; Salzberg, S.L.; Phillippy, A.M. Using mummer to identify similar regions in large sequence sets. Curr. Protoc. Bioinform. 2003. [Google Scholar] [CrossRef] [PubMed]
- Luo, R.; Liu, B.; Xie, Y.; Li, Z.; Huang, W.; Yuan, J.; He, G.; Chen, Y.; Pan, Q.; Liu, Y.; et al. Soapdenovo2: An empirically improved memory-efficient short-read de novo assembler. Gigascience 2012, 1, 18. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with burrows-wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef] [PubMed]
- Tillich, M.; Lehwark, P.; Pellizzer, T.; Ulbricht-Jones, E.S.; Fischer, A.; Bock, R.; Greiner, S. Geseq—Versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 2017, 45, W6–W11. [Google Scholar] [CrossRef] [PubMed]
- Carver, T.; Berriman, M.; Tivey, A.; Patel, C.; Bohme, U.; Barrell, B.G.; Parkhill, J.; Rajandream, M.A. Artemis and act: Viewing, annotating and comparing sequences stored in a relational database. Bioinformatics 2008, 24, 2672–2676. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Eddy, S.R. Trnascan-se: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Gurusaran, M.; Ravella, D.; Sekar, K. Repex: Repeat extractor for biological sequences. Genomics 2013, 102, 403–408. [Google Scholar] [CrossRef] [PubMed]
- Lohse, M.; Drechsel, O.; Bock, R. OrganellarGenomeDRAW (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. Mega6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 2007, 24, 1586–1591. [Google Scholar] [CrossRef] [PubMed]
- Thiel, T. Misa—Microsatellite Identification Tool. 2003. Available online: http://pgrc.ipk-gatersleben.de/misa/ (accessed on 17 May 2018).
- Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 1999, 27, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Warburton, P.E.; Giordano, J.; Cheung, F.; Gelfand, Y.; Benson, G. Inverted repeat structure of the human genome: The x-chromosome contains a preponderance of large, highly homologous inverted repeats that contain testes genes. Genome Res. 2004, 14, 1861–1869. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. Mafft: A novel method for rapid multiple sequence alignment based on fast fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. Jmodeltest 2: More models, new heuristics and parallel computing. Nat. Methods 2012, 9, 772. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Hohna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. Mrbayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Coulouris, G.; Zaretskaya, I.; Cutcutache, I.; Rozen, S.; Madden, T. Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinform. 2012, 13, 134. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of D. asper, D. japonicus, and P. umbrosa are available from the authors and the herbarium of KIOM. |





| Species | DNA | AL 1 Length (bp) | Parsimony Informative Sites | Variable Sites | Nucleotide Diversity (Pi) | No. of Indels | No. of Haplotypes | ||
|---|---|---|---|---|---|---|---|---|---|
| No. | % | No. | % | ||||||
| D. asper vs. D. japonicus vs. P. umbrosa | ITS2 | 390 | 91 | 23.3 | 145 | 37.2 | 0.12842 | 8 | 8 |
| matK | 1276 | 234 | 18.3 | 242 | 19.0 | 0.10377 | 2 | 8 | |
| rbcL | 1511 | 106 | 7.0 | 113 | 7.5 | 0.0394 | 1 | 6 | |
| D. asper vs. D. japonicus | ITS2 | 379 | 7 | 1.8 | 9 | 2.4 | 0.00718 | 0 | 4 |
| matK | 1264 | 5 | 0.4 | 8 | 0.6 | 0.00332 | 0 | 4 | |
| rbcL | 1501 | 3 | 0.2 | 6 | 0.4 | 0.00233 | 0 | 4 | |
| Characteristic | D. asper | D. japonicus |
|---|---|---|
| Accession number | MH074864 | MH074865 |
| Total CP genome size (bp) | 160,530 | 160,371 |
| LSC region (bp) | 86,979 | 87,193 |
| IR region (bp) | 27,821 | 27,664 |
| SSC region (bp) | 17,909 | 17,850 |
| Number of genes | 112 | 112 |
| Protein-coding genes | 79 | 79 |
| rRNA | 4 | 4 |
| tRNA | 29 | 29 |
| Introns (bp) | 14,368 | 14,392 |
| Intergenic spacers (bp) | 50,578 | 51,526 |
| GC content (%) | 38.8 | 38.8 |
| LSC (%) | 37.2 | 37.1 |
| IR (%) | 42.8 | 42.9 |
| SSC (%) | 34.2 | 34.2 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Park, I.; Yang, S.; Kim, W.J.; Noh, P.; Lee, H.O.; Moon, B.C. Authentication of Herbal Medicines Dipsacus asper and Phlomoides umbrosa Using DNA Barcodes, Chloroplast Genome, and Sequence Characterized Amplified Region (SCAR) Marker. Molecules 2018, 23, 1748. https://doi.org/10.3390/molecules23071748
Park I, Yang S, Kim WJ, Noh P, Lee HO, Moon BC. Authentication of Herbal Medicines Dipsacus asper and Phlomoides umbrosa Using DNA Barcodes, Chloroplast Genome, and Sequence Characterized Amplified Region (SCAR) Marker. Molecules. 2018; 23(7):1748. https://doi.org/10.3390/molecules23071748
Chicago/Turabian StylePark, Inkyu, Sungyu Yang, Wook Jin Kim, Pureum Noh, Hyun Oh Lee, and Byeong Cheol Moon. 2018. "Authentication of Herbal Medicines Dipsacus asper and Phlomoides umbrosa Using DNA Barcodes, Chloroplast Genome, and Sequence Characterized Amplified Region (SCAR) Marker" Molecules 23, no. 7: 1748. https://doi.org/10.3390/molecules23071748






