1. Introduction
Traditional Chinese medicines (TCMs) have long played an important role in health care. The therapeutic efficacy of TCMs has been proven by medical practices in China and has also been confirmed in many countries worldwide. However, the frequent adverse reactions of TCMs have raised public concern on the safety and toxicity of TCMs [
1]. Although the adverse reactions caused by TCMs are mostly transient and reversible with the cessation of treatment, their hepatic adverse effects may be severe and sometimes even permanent [
2]. In recent years, liver injury cases associated with the use of TCMs have increasingly been reported [
3]. Examples of some known hepatotoxic TCMs are
Polygonum multiflorum,
Dioscorea bulbifera,
Dichroa febrifuga,
Mistletoe Viscum,
Melia azedarach,
Tripterygium wilfordii,
Gynura segetum, and
Senecio vulgaris. At present, the mechanisms of liver injury induced by TCMs are still not very clear. Fortunately, some specific phytochemical ingredients that may cause liver injury have been identified, such as diterpenoid-lactones in
T. wilfordii and
D. bulbifera and pyrrolizidine alkaloids in
G. segetum and
S. vulgaris [
1]. The identification of hepatotoxic components in TCMs would be helpful to elucidate the hepatoxicity basis of TCMs and the mechanisms of TCMs-induced liver injury.
The traditional strategy to screen toxic components from TCMs often involves the isolation and purification of TCM components and their subsequent in vitro or in vivo toxicity evaluation [
4]. These experimental approaches are very time- and resource-consuming [
5]. With the advantages of time-efficiency and cost-effectiveness, in silico screening methods based on quantitative structure–toxicity relationship (QSAR) analysis have been used as a complementary tool to identify hepatotoxic compounds in TCMs [
6,
7]. Nevertheless, a common drawback of the above experimental and computational methods is that the hepatotoxicity of each compound is evaluated individually with ignorance of the synergistic effects of multiple constituents in TCMs. Recently, fingerprint-toxicity modeling methods have been employed to screen the major hepatotoxic components in
Euodia rutaecarpa,
D. bulbifera, and
P. multiflorum [
8,
9,
10]. These methods are based on the characterization of the correlations between characteristic chromatographic peaks and the holistic hepatotoxicity of TCMs by multivariate statistical analysis. However, these methods suffer from the disadvantages of extensive workload and high cost (for example, linked to the increased use of animals).
In recent years, screening methods based on live cell extraction and high-performance liquid chromatography–mass spectrometry analysis have been applied to identify potential bioactive components in TCMs [
11,
12,
13]. These methods demonstrated the advantages of high efficiency and specificity over previous methods. Inspired by these methods, hepatotoxic components in TCMs could also be screened using cell-based screening strategies. Unlike the screening of bioactive components of TCMs, however, the screening of hepatotoxic components should not be restricted to the identification of the compounds accumulated in hepatocytes. A further determination of intracellular accumulation of the identified compounds is necessary. It is well known that the dose makes the poison. Drug-induced toxicity is thought to be mediated by the drug accumulation in target cells. Intracellular drug concentrations were reported to have significant impact on cytotoxicity [
14]. It remains to be seen whether intracellular accumulation can be used as an indicator of cytotoxicity for a fast screening of hepatotoxic components in TCMs.
In the present study, we attempted to develop a whole-cell-based strategy to screen and predict potential hepatotoxic components in
Chelidonium majus L., a perennial herbaceous plant of the family
Papaveraceae. Ch. majus, also known as greater celandine, is widely used in Asia and Europe. It has been used as a folk medicine due to its various pharmacological effects [
15], such as antibacterial, anti-inflammatory, and anti-tumour actions [
16,
17]. Isoquinoline alkaloids are the main chemical components of
Ch. majus, among which sanguinarine, coptisine, and chelerythrine showed cytotoxicity in rat hepatocytes [
18].
Ch. majus may also induce liver injury after long-term administration in large doses [
19]. To identify the components contributed to the holistic hepatotoxicity of
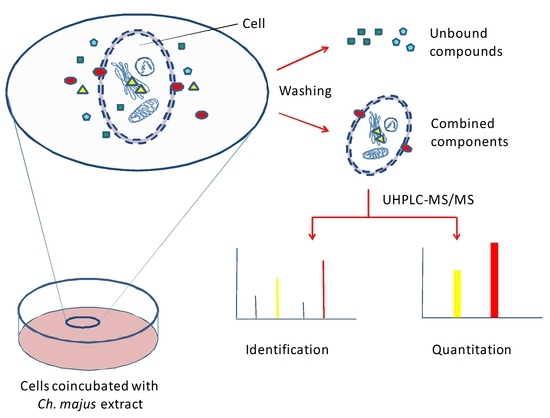
Ch. majus, the proposed strategy involved: (1) exposing HL-7702 cells to
Ch. majus extract; (2) identifying the compounds accumulated in HL-7702 cells; (3) determining the intracellular accumulation of potential hepatotoxic compounds. Liquid chromatography–tandem mass spectrometry (LC–MS/MS) was used for the identification of hepatotoxic components and the determination of their intracellular accumulation.
3. Materials and Methods
3.1. Chemicals and Materials
Ch. majus was obtained from Tongrentang Chinese Medicne (Shenyang, China). The reference standards of protopine, allocryptopine, chelerythrine, chelidonine, and magnoflorine were purchased from Shenzhen Medherb Biotechnology Co., Ltd. (Shenzhen, China). Sanguinarine and coptisine were obtained from Chengdu Mansite Pharmaceutical Co., Ltd. (Sichuan, China). The IS, palmatine, was purchased from the National Institute for the Control of Pharmaceutical and Biological Products (Beijing, China). Trypsin and RPMI-1640 medium were purchased from Gibco Life Technologies Corporation (Grand Island, NY, USA), and newborn calf serum from Gibco Life Technologies Corporation (Auckland, New Zealand). Penicillin, streptomycin, and PBS(-) were obtained from Sangon Biotech (Shanghai, China). Dimethyl sulphoxide (DMSO) was obtained from Tianjin Bodi Chemical Co., Ltd. (Tianjin, China). The commercial Cell Counting Kit-8 (CCK8) was obtained from Dojindo (Kumamoto, Japan). All of the cell culture plastics were bought from Corning incorporated (Corning, NY, USA). HL-7702 cells were bought from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). Acetonitrile and formic acid of HPLC grade were obtained from Sigma-Aldrich (St. Louis, MO, USA) and Dikma Technologies (Lake Forest, CA, USA), respectively. Water used throughout the study was prepared by a Millipore Milli-Q Academic system (Millipore, Bedford, MA, USA). All other reagents were of analytical grade.
3.2. Preparation of Calibration Standards and QC Samples
Stock solutions of each analyte were prepared in acetonitrile/water (1:1, v/v). A mixed stock solution was prepared by combining appropriate aliquots of each stock solution and diluting with acetonitrile/water (1:1, v/v). A series of mixed working solutions were made by successively diluting the mixed stock solution with acetonitrile/water. An IS solution was prepared at the concentration of 80 ng/mL. QC solutions were made from a separately prepared mixed stock solution. All solutions were stored at 4 °C before use.
Calibration standards were prepared daily by spiking each mixed working solution into blank cell lysates to yield the desired final concentrations. QC samples were prepared by the same procedures to yield low, medium, and high final concentrations.
3.3. Sample Preparation
The powder of Ch. majus was extracted with five-fold (w/v) 70% ethanol for 1 h three times in an ultrasonic bath and then evaporated to dryness. The dried extract was dissolved in DMSO and filtered through a 0.22 μm cellulose acetate membrane. The filtrate was diluted with RPMI-1640 medium to yield a final concentration equivalent to 2.5 mg/mL of raw herb.
3.4. Cell Culture
HL-7702 cells were dissociated with 0.05% trypsin–EDTA and subcultured in 10 cm-diameter dishes with RPMI-1640 medium containing 10% newborn calf serum, 100 µg/mL streptomycin, and 100 U/mL penicillin. The cells were maintained at 37 °C in a humidified atmosphere of 5% CO2 and 95% air. The culture medium was replaced every 48 h and removed until the cells reached about 80% confluence.
3.5. Cell Exposure to the Test Sample
HL-7702 were incubated in the medium containing Ch.majus for 30 min. Then, the cells were washed and lysed (following the procedures described below). The cell lysate was used for the identification study. HL-7702 were incubated in blank medium and in medium containing Ch. majus for 5, 10, 30, 60, 90, 120, 180, and 360 min. Then, the cells were washed and lysed for the determination study.
3.6. Cell Lysate Preparation
After incubation, the cells were washed three times with PBS(-) and then harvested with 0.05% trypsin–EDTA. The cell suspension (about 5 × 106 cells) was centrifuged at 1000 rpm for 5 min and further washed three times with PBS(-). The cell pellets were lysed in 200 µL of ice-cold methanol by vortex-mixing for 2 min. The cell lysate was centrifuged at 12,000 rpm for 4 min. To identify the compounds accumulated in the cells, 2 µL of supernatant was directly injected into the LC–MS/MS system. For the purpose of determining the intracellular accumulation of the compounds, 20 µL of supernatant was placed into a 1.5 mL Eppendorf tube and then 20 µL of IS working solution and 100 µL of acetonitrile were added. The mixture was vortex-mixed for 1 min. A 2 µL aliquot of the mixture was injected into the LC–MS/MS system.
3.7. LC–MS/MS Analysis
LC–MS/MS analysis was performed on a Shimadzu Nexera LC system coupled with a Shimadzu LCMS-8050 triple quadrupole tandem mass spectrometer equipped with an ESI interface in positive ionization mode (Shimadzu, Kyoto, Japan). Chromatographic separation was performed on an ACQUITY UPLC C18 column (2.1 × 50 mm, 1.7 µm) maintained at 40 °C by gradient elution, using water containing 0.1% formic acid (solvent A) and acetonitrile containing 0.1% formic acid (solvent B) as the mobile phase at a flow rate of 0.3 mL/min. The gradient program for compound identification was: 0–4.0 min, 20%–30% B, 4.0–8.0 min, 30%–90% B, 8.0–10.0 min, 20% B. The gradient program for compound quantitation was: 0–1.0 min 20% B, 1.0–3.5 min, 60% B, 3.5–4.0 min, 20% B. The MS conditions were set as follows: nebulizing gas flow, 3.0 L/min; drying gas flow, 10.0 L/min; heat-block temperature, 400 °C; desolvation line temperature, 250 °C; interface voltage, 4 kV. Product ion scan and MRM mode were used for identification and quantitation, respectively. The ion transitions and collision energies are listed in
Table 6. Data acquisition and processing were conducted using LabSolutions LCMS Ver.5.82 SP1 (Shimadzu, Kyoto, Japan).
3.8. Cytotoxicity Assay
Cell viability was performed using the Cell Counting Kit-8 (CCK-8) assay according to the manufacturer’s instructions. In brief, HL-7702 cells were seeded in a 96-well plate at a density of 25,000 per well. After 24 h, the cell growing medium was removed and replaced with fresh medium containing Ch. majus or the test compound at various concentrations. After an additional 24 h, 10 µL of WST reagent was added into each well and incubated with the cells for 2 h. The optical density of each well was measured by a microplate reader (Corona Electric, Ibaraki-Ken, Japan) at a wave length of 450 nm.