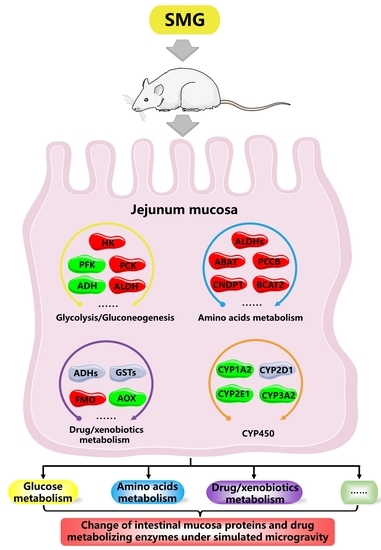
Investigation on Intestinal Proteins and Drug Metabolizing Enzymes in Simulated Microgravity Rats by a Proteomics Method
Abstract
:1. Introduction
2. Results and Discussion
2.1. Proteomic Analysis
2.1.1. Gene-Ontology Classification
2.1.2. Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Analysis
2.2. Metabolism-Related DEPs
2.2.1. Glucose Metabolism
2.2.2. Amino Acids Metabolism
2.2.3. Drugs and Xenobiotics Metabolism
SMG Effect on ADH Enzyme System (ADHs)
SMG Effect on GST Enzyme System (GSTs)
SMG Effect on FMO and AOX
2.3. Determination of Drug Metabolism CYP450
3. Materials and Methods
3.1. Animals Treatment and Samples Collection
3.2. Histomorphology
3.3. Protein Extraction and In-Gel Digestion
3.4. HPLC-MS/MS Analysis
3.5. Protein Identification and Bioinformatic Analysis
3.6. Western-Blot Analysis of IDMEs
3.7. Determination of ADH, ALDH, and GST Activity
3.8. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Riva, D.; Rossitto, F.; Battocchio, L. Postural muscle atrophy prevention and recovery and bone remodelling through high frequency proprioception for astronauts. Acta Astronaut. 2009, 65, 813–819. [Google Scholar] [CrossRef]
- Hackney, K.J.; Brown, L.T.C.; Stone, K.A.; Tennent, D.J. The role of blood flow restriction training to mitigate sarcopenia, dynapenia, and enhance clinical recovery. Tech. Orthop. 2018, 33, 98–105. [Google Scholar] [CrossRef]
- Ting, H.Y.; Du, Y.Y.; Peng, H.R.; Hu, X.J.; Wu, C.R.; Li, Q.; Dong, D.D.; Li, J.B.; Shang, P. Evaluation of osteoclast-derived exosomal miRNA under simulated microgravity conditions using next-generation sequencing. Acta Astronaut. 2019, 161, 75–86. [Google Scholar]
- Blaber, E.A.; Dvorochkin, N.; Chialing, L.; Alwood, J.S.; Yousuf, R.; Pianetta, P.; Globus, R.K.; Burns, B.P.; Almeida, E.A.C. Microgravity induces pelvic bone loss through osteoclastic activity, osteocytic osteolysis, and osteoblastic cell cycle inhibition by CDKN1a/p21. PLoS ONE 2013, 8, e61372. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pastushkova, L.K.; Kashirina, D.N.; Brzhozovskiy, A.G.; Kononikhin, A.S.; Tiys, E.S.; Ivanisenko, V.A.; Koloteva, M.I.; Nikolaev, E.N.; Larina, I.M. Evaluation of cardiovascular system state by urine proteome after manned space flight. Acta Astronaut. 2019, 160, 594–600. [Google Scholar] [CrossRef]
- Lu, Y.M.; Jiao, B.; Lee, J.; Zhang, L.; Yu, Z.B. Simulated microgravity increases myocardial susceptibility to ischemia-reperfusion injury via a deficiency of AMP-activated protein kinase. Can. J. Physiol. Pharmacol. 2016, 95, 59–71. [Google Scholar] [CrossRef]
- Hatzistergos, K.E.; Jiang, Z.J.; Valasaki, K.; Takeuchi, L.M.; Balkan, W.; Atluri, P.; Saur, D.; Seidler, B.; Tsinoremas, N.; Difede, D.L.; et al. Simulated microgravity impairs cardiac autonomic neurogenesis from neural crest cells. Stem Cells Dev. 2018, 27, 819–830. [Google Scholar] [CrossRef]
- Cooke, W.H.; Convertino, V.A. Sympathetic nervous system and spaceflight. Acta Astronaut. 2007, 60, 223–233. [Google Scholar] [CrossRef]
- Tauber, S.; Lauber, B.A.; Paulsen, K.; Layer, L.E.; Oliver, U. Cytoskeletal stability and metabolic alterations in primary human macrophages in long-term microgravity. PLoS ONE 2017, 12, e0175599. [Google Scholar] [CrossRef] [Green Version]
- Paulsen, K.; Thiel, C.; Timm, J.; Schmidt, P.M.; Huber, K.; Tauber, S.; Hemmersbach, R.; Seibt, D.; Kroll, H.; Grote, K.; et al. Microgravity-induced alterations in signal transduction in cells of the immune system. Acta Astronaut. 2010, 67, 1116–1125. [Google Scholar] [CrossRef]
- Wang, S.B.; Zhang, Y.S.; Guo, J.J.; Kang, L.T.; Deng, Y.L.; Li, Y.J. Investigation on rat intestinal homeostasis alterations induced by 7-day simulated microgravity effect based on a proteomic approach. Acta Astronaut. 2020, 166, 560–566. [Google Scholar] [CrossRef]
- Jin, M.L.; Zhang, H.; Zhao, K.; Xu, C.L.; Shao, D.L.; Huang, Q.S.; Shi, J.L.; Yang, H. Responses of intestinal mucosal barrier functions of rats to simulated weightlessness. Front. Physiol. 2018, 9, 729–741. [Google Scholar] [CrossRef]
- Li, P.P.; Shi, J.X.; Zhang, P.; Wang, K.; Li, J.L.; Liu, H.J.; Zhou, Y.; Xu, X.; Hao, J.; Sun, X.Y.; et al. Simulated microgravity disrupts intestinal homeostasis and increases colitis susceptibility. FASEB J. 2015, 29, 3263–3273. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.J.; Qu, L.N.; Li, Y.X.; Chen, X.P.; Li, Y.H. Progresses and prospects of weightlessness physiology in China. Space Med. Med. Eng. 2018, 31, 131–139. [Google Scholar]
- Ramanathan, R.; Geary, R.S.; Bourne, D.W.A.; Putcha, L. Bioavailability fo intranasal promethazine dosage forms in dogs. Pharmacol. Res. 1998, 38, 35–39. [Google Scholar] [CrossRef] [PubMed]
- Wotring, V.E. Medication use by U.S. crewmembers on the international space station. FASEB J. 2015, 29, 4417–4423. [Google Scholar] [CrossRef] [Green Version]
- Barger, L.K.; Flynn-Evans, E.E.; Kubey, A.; Walsh, L.; Ronda, J.M.; Wang, W.; Wright, K.P.; Czeisler, C.A. Prevalence of sleep deficiency and use of hypnotic drugs in astronauts before, during, and after spaceflight: An observational study. Lancet Neurol. 2014, 13, 904–912. [Google Scholar] [CrossRef] [Green Version]
- Blue, R.S.; Bayuse, T.M.; Daniels, V.R.; Wotring, V.E.; Suresh, R.; Mulcahy, R.A.; Antonsen, E.L. Supplying a pharmacy for NASA exploration spaceflight: Challenges and current understanding. NPJ Microgravity 2019, 5, 14. [Google Scholar] [CrossRef] [Green Version]
- Mithieux, G.; Gautier-Stein, A. Intestinal glucose metabolism revisited. Diabetes Res. Clin. Pract. 2014, 105, 295–301. [Google Scholar] [CrossRef]
- Zhang, J.; Dai, Z.L.; Zhu, W.Y. Recent progress in intestinal essential amino acids metabolism and its function. Parenter. Enter. 2010, 17, 55–59. [Google Scholar]
- Varma, M.V. Role of intestinal transporters and metabolism in the oral absorption of drug and prodrugs. Curr. Drug Metab. 2010, 11, 715. [Google Scholar]
- Sáenz-Robles, M.T.; Toma, D.; Cantalupo, P.; Zhou, J.; Gong, H.; Edwards, C.; Pipas, J.M.; Xie, W. Repression of intestinal drug metabolizing enzymes by the SV40 large T antigen. Oncogene 2007, 26, 5124–5131. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rushmore, T.H.; Tony Kong, A. Pharmacogenomics, regulation and signaling pathways of phase I and II drug metabolizing enzymes. Curr. Drug Metab. 2002, 3, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Iyanagi, T. Molecular mechanism of phase I and phase II drug-metabolizing enzymes: Implications for detoxification. Int. Rev. Cytol. 2007, 260, 35–112. [Google Scholar] [PubMed]
- Drozdzik, M.; Busch, D.; Lapczuk, J.; Müller, J.; Ostrowski, M.; Kurzawski, M.; Oswald, S. Protein abundance of clinically relevant drug-metabolizing enzymes in the human liver and intestine: A comparative analysis in paired tissue specimens. Clin. Pharmacol. Ther. 2018, 104, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Groza, P.; Bordeianu, A.; Boca, A. Modifications of the digestive tract in rats submitted to an orbital flight aboard the Soviet satellite Cosmos 1129. Physiologie 1983, 20, 35–44. [Google Scholar] [PubMed]
- Groza, P.; Bordeianu, A.; Boca, A. The digestive tract of rat after flight in the biosatellite Cosmos 1667. Physiologie 1987, 24, 187–190. [Google Scholar]
- Groza, P.; Bordeianu, A.; Boca, A. Histochemical alterations in the digestive tract of rats after 13 days of flight in the "Cosmos 1887" biosatellite. Physiologie 1989, 26, 3–6. [Google Scholar]
- Merrill, A.H., Jr.; Hoel, M.; Wang, E.; Mullins, R.E.; Popova, I.A. Altered carbohydrate, lipid, and xenobiotic metabolism by liver from rats flown on Cosmos 1887. FASEB J. 1990, 4, 95–100. [Google Scholar] [CrossRef]
- Moskaleva, N.; Moysa, A.; Novikova, S.; Tikhonova, O.; Zgoda, V.; Archakov, A. Spaceflight Effects on Cytochrome P450 Content in Mouse Liver. PLoS ONE 2015, 10, e0142374. [Google Scholar] [CrossRef] [Green Version]
- Chen, B.; Guo, J.J.; Wang, S.B.; Kang, L.T.; Deng, Y.L.; Li, Y.J. Simulated microgravity altered the metabolism of loureirin B and the expression of major cytochrome P450 in liver of rats. Front. Pharmacol. 2019, 9, 1130. [Google Scholar]
- Kast, J.; Yu, Y.; Seubert, C.N.; Wotring, V.E.; Derendorf, H. Drugs in space: Pharmacokinetics and pharmacodynamics in astronauts. Eur. J. Pharm. Sci. 2017, 109S, S2–S8. [Google Scholar] [PubMed]
- Greenblatt, D.J.; von Moltke, L.L.; Harmatz, J.S.; Mertzanis, P.; Graf, J.A.; Durol, A.L.B.; Counihan, M.; Roth-Schechter, B.; Shader, R.I. Kinetic and dynamic interaction study of zolpidem with ketoconazole, itraconazole, and fluconazole. Clin. Pharmacol. Ther. 1998, 64, 661–671. [Google Scholar]
- Uehara, S.; Uno, Y.; Nakanishi, K.; Ishii, S.; Inoue, T.; Sasaki, E.; Yamazaki, H. Marmoset cytochrome P450 3A4 ortholog expressed in liver and small-Intestine tissues efficiently metabolizes midazolam, alprazolam, nifedipine, and testosterone. Drug Metab. Dispos. 2017, 45, 457. [Google Scholar]
- Pichard, L.; Gillet, G.; Thenot, J.P.; Maurel, P. Zolpidem metabolism by human cytochrome P450S. Biol. Psychiat. 1997, 42, 45. [Google Scholar]
- Wotring, V.E. Space Pharmacology; Springer Science & Business Media: New York, NY, USA, 2012. [Google Scholar]
- Morey-Holton, E.; Wronski, T.J. Animal models for simulating weightlessness. Physiologist 1982, 24, 45–48. [Google Scholar]
- Yao, Y.J.; Yue, Y.; Sun, H.P.; Jiang, S.Z.; Wu, X.Y. Compliance changes in femoral veins of rabbits after 21 days of simulated weightlessness. Aerosp. Environ. Med. 2006, 40, 29–33. [Google Scholar]
- Tang, Z.Z.; Li, Y.Z.; Bai, G.E. Effects of 7 d head-down bed rest on human vectorcardiogram (VCG). Space Med. Med. Eng. 2003, 16, 410–413. [Google Scholar]
- Morey-Holton, E.R.; Globus, R.K. Hindlimb unloading rodent model: Technical aspects. J. Appl. Physiol. 2002, 92, 1367–1377. [Google Scholar]
- Ma, J.; Chen, T.; Wu, S.F.; Yang, C.Y.; Bai, M.Z.; Shu, K.X.; Li, K.L.; Zhang, G.Q.; Jin, Z.; He, F.C.; et al. iProX: An integrated proteome resource. Nucleic Acids Res. 2019, 47, D1211–D1217. [Google Scholar]
- Jonscher, K.R.; Alfonso-Garcia, A.; Suhalim, J.L.; Orlicky, D.J.; Potma, E.O.; Ferguson, V.L.; Bouxsein, M.L.; Bateman, T.A.; Stodieck, L.S.; Levi, M.; et al. Spaceflight Activates Lipotoxic Pathways in Mouse Liver. PLoS ONE 2016, 11, e0152877. [Google Scholar]
- Souza, M.A.; Ribeiro, M.Z.; Silva, D.P.; Pessoa, A.; Vitolo, M. Effect of pH on the stability of hexokinase and glucose 6-phosphate dehydrogenase. Appl. Biochem. Biotechnol. 2002, 98, 265–272. [Google Scholar] [CrossRef]
- Webb, B.A.; Forouhar, F.; Szu, F.E.; Seetharaman, J.; Tong, L.; Barber, D.L. Structures of human phosphofructokinase-1 and atomic basis of cancer-associated mutations. Nature 2015, 523, 111–114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramirez, J.; Periyakaruppan, A.; Sarkar, S.; Ramesh, G.T.; Sharma, S.C. Effect of simulated microgravity on the activity of regulatory enzymes of glycolysis and gluconeogenesis in mice liver. Microgravity Sci. Technol. 2014, 25, 303–309. [Google Scholar] [CrossRef] [Green Version]
- Chen, B.; Li, G.Q.; Li, Y.Z.; Cho, J.L.; Wang, J.P.; Gao, J.Y.; Deng, Y.L.; Li, Y.J. Label-free quantitative proteomics of rat liver exposed to simulated microgravity. Acta Astronaut. 2020, 170, 251–260. [Google Scholar] [CrossRef]
- Stein, T.P.; Schluter, M.D.; Galante, A.T.; Soteropoulos, P.; Tolias, P.P.; Grindeland, R.E.; Moran, M.M.; Wang, T.J.; Polansky, M.; Wade, C.E. Energy metabolism pathways in rat muscle under conditions of simulated microgravity. J. Nutr. Biochem. 2002, 13, 471–478. [Google Scholar] [CrossRef]
- Zhang, Q.; Koser, S.L.; Donkin, S.S. Propionate induces the bovine cytosolic phosphoenolpyruvate carboxykinase promoter activity. J. Dairy Sci. 2016, 99, 6654–6664. [Google Scholar] [CrossRef]
- Pedro, L.M.; Josue, B.; Eric, A.A.; Ramon, H.G.; Francisco, C.; Lindsay, E.W.; David, A.S.; Pascual, L.B.; José, C.; John, M.D. Dynamic acetylation of phosphoenolpyruvate carboxykinase toggles enzyme activity between gluconeogenic and anaplerotic reactions. Mol. Cell 2018, 71, 718–732. [Google Scholar]
- Rees, S.D.; Britten, A.C.; Bellary, S.; O’hare, J.P.; Kelly, M.A. The promoter polymorphism -232C/G of the PCK1 gene is associated with type 2 diabetes in a UK-resident South Asian population. BMC Med. Genet. 2009, 10, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Schlicker, L.; Szebenyi, M.; Ortiz, S.R.; Heinz, A.; Hiller, K.; Field, M.S. Unexpected roles for ADH1 and SORD in catalyzing the final step of erythritol biosynthesis. J. Biol. Chem. 2019, 294, 16095–16108. [Google Scholar] [CrossRef]
- Burke, S.J.; Batdorf, H.M.; Burk, D.H.; Martin, T.M.; Mendoza, T.; Stadler, K.; Alami, W.; Karlstad, M.D.; Robson, M.J.; Blakely, R.D.; et al. Pancreatic deletion of the interleukin-1 receptor disrupts whole body glucose homeostasis and promotes islet β-cell de-differentiation. Mol. Metab. 2018, 14, 95–107. [Google Scholar] [PubMed]
- Jiang, P.Z.; Dong, Z.; Ma, B.C.; Ni, Z.Z.; Duan, H.K.; Li, X.D.; Wang, B.; Ma, X.F.; Ji, X.Z.; Li, M.G. Effect of vanadyl rosiglitazone, a new insulin-mimetic vanadium complexes, on glucose homeostasis of diabetic mice. Appl. Biochem. Biotechnol. 2016, 180, 841–851. [Google Scholar] [CrossRef] [PubMed]
- Tanner, J.J. SAXS fingerprints of aldehyde dehydrogenase oligomers. Data Brief 2015, 5, 745–751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Uchakin, P.N.; Smith, S.M.; Uchakina, O.N.; Tobin, B.W. Amino acid metabolism and immune responsiveness in vitro in a microgravity analog. STAR 2007, 45, 1–16. [Google Scholar]
- Zhao, G.J. The research progress of ABAT gene in human diseases. Fudan U. J. Med. Sci. 2016, 43, 357–361, 367. [Google Scholar]
- Ita-Pérez, D.D.; Méndez, I.; Vázquez-Martínez, O.; Villalobos-Leal, M.; Díaz-Muñoz, M. Synchronization by food access modifies the daily variations in expression and activity of liver GABA transaminase. BioMed Res. Int. 2014, 2014, 590581. [Google Scholar]
- Jiang, H.; Rao, K.S.; Yee, V.C.; Kraus, J.P. Characterization of four variant forms of human propionyl-CoA carboxylase expressed in Escherichia coli. J. Biol. Chem. 2005, 280, 27719–27727. [Google Scholar] [CrossRef] [Green Version]
- Schomburg, D.; Stephan, D. beta-Ala-His dipeptidase. Enzym. Handb. 1998, 15, 423–426. [Google Scholar]
- Schuster, J.; Binder, S. The mitochondrial branched-chain aminotransferase (AtBCAT-1) is capable to initiate degradation of leucine, isoleucine and valine in almost all tissues in Arabidopsis thaliana. Plant Mol. Biol. 2005, 57, 241–254. [Google Scholar] [CrossRef]
- Burrin, D.G.; Reeds, P.J. Alternative fuels in the gastrointestinal tract. Curr. Opin. Gastroenterol. 1997, 13, 165–170. [Google Scholar] [CrossRef]
- Seiler, N.; Raul, F. Polyamines and the intestinal tract. Crit. Rev. Clin. Lab. Sci. 2007, 44, 365–411. [Google Scholar] [CrossRef] [PubMed]
- Kakoki, M.; Kim, H.S.; Edgell, C.J.S.; Maeda, N.; Smithies, O.; Mattson, D.L. Amino acids as modulators of endothelium-derived nitric oxide. Am. J. Physiol. Renal Physiol. 2006, 291, 297–304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reeds, P.J.; Burrin, D.G.; Jahoor, F.; Wykes, L.; Henry, J.; Frazer, E.M. Enteral glutamate is almost completely metabolized in first pass by the gastrointestinal tract of infant pigs. Am. J. Physiol. 1996, 270, 413–418. [Google Scholar] [CrossRef]
- Su, N.; Zhou, A.G. Catabolism of amino acids in intestinal mucosa and its significance. J. Sichuan I. Anim. Husb. Vet. Med. 2000, 2, 47–52. [Google Scholar]
- Höög, J.O.; Hedberg, J.J.; Strömberg, P.; Svensson, S. Mammalian alcohol dehydrogenase—Functional and structural implications. J. Biomed. Sci. 2001, 8, 71–76. [Google Scholar] [CrossRef]
- Zhu, R.; Liu, J.G.; Yang, L. Effect of Zhihuang decoction on liver and stomach ADH_1 mRNA expression in rats with alcohol-induced liver disease. Chin. J. Integr. Tradit. West. Med. Liver Dis. 2007, 17, 352–354. [Google Scholar]
- Buervenich, S.; Sydow, O.; Carmine, A.; Zhang, Z.P.; Anvret, M.; Olson, L. Alcohol dehydrogenase alleles in Parkinson’s disease. Mov. Disord. 2000, 15, 813–818. [Google Scholar] [CrossRef]
- Buervenich, S.; Carmine, A.; Galter, D.; Shahabi, H.N.; Johnels, B.; Holmberg, B.; Ahlberg, J.; Nissbrandt, H.; Eerola, J.; Hellström, O.; et al. A rare truncating mutation in ADH1C (G78Stop) shows significant association with parkinson disease in a large international sample. Arch. Neurol. 2005, 62, 74–78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Westerlund, M.; Belin, A.C.; Felder, M.R.; Olson, L.; Galter, D. High and complementary expression patterns of alcohol and aldehyde dehydrogenases in the gastrointestinal tract: Implications for Parkinson’s disease. FEBS J. 2007, 274, 1212–1223. [Google Scholar] [CrossRef]
- Chen, L.J.; Xu, Y.Q. Research progress of glutathione S-transferase gene family. Acta Acad. Med. Wannan 2003, 22, 144–146. [Google Scholar]
- Wu, Z.L.; Yu, X.B.; Wu, Z.D. Recent advances of cytosolic glutathione S-transferase. Chin. Trop. Med. 2004, 4, 285–286. [Google Scholar]
- Lei, A.P.; Chen, H.; Li, S.F.; Hu, Z.L. Function, application, cloning and expression of glutatllione S-transferases. Environ. Sci. Technol. 2009, 32, 85–91. [Google Scholar]
- Mcdonagh, P.D.; Judah, D.J.; Hayes, J.D.; Lian, L.Y.; Neal, G.E.; Wolf, C.R.; Roberts, G.C.K. Determinants of specificity for aflatoxin B1-8,9-epoxide in alpha-class glutathione S-transferases. Biochem. J. 1999, 339, 95–101. [Google Scholar] [CrossRef] [PubMed]
- Nan, H.M.; Kim, H.; Lim, H.S.; Choi, J.K.; Kawamoto, T.; Kang, J.W.; Lee, C.H.; Kim, Y.D.; Kwon, E.H. Effects of occupation, life style and genetic polymorphism of CYP1A1, GSTM1, and GSTT1 on urinary 1-hydroxypyrene and 2-naphthol concentration. Carcinogenesis 2001, 22, 787–793. [Google Scholar] [CrossRef]
- Chen, Y.; Li, G.L.; Ji, Z.Y.; Xu, J.M.; Wu, C.L. Genetic polymorphisms of NQO1, GSTT1, GSTM1 and susceptibility to chronic benzene poisoning. Chin. J. Ind. Hyg. Occup. Dis. 2005, 23, 1–5. [Google Scholar]
- Rojas, M.; Cascorbi, I.; Alexandrov, K.; Kriek, E.; Auburtin, G.; Mayer, L.; Kopp-Schneider, A.; Roots, I.; Bartsch, H. Modulation of benzo[a]pyrene diolepoxide-DNA adduct levels in human white blood cells by CYP1A1, GSTM1 and GSTT1 polymorphism. Carcinogenesis 2000, 21, 35–41. [Google Scholar] [CrossRef] [Green Version]
- Cummings, B.S.; Parker, J.C.; Lash, L.H. Role of cytochrome P450 and glutathione S-transferase α in the metabolism and cytotoxicity of trichloroethylene in rat kidney. Biochem. Pharmacol. 2000, 59, 531–543. [Google Scholar] [CrossRef]
- Mathew, N.; Srinivasan, L.; Karunan, T.; Ayyanar, E.; Muthuswamy, K. Studies on filarial GST as a target for antifilarial drug development—In silico and in vitro inhibition of filarial GST by substituted 1,4-naphthoquinones. J. Mol. Model. 2011, 17, 2651–2657. [Google Scholar] [CrossRef]
- Eve, D.L.; Claude, G. Impact of reactive oxygen species on spermatozoa: A balancing act between beneficial and detrimental effects. Hum. Reprod. 1995, 10, 15–21. [Google Scholar]
- Carcia-Ruiz, C.; José, C.F. Mitochondrial glutathione: Hepatocellular survival-death switch. J. Gastro. Hepat. 2006, 21, 3–6. [Google Scholar] [CrossRef]
- Hayes, J.D.; Pulford, D.J. The glutathione S-transferase supergene family: Regulation of GST and the contribution of the isoenzymes to cancer chemoprotection and drug resistance part I. Crit. Rev. Biochem. Mol. Biol. 1995, 30, 521–600. [Google Scholar] [CrossRef]
- Rabot, S.; Szylit, O.; Nugon-Baudon, L.; Meslin, J.C.; Vaissade, P.; Popot, F.; Viso, M. Variations in digestive physiology of rats after short duration flights aboard the US space shuttle. Dig. Dis. Sci. 2000, 45, 1687–1695. [Google Scholar] [CrossRef]
- Hao, D.C.; Yang, L. Role of flavin-containing monooxygenase 3 in drug metabolism. Chin. J. Pharmacol. Toxicol. 2008, 22, 391–395. [Google Scholar]
- Mi, J.Q.; Li, Y. Advances in the study of aldehyde oxidases. Acta Pharm. Sin. 2014, 49, 582–589. [Google Scholar]
- Furge, L.L.; Guengerich, F.P. Cytochrome P450 enzymes in drug metabolism and chemical toxicology: An introduction. Biochem. Mol. Biol. Educ. 2006, 34, 66–74. [Google Scholar] [CrossRef]
- Park, B.K. Cytochrome P450 enzymes in the heart. Lancet 2000, 355, 945–946. [Google Scholar] [CrossRef]
- Hoen, P.A.; Commandeur, J.N.; Vermeulen, N.P.; Van Berkel, T.J.; Bijsterbosch, M.K. Selective induction of cytochrome P450 3A1 by dexamethasone in cultured rat hepatocytes: Analysis with a novel reverse transcriptase-polymerase chain reaction assay section sign. Biochem. Pharmacol. 2000, 60, 1509–1518. [Google Scholar] [CrossRef]
- Elbarbry, F.A.; Mcnamara, P.J.; Alcorn, J. Ontogeny of hepatic CYP1A2 and CYP2E1 expression in rat. J. Biochem. Mol. Toxicol. 2007, 21, 41–50. [Google Scholar] [CrossRef]
- Su, J.H.; Chang, C.; Xiang, Q.; Zhou, Z.W.; Luo, R.; Yang, L.; He, Z.X.; Yang, H.T.; Li, J.N.; Bei, Y.; et al. Xyloketal B, a marine compound, acts on a network of molecular proteins and regulates the activity and expression of rat cytochrome P450 3a: A bioinformatic and animal study. Drug Des. Dev. Ther. 2014, 8, 2555–2602. [Google Scholar]
- Chen, C.L.; You, X.F. Research progress of cytochrome P450 3A. World Notes Antibiot. 2004, 25, 124–128. [Google Scholar]
- Gonzalez, F.J.; Meyer, U.A. Molecular genetics of the debrisoquin-sparteine polymorphism. Clin. Pharmacol. Ther. 1991, 50, 233–238. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.G.; Chowdhury, S.K. Drug metabolism in drug design and development: Basic concepts and practice. Am. J. Ther. 2008, 16, 466–467. [Google Scholar]
- Bertilsson, L.; Dahl, M.L.; Dalén, P.; Al-Shurbaji, A. Molecular genetics of CYP2D6: Clinical relevance with focus on psychotropic drugs. Br. J. Clin. Pharmacol. 2002, 53, 111–122. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, X.Y.; Liu, G.L. Research advance on the characteristics and application of CYP450 metabolic enzymes. Chin. J. Clin. Pharmacol. Ther. 2008, 13, 942–946. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.L.; Xie, H.G. Research progress of cytochrome P450 2E1. Chin. J. Clin. Pharmacol. 1997, 13, 57–62. [Google Scholar]
- Davis, R.; Markham, A.; Balfour, J.A. Ciprofloxacin. Drugs 1996, 51, 1019–1074. [Google Scholar] [CrossRef]
- Kuang, Z.J. Side effects of ciprofloxacin. Guangdong Trace Elem. Sci. 2001, 8, 15–20. [Google Scholar]
- Bagian, J.P.; Beck, B.G. Promethazine and its use as a treatment for space motion sickness. In Sixth Annual Workshop on Space Operations Applications and Research (SOAR 1992); NASA Lyndon B. Johnson Space Center: Houston, TX, USA, 1992; Volume 2, p. 574. [Google Scholar]
- Nakamura, K.; Yokoi, T.; Inoue, K.; Shimada, N.; Ohashi, N.; Kume, T.; Kamataki, T. CYP2D6 is the principal cytochrome P450 responsible for metabolism of the histamine 111 antagonist promethazine in human liver microsomes. Pharm. Genom. 1996, 6, 449–457. [Google Scholar] [CrossRef]
- Zhang, R.T.; Wang, Q.W. Metabolic properties of promethazine in rat’s isolated gastrointestinal tract. Medi. J. Nat. Def. For. Northwest Chin. 2017, 38, 431–435. [Google Scholar]
- Risberg, A.M.; Risberg, J.; Ingvar, D.H. Effects of promethazine on nocturnal sleep in normal man. Psychopharmacologia 1975, 43, 279–284. [Google Scholar] [CrossRef]
- Schepp, W.; Classen, M. Omeprazole in the acute treatment of gastric ulcer. Scand. J. Gastroentero. 1989, 24, 58–62. [Google Scholar] [CrossRef] [PubMed]
- Fang, H.M.; Mei, Q.; Jin, J.; Diao, L.; Chen, M.L.; Xu, X.H.; Xu, J.M. Study on intestinal first pass effect and possible mechanism of omeprazole. Chin. Pharmacol. Bull. 2009, 25, 1679–1680. [Google Scholar]
- Saveliev, S.V.; Woodroofe, C.C.; Sabat, G.; Adams, C.M.; Klaubert, D.; Wood, K.; Urh, M. Mass spectrometry compatible surfactant for optimized in-gel protein digestion. Anal. Chem. 2013, 85, 907–914. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Carlson, A.; Sinitcyn, P.; Mann, M.; Cox, J. Visualization of LC-MS/MS proteomics data in MaxQuant. Proteomics 2015, 15, 1453–1456. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moritz, C.P. Tubulin or not tubulin: Heading towards total protein staining as loading control in Western blots. Proteomics 2017, 17, 1600189. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Sample Availability: Samples of the compounds are not available from the authors. |







| Gene IDs | Protein Names | Fold Change | p-Value |
|---|---|---|---|
| Q8K4D8 | Aldehyde dehydrogenase family 1 member A3 (ALDH1A3) | 3.37 | 0.0121 |
| P07379 | Cytosolic phosphoenolpyruvate carboxykinase (PCK1) | 3.08 | 0.0122 |
| P27881 | Hexokinase-2 (HK2) | 2.58 | 0.0214 |
| P05708 | Hexokinase-1 (HK1) | 2.40 | 0.0031 |
| P11884 | mitochondrial Aldehyde dehydrogenase (ALDH2) | 2.33 | 0.0005 |
| P30839 | Aldehyde dehydrogenase family 3 member A2 (ALDH3A2) | 2.17 | 0.0005 |
| P30835 | ATP-dependent 6-phosphofructokinase, liver type (PFK) | 0.49 | 0.0078 |
| P00884 | Fructose-bisphosphate aldolase B (ALDOB) | 0.20 | 0.0002 |
| P06757 | Alcohol dehydrogenase 1 (ADH1) | 0.12 | 0.0007 |
| Gene IDs | Protein Names | Fold Change | p-Value |
|---|---|---|---|
| P50554 | Mitochondrial 4-aminobutyrate aminotransferase (ABAT) | 8.96 | 0.0005 |
| Q02253 | Aldehyde dehydrogenase family 6 member A1 (ALDH6A1) | 5.38 | 0.0031 |
| P07633 | Mitochondrial Propionyl-CoA carboxylase beta chain (PCCB) | 5.33 | 0.0012 |
| Q8K4D8 | Aldehyde dehydrogenase family 1 member A3 (ALDH1A3) | 3.37 | 0.0121 |
| P07896 | Peroxisomal bifunctional enzyme (EHHADH) | 2.61 | 0.0353 |
| O08590 | Membrane primary amine oxidase (AOC3) | 2.49 | 0.0001 |
| P11884 | mitochondrial Aldehyde dehydrogenase (ALDH2) | 2.33 | 0.0005 |
| Q66HG3 | Beta-Ala-His dipeptidase (CNDP1) | 2.33 | 0.0146 |
| P30839 | Aldehyde dehydrogenase family 3 member A2 (ALDH3A2) | 2.17 | 0.0005 |
| O35854 | Mitochondrial branched-chain-amino-acid aminotransferase (BCAT2) | 2.06 | 0.0042 |
| Q5QE78 | Aldehyde oxidase 2 (AOX2) | 0.41 | 0.00002 |
| Metabolic Enzymes | CON Group (U/min mg Prot) | SMG Group (U/min mg Prot) |
|---|---|---|
| ADH | 4.75 ± 0.26 | 2.93 ± 0.57 *,1 |
| ALDH | 15.83 ± 1.82 | 16.30 ± 2.11 |
| GST | 20.67 ± 1.49 | 16.55 ± 1.17 |
| Gene IDs | Protein Names | Fold Change | p-Value |
|---|---|---|---|
| Q9Z1B2 | Glutathione S-transferase Mu 5 (GSTM5) | 7.54 | 0.0041 |
| P08011 | Microsomal glutathione S-transferase 1 (MGST1) | 4.92 | 0.0089 |
| Q8K4C0 | Dimethylaniline monooxygenase [N-oxide-forming] 5 (FMO5) | 3.57 | 0.0016 |
| Q8K4D8 | Aldehyde dehydrogenase family 1 member A3 (ALDH1A3) | 3.37 | 0.0121 |
| P04041 | Glutathione peroxidase 1 (GPX1) | 2.40 | 0.0425 |
| P15684 | Aminopeptidase N (APN) | 2.01 | 0.0097 |
| Q5QE78 | Aldehyde oxidase 2 (AOX2) | 0.41 | 0.0000 |
| P14942 | Glutathione S-transferase alpha-4 (GSTA4) | 0.32 | 0.0004 |
| P04903 | Glutathione S-transferase alpha-2 (GSTA2) | 0.21 | 0.0003 |
| P46418 | Glutathione S-transferase alpha-5 (GSTA5) | 0.14 | 0.0005 |
| P06757 | Alcohol dehydrogenase 1 (ADH1) | 0.12 | 0.0007 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, H.; Guo, J.; Li, Y.; Zhang, Y.; Wang, J.; Gao, J.; Deng, Y.; Li, Y. Investigation on Intestinal Proteins and Drug Metabolizing Enzymes in Simulated Microgravity Rats by a Proteomics Method. Molecules 2020, 25, 4391. https://doi.org/10.3390/molecules25194391
Liu H, Guo J, Li Y, Zhang Y, Wang J, Gao J, Deng Y, Li Y. Investigation on Intestinal Proteins and Drug Metabolizing Enzymes in Simulated Microgravity Rats by a Proteomics Method. Molecules. 2020; 25(19):4391. https://doi.org/10.3390/molecules25194391
Chicago/Turabian StyleLiu, Huayan, Jingjing Guo, Yujuan Li, Yushi Zhang, Jiaping Wang, Jianyi Gao, Yulin Deng, and Yongzhi Li. 2020. "Investigation on Intestinal Proteins and Drug Metabolizing Enzymes in Simulated Microgravity Rats by a Proteomics Method" Molecules 25, no. 19: 4391. https://doi.org/10.3390/molecules25194391