The Discovery of New Drug-Target Interactions for Breast Cancer Treatment
Abstract
:1. Introduction
2. Results
2.1. Performance Evaluation
2.2. Features Generation
2.2.1. Parameter Setting for PsePSSM and DCCA Coefficient
2.2.2. The Dimensionality of the Generated Features
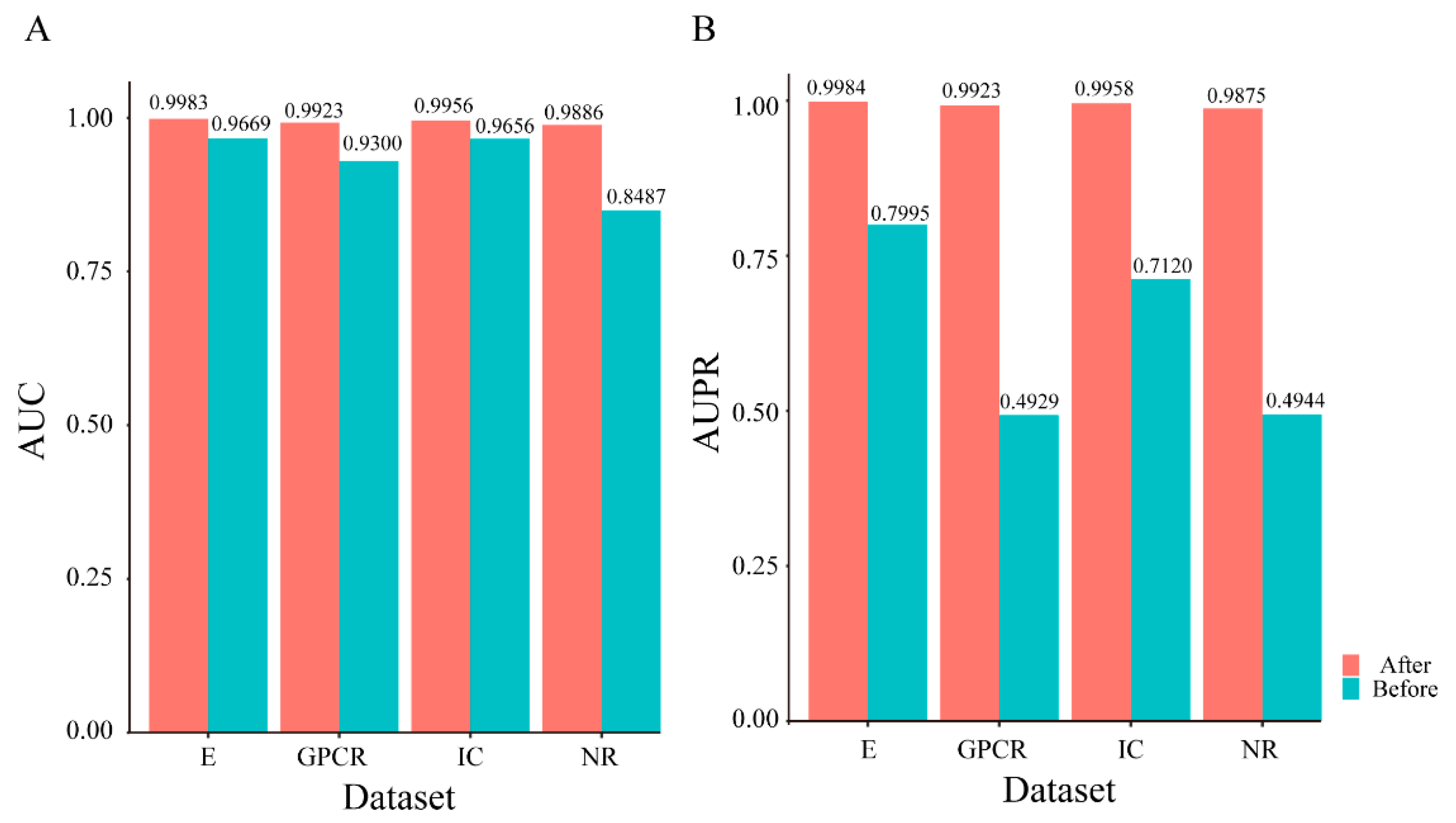
2.3. Predictive Performance of Lasso for Dimensionality Reduction
2.4. Predictive Performance of SMOTE for Imbalanced Datasets
2.5. Predictive Performance of RF for DTIs Prediction
2.6. Predictive Performance of PsePDC-DTIs Compared with State-of-the-Art Methods
2.7. Predictive Performance of PsePDC-DTIs Compared with State-of-the-Art Methods
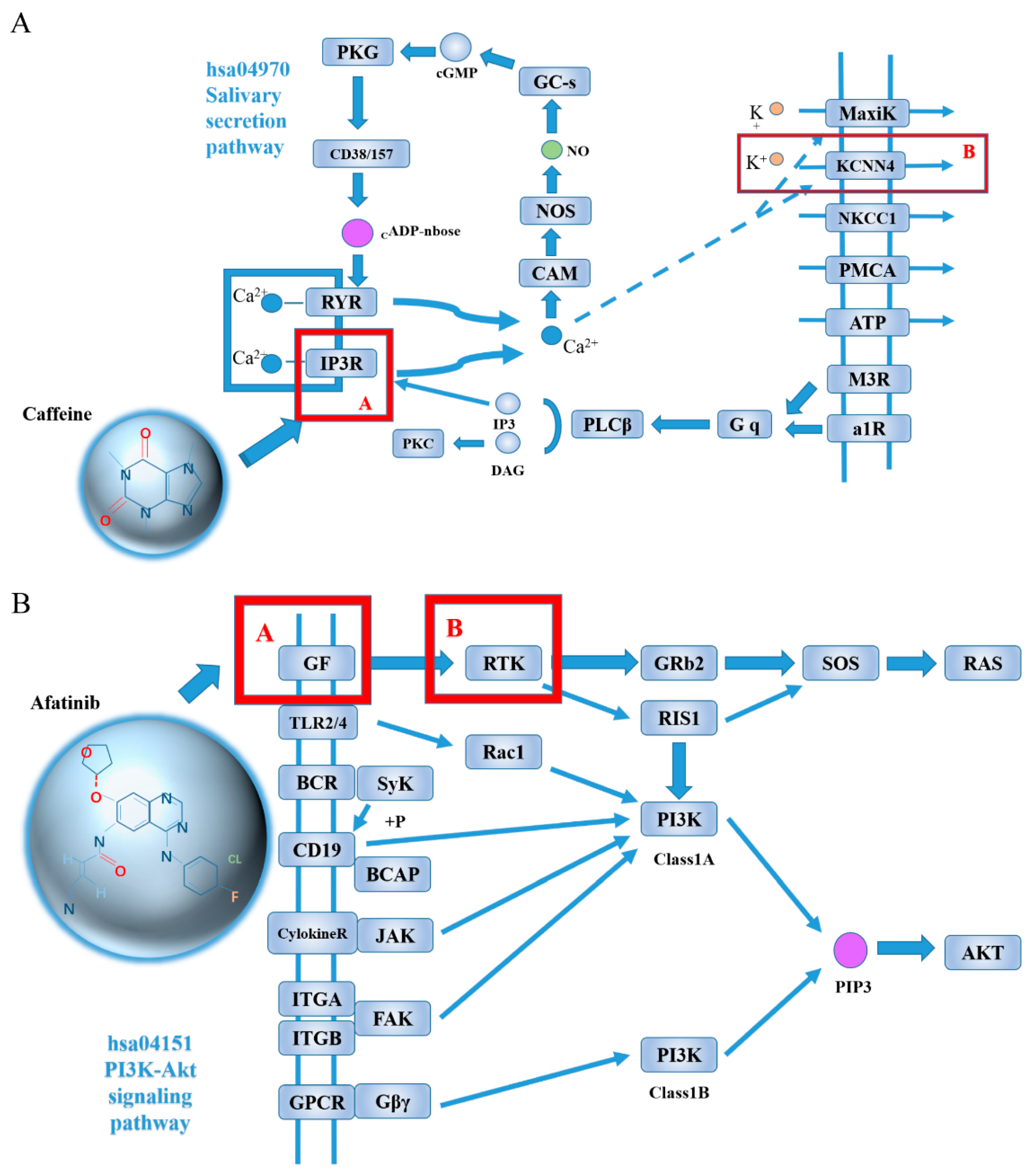
3. Discussion
4. Materials and Methods
4.1. Datasets
4.1.1. Benchmark Datasets
4.1.2. DTIs Dataset Constructed by Drugs of FDA-Approved and Targets of Breast Cancer
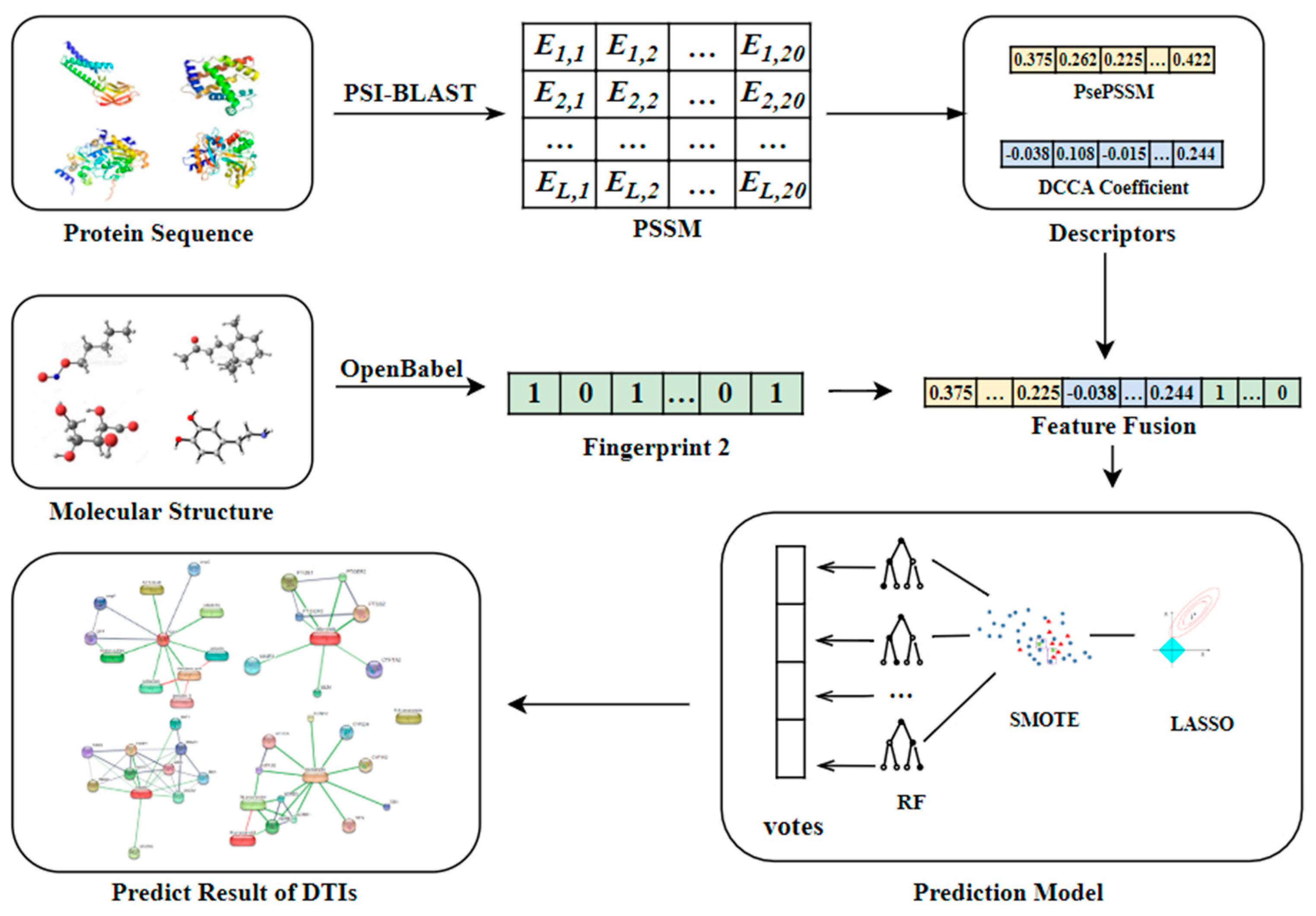
4.2. Methods for Features Generation
4.2.1. Pseudo-Position Specific Scoring Matrix (PsePSSM)
4.2.2. Detrended Cross-Correlation Analysis Coefficient (DCCA Coefficient)
4.2.3. FP2 Molecular Fingerprint
4.3. Lasso for Dimensionality Reduction of Features
4.4. SMOTE for High-Dimensional Class-Imbalanced Data
4.5. RF for DTIs Prediction
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
References
- Maurya, A.P.; Brahmachari, S. Current Status of Breast Cancer Management in India. Indian J. Surg. 2020. [Google Scholar] [CrossRef]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [Green Version]
- Green, M.; Raina, V. Epidemiology, Screening and Diagnosis of Breast Cancer in the Asia–Pacific Region: Current Perspectives and Important Considerations. Asia Pac. J. Clin. Oncol. 2008, 4, S5–S13. [Google Scholar] [CrossRef]
- You, J.; McLeod, R.D.; Hu, P. Predicting Drug-Target Interaction Network Using Deep Learning Model. Comput. Biol. Chem. 2019, 80, 90–101. [Google Scholar] [CrossRef]
- Chong, C.R.; Sullivan, D.J. New Uses for Old Drugs. Nature 2007, 448, 645–646. [Google Scholar] [CrossRef]
- Kola, I.; Landis, J. Can the Pharmaceutical Industry Reduce Attrition Rates? Nat. Rev. Drug Discov. 2004, 3, 711–715. [Google Scholar] [CrossRef]
- Kamb, A.; Wee, S.; Lengauer, C. Why Is Cancer Drug Discovery so Difficult? Nat. Rev. Drug Discov. 2007, 6, 115–120. [Google Scholar] [CrossRef]
- Chen, H.-R.; Sherr, D.H.; Hu, Z.; DeLisi, C. A Network Based Approach to Drug Repositioning Identifies Plausible Candidates for Breast Cancer and Prostate Cancer. BMC Med. Genom. 2016, 9, 51. [Google Scholar] [CrossRef] [Green Version]
- DiMasi, J.A.; Hansen, R.W.; Grabowski, H.G. The Price of Innovation: New Estimates of Drug Development Costs. J. Health Econ. 2003, 22, 151–185. [Google Scholar] [CrossRef] [Green Version]
- Baxter, J.S.; Leavy, O.C.; Dryden, N.H.; Maguire, S.; Johnson, N.; Fedele, V.; Simigdala, N.; Martin, L.-A.; Andrews, S.; Wingett, S.W.; et al. Capture Hi-C Identifies Putative Target Genes at 33 Breast Cancer Risk Loci. Nat. Commun. 2018, 9, 1028. [Google Scholar] [CrossRef]
- Wu, L.; Shi, W.; Long, J.; Guo, X.; Michailidou, K.; Beesley, J.; Bolla, M.K.; Shu, X.-O.; Lu, Y.; Cai, Q.; et al. A Transcriptome-Wide Association Study of 229,000 Women Identifies New Candidate Susceptibility Genes for Breast Cancer. Nat. Genet. 2018, 50, 968–978. [Google Scholar] [CrossRef]
- Takenaka, T. Classical vs Reverse Pharmacology in Drug Discovery. BJU Int. 2008, 88, 7–10. [Google Scholar] [CrossRef]
- Ezzat, A.; Wu, M.; Li, X.-L.; Kwoh, C.-K. Computational Prediction of Drug–Target Interactions Using Chemogenomic Approaches: An Empirical Survey. Brief. Bioinform. 2019, 20, 1337–1357. [Google Scholar] [CrossRef] [Green Version]
- Mitchell, J.B. The Relationship between the Sequence Identities of Alpha Helical Proteins in the PDB and the Molecular Similarities of Their Ligands. J. Chem. Inf. Comput. Sci. 2001, 41, 1617–1622. [Google Scholar] [CrossRef]
- Keiser, M.J.; Roth, B.L.; Armbruster, B.N.; Ernsberger, P.; Irwin, J.J.; Shoichet, B.K. Relating Protein Pharmacology by Ligand Chemistry. Nat. Biotechnol. 2007, 25, 197–206. [Google Scholar] [CrossRef] [Green Version]
- Mahmud, S.M.H.; Chen, W.; Meng, H.; Jahan, H.; Liu, Y.; Hasan, S.M.M. Prediction of Drug-Target Interaction Based on Protein Features Using Undersampling and Feature Selection Techniques with Boosting. Anal. Biochem. 2020, 589, 113507. [Google Scholar] [CrossRef]
- Sachdev, K.; Gupta, M.K. A Comprehensive Review of Feature Based Methods for Drug Target Interaction Prediction. J. Biomed. Inform. 2019, 93, 103159. [Google Scholar] [CrossRef]
- Xie, L.; Evangelidis, T.; Xie, L.; Bourne, P.E. Drug Discovery Using Chemical Systems Biology: Weak Inhibition of Multiple Kinases May Contribute to the Anti-Cancer Effect of Nelfinavir. PLoS Comput. Biol. 2011, 7, e1002037. [Google Scholar] [CrossRef]
- Mousavian, Z.; Masoudi-Nejad, A. Drug-Target Interaction Prediction via Chemogenomic Space: Learning-Based Methods. Expert Opin. Drug Metab. Toxicol. 2014, 10, 1273–1287. [Google Scholar] [CrossRef]
- Rayhan, F.; Ahmed, S.; Shatabda, S.; Farid, D.M.; Mousavian, Z.; Dehzangi, A.; Rahman, M.S. IDTI-ESBoost: Identification of Drug Target Interaction Using Evolutionary and Structural Features with Boosting. Sci. Rep. 2017, 7, 17731. [Google Scholar] [CrossRef]
- Mousavian, Z.; Khakabimamaghani, S.; Kavousi, K.; Masoudi-Nejad, A. Drug–Target Interaction Prediction from PSSM Based Evolutionary Information. J. Pharmacol. Toxicol. Methods 2016, 78, 42–51. [Google Scholar] [CrossRef]
- Yu, B.; Li, S.; Qiu, W.; Wang, M.; Du, J.; Zhang, Y.; Chen, X. Prediction of Subcellular Location of Apoptosis Proteins by Incorporating PsePSSM and DCCA Coefficient Based on LFDA Dimensionality Reduction. BMC Genom. 2018, 19, 478. [Google Scholar] [CrossRef] [Green Version]
- Shi, H.; Liu, S.; Chen, J.; Li, X.; Ma, Q.; Yu, B. Predicting Drug-Target Interactions Using Lasso with Random Forest Based on Evolutionary Information and Chemical Structure. Genomics 2019, 111, 1839–1852. [Google Scholar] [CrossRef]
- Sun, Y.; Wong, A.K.C.; Kamel, M.S. Classification of Imbalanced Data: A Review. Int. J. Pattern Recognit. Artif. Intell. 2009, 23, 687–719. [Google Scholar] [CrossRef]
- Yamanishi, Y.; Kotera, M.; Kanehisa, M.; Goto, S. Drug-Target Interaction Prediction from Chemical, Genomic and Pharmacological Data in an Integrated Framework. Bioinformatics 2010, 26, i246–i254. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, Z. A Semi-Supervised Method for Drug-Target Interaction Prediction with Consistency in Networks. PLoS ONE 2013, 8, e62975. [Google Scholar] [CrossRef] [Green Version]
- Huang, Y.-A.; You, Z.-H.; Chen, X. A Systematic Prediction of Drug-Target Interactions Using Molecular Fingerprints and Protein Sequences. Curr. Protein Pept. Sci. 2018, 19, 468–478. [Google Scholar] [CrossRef]
- Li, Z.; Han, P.; You, Z.-H.; Li, X.; Zhang, Y.; Yu, H.; Nie, R.; Chen, X. In Silico Prediction of Drug-Target Interaction Networks Based on Drug Chemical Structure and Protein Sequences. Sci. Rep. 2017, 7, 11174. [Google Scholar] [CrossRef] [Green Version]
- Gönen, M. Predicting Drug-Target Interactions from Chemical and Genomic Kernels Using Bayesian Matrix Factorization. Bioinformatics 2012, 28, 2304–2310. [Google Scholar] [CrossRef]
- Liu, Y.; Wu, M.; Miao, C.; Zhao, P.; Li, X.-L. Neighborhood Regularized Logistic Matrix Factorization for Drug-Target Interaction Prediction. PLoS Comput. Biol. 2016, 12, e1004760. [Google Scholar] [CrossRef]
- Yamanishi, Y.; Araki, M.; Gutteridge, A.; Honda, W.; Kanehisa, M. Prediction of Drug–Target Interaction Networks from the Integration of Chemical and Genomic Spaces. Bioinformatics 2008, 24, i232–i240. [Google Scholar] [CrossRef]
- Kanehisa, M. From Genomics to Chemical Genomics: New Developments in KEGG. Nucleic Acids Res. 2006, 34, D354–D357. [Google Scholar] [CrossRef] [PubMed]
- Schomburg, I.; Chang, A.; Ebeling, C.; Gremse, M.; Heldt, C.; Huhn, G.; Schomburg, D. BRENDA, the Enzyme Database: Updates and Major New Developments. Nucleic Acids Res. 2004, 32, D431–D433. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Günther, S.; Kuhn, M.; Dunkel, M.; Campillos, M.; Senger, C.; Petsalaki, E.; Ahmed, J.; Urdiales, E.G.; Gewiess, A.; Jensen, L.J.; et al. SuperTarget and Matador: Resources for Exploring Drug-Target Relationships. Nucleic Acids Res. 2008, 36, D919–D922. [Google Scholar] [CrossRef]
- Wishart, D.S.; Knox, C.; Guo, A.C.; Cheng, D.; Shrivastava, S.; Tzur, D.; Gautam, B.; Hassanali, M. DrugBank: A Knowledgebase for Drugs, Drug Actions and Drug Targets. Nucleic Acids Res. 2008, 36, D901–D906. [Google Scholar] [CrossRef] [PubMed]
- Ba-alawi, W.; Soufan, O.; Essack, M.; Kalnis, P.; Bajic, V.B. DASPfind: New Efficient Method to Predict Drug–Target Interactions. J. Cheminform. 2016, 8, 15. [Google Scholar] [CrossRef] [Green Version]
- Chou, K.-C. Prediction of Protein Cellular Attributes Using Pseudo-Amino Acid Composition. Proteins Struct. Funct. Genet. 2001, 43, 246–255. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T. Protein Secondary Structure Prediction Based on Position-Specific Scoring Matrices. J. Mol. Biol. 1999, 292, 195–202. [Google Scholar] [CrossRef] [Green Version]
- Altschul, S. Gapped BLAST and PSI-BLAST: A New Generation of Protein Database Search Programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [Green Version]
- Podobnik, B.; Stanley, H.E. Detrended Cross-Correlation Analysis: A New Method for Analyzing Two Nonstationary Time Series. Phys. Rev. Lett. 2008, 100, 084102. [Google Scholar] [CrossRef] [Green Version]
- Zebende, G.F. DCCA Cross-Correlation Coefficient: Quantifying Level of Cross-Correlation. Phys. Stat. Mech. Its Appl. 2011, 390, 614–618. [Google Scholar] [CrossRef]
- Liang, Y.; Liu, S.; Zhang, S. Geary Autocorrelation and DCCA Coefficient: Application to Predict Apoptosis Protein Subcellular Localization via PSSM. Phys. Stat. Mech. Its Appl. 2017, 467, 296–306. [Google Scholar] [CrossRef]
- Podobnik, B.; Jiang, Z.-Q.; Zhou, W.-X.; Stanley, H.E. Statistical Tests for Power-Law Cross-Correlated Processes. Phys. Rev. E 2011, 84, 066118. [Google Scholar] [CrossRef] [Green Version]
- Tibshirani, R. Regression Shrinkage and Selection via the Lasso. J. R. Stat. Soc. Ser. B Methodol. 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Friedman, J.; Hastie, T.; Tibshirani, R. Regularization Paths for Generalized Linear Models via Coordinate Descent. J. Stat. Softw. 2010, 33, 1–22. [Google Scholar] [CrossRef] [Green Version]
- Tibshirani, R. Regression Shrinkage and Selection via the Lasso: A Retrospective. J. R. Stat. Soc. Ser. B Stat. Methodol. 2011, 73, 273–282. [Google Scholar] [CrossRef]
- Shrivastava, S.; Jeyanthi, P.M.; Singh, S. Failure Prediction of Indian Banks Using SMOTE, Lasso Regression, Bagging and Boosting. Cogent Econ. Financ. 2020, 8, 1729569. [Google Scholar] [CrossRef]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic Minority Over-Sampling Technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Raghuwanshi, B.S.; Shukla, S. SMOTE Based Class-Specific Extreme Learning Machine for Imbalanced Learning. Knowl.-Based Syst. 2020, 187, 104814. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests. Mach. Learn 2001, 45, 5–32. [Google Scholar] [CrossRef] [Green Version]
- Shar, P.A.; Tao, W.; Gao, S.; Huang, C.; Li, B.; Zhang, W.; Shahen, M.; Zheng, C.; Bai, Y.; Wang, Y. Pred-Binding: Large-Scale Protein–Ligand Binding Affinity Prediction. J. Enzyme Inhib. Med. Chem. 2016, 31, 1443–1450. [Google Scholar] [CrossRef]
- Cao, D.-S.; Zhang, L.-X.; Tan, G.-S.; Xiang, Z.; Zeng, W.-B.; Xu, Q.-S.; Chen, A.F. Computational Prediction of Drug Target Interactions Using Chemical, Biological, and Network Features. Mol. Inform. 2014, 33, 669–681. [Google Scholar] [CrossRef]
- Gornitsky, M.; Velly, A.M.; Mohit, S.; Almajed, M.; Su, H.; Panasci, L.; Schipper, H.M. Altered Levels of Salivary 8-Oxo-7-Hydrodeoxyguanosine in Breast Cancer. JDR Clin. Transl. Res. 2016, 1, 171–177. [Google Scholar] [CrossRef] [PubMed]
- Jeschke, U. Can We Find Breast Cancer via Salivary Fluid Glycosylation Analyses? EBioMedicine 2018, 28, 4. [Google Scholar] [CrossRef] [Green Version]
- Zeitzer, J.M.; Nouriani, B.; Neri, E.; Spiegel, D. Correspondence of Plasma and Salivary Cortisol Patterns in Women with Breast Cancer. Neuroendocrinology 2014, 100, 153–161. [Google Scholar] [CrossRef] [Green Version]
- Bretschneider, N.; Brand, H.; Miller, N.; Lowery, A.J.; Kerin, M.J.; Gannon, F.; Denger, S. Estrogen Induces Repression of the Breast Cancer and Salivary Gland Expression Gene in an Estrogen Receptor Dependent Manner. Cancer Res. 2008, 68, 106–114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- In der Maur, C.D.; Klokman, W.J.; van Leeuwen, F.E.; Tan, I.B.; Rutgers, E.J.T.; Balm, A.J.M. Increased Risk of Breast Cancer Development after Diagnosis of Salivary Gland Tumour. Eur. J. Cancer 2005, 41, 1311–1315. [Google Scholar] [CrossRef]
- Sawczuk, B.; Maciejczyk, M.; Sawczuk-Siemieniuk, M.; Posmyk, R.; Zalewska, A.; Car, H. Salivary Gland Function, Antioxidant Defence and Oxidative Damage in the Saliva of Patients with Breast Cancer: Does the BRCA1 Mutation Disturb the Salivary Redox Profile? Cancers 2019, 11, 1501. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, X.; Yu, H.; Qiao, Y.; Yang, J.; Shu, J.; Zhang, J.; Zhang, Z.; He, J.; Li, Z. Salivary Glycopatterns as Potential Biomarkers for Screening of Early-Stage Breast Cancer. EBioMedicine 2018, 28, 70–79. [Google Scholar] [CrossRef] [Green Version]
- Streckfus, C.F.; Arreola, D.; Edwards, C.; Bigler, L. Salivary Protein Profiles among HER2/Neu-Receptor-Positive and -Negative Breast Cancer Patients: Support for Using Salivary Protein Profiles for Modeling Breast Cancer Progression. J. Oncol. 2012, 2012, 1–9. [Google Scholar] [CrossRef]
- Chang, P.-H.; Hwang-Verslues, W.W.; Chang, Y.-C.; Chen, C.-C.; Hsiao, M.; Jeng, Y.-M.; Chang, K.-J.; Lee, E.Y.-H.P.; Shew, J.-Y.; Lee, W.-H. Activation of Robo1 Signaling of Breast Cancer Cells by Slit2 from Stromal Fibroblast Restrains Tumorigenesis via Blocking PI3K/Akt/ -Catenin Pathway. Cancer Res. 2012, 72, 4652–4661. [Google Scholar] [CrossRef] [Green Version]
- Smit, L.; Berns, K.; Spence, K.; Ryder, W.D.; Zeps, N.; Madiredjo, M.; Beijersbergen, R.; Bernards, R.; Clarke, R.B. An Integrated Genomic Approach Identifies That the PI3K/AKT/FOXO Pathway Is Involved in Breast Cancer Tumor Initiation. Oncotarget 2016, 7, 2596–2610. [Google Scholar] [CrossRef] [Green Version]
- Tao, J.J.; Castel, P.; Radosevic-Robin, N.; Elkabets, M.; Auricchio, N.; Aceto, N.; Weitsman, G.; Barber, P.; Vojnovic, B.; Ellis, H.; et al. Antagonism of EGFR and HER3 Enhances the Response to Inhibitors of the PI3K-Akt Pathway in Triple-Negative Breast Cancer. Sci. Signal. 2014, 7, ra29. [Google Scholar] [CrossRef] [Green Version]
- Gonzalez-Angulo, A.M.; Blumenschein, G.R. Defining Biomarkers to Predict Sensitivity to PI3K/Akt/MTOR Pathway Inhibitors in Breast Cancer. Cancer Treat. Rev. 2013, 39, 313–320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pierobon, M.; Ramos, C.; Wong, S.; Hodge, K.A.; Aldrich, J.; Byron, S.; Anthony, S.P.; Robert, N.J.; Northfelt, D.W.; Jahanzeb, M.; et al. Enrichment of PI3K-AKT–MTOR Pathway Activation in Hepatic Metastases from Breast Cancer. Clin. Cancer Res. 2017, 23, 4919–4928. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chandarlapaty, S.; Sakr, R.A.; Giri, D.; Patil, S.; Heguy, A.; Morrow, M.; Modi, S.; Norton, L.; Rosen, N.; Hudis, C.; et al. Frequent Mutational Activation of the PI3K-AKT Pathway in Trastuzumab-Resistant Breast Cancer. Clin. Cancer Res. 2012, 18, 6784–6791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gris-Oliver, A.; Palafox, M.; Monserrat, L.; Brasó-Maristany, F.; Òdena, A.; Sánchez-Guixé, M.; Ibrahim, Y.H.; Villacampa, G.; Grueso, J.; Parés, M.; et al. Genetic Alterations in the PI3K/AKT Pathway and Baseline AKT Activity Define AKT Inhibitor Sensitivity in Breast Cancer Patient-Derived Xenografts. Clin. Cancer Res. 2020, 26, 3720–3731. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ramaswamy, B.; Lu, Y.; Teng, K.-Y.; Nuovo, G.; Li, X.; Shapiro, C.L.; Majumder, S. Hedgehog Signaling Is a Novel Therapeutic Target in Tamoxifen-Resistant Breast Cancer Aberrantly Activated by PI3K/AKT Pathway. Cancer Res. 2012, 72, 5048–5059. [Google Scholar] [CrossRef] [Green Version]
- Le Rhun, E.; Bertrand, N.; Dumont, A.; Tresch, E.; Le Deley, M.-C.; Mailliez, A.; Preusser, M.; Weller, M.; Revillion, F.; Bonneterre, J. Identification of Single Nucleotide Polymorphisms of the PI3K-AKT-MTOR Pathway as a Risk Factor of Central Nervous System Metastasis in Metastatic Breast Cancer. Eur. J. Cancer 2017, 87, 189–198. [Google Scholar] [CrossRef] [Green Version]
- Yi, Y.W.; Hong, W.; Kang, H.J.; Kim, H.J.; Zhao, W.; Wang, A.; Seong, Y.-S.; Bae, I. Inhibition of the PI3K/AKT Pathway Potentiates Cytotoxicity of EGFR Kinase Inhibitors in Triple-Negative Breast Cancer Cells. J. Cell. Mol. Med. 2013, 17, 648–656. [Google Scholar] [CrossRef]
- Yang, S.X.; Polley, E.; Lipkowitz, S. New Insights on PI3K/AKT Pathway Alterations and Clinical Outcomes in Breast Cancer. Cancer Treat. Rev. 2016, 45, 87–96. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cavazzoni, A.; Bonelli, M.A.; Fumarola, C.; La Monica, S.; Airoud, K.; Bertoni, R.; Alfieri, R.R.; Galetti, M.; Tramonti, S.; Galvani, E.; et al. Overcoming Acquired Resistance to Letrozole by Targeting the PI3K/AKT/MTOR Pathway in Breast Cancer Cell Clones. Cancer Lett. 2012, 323, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Riggio, M.; Polo, M.L.; Blaustein, M.; Colman-Lerner, A.; Lüthy, I.; Lanari, C.; Novaro, V. PI3K/AKT Pathway Regulates Phosphorylation of Steroid Receptors, Hormone Independence and Tumor Differentiation in Breast Cancer. Carcinogenesis 2012, 33, 509–518. [Google Scholar] [CrossRef] [Green Version]
- Khan, M.A.; Jain, V.K.; Rizwanullah, M.; Ahmad, J.; Jain, K. PI3K/AKT/MTOR Pathway Inhibitors in Triple-Negative Breast Cancer: A Review on Drug Discovery and Future Challenges. Drug Discov. Today 2019, 24, 2181–2191. [Google Scholar] [CrossRef] [PubMed]
- Sharma, V.; Sharma, A.K.; Punj, V.; Priya, P. Recent Nanotechnological Interventions Targeting PI3K/Akt/MTOR Pathway: A Focus on Breast Cancer. Semin. Cancer Biol. 2019, 59, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Delaloge, S.; DeForceville, L. Targeting PI3K/AKT Pathway in Triple-Negative Breast Cancer. Lancet Oncol. 2017, 18, 1293–1294. [Google Scholar] [CrossRef]
- Basho, R.K.; Gilcrease, M.; Murthy, R.K.; Helgason, T.; Karp, D.D.; Meric-Bernstam, F.; Hess, K.R.; Herbrich, S.M.; Valero, V.; Albarracin, C.; et al. Targeting the PI3K/AKT/MTOR Pathway for the Treatment of Mesenchymal Triple-Negative Breast Cancer: Evidence From a Phase 1 Trial of MTOR Inhibition in Combination With Liposomal Doxorubicin and Bevacizumab. JAMA Oncol. 2017, 3, 509. [Google Scholar] [CrossRef]
- Ciruelos Gil, E.M. Targeting the PI3K/AKT/MTOR Pathway in Estrogen Receptor-Positive Breast Cancer. Cancer Treat. Rev. 2014, 40, 862–871. [Google Scholar] [CrossRef]
- Costa, R.L.B.; Han, H.S.; Gradishar, W.J. Targeting the PI3K/AKT/MTOR Pathway in Triple-Negative Breast Cancer: A Review. Breast Cancer Res. Treat. 2018, 169, 397–406. [Google Scholar] [CrossRef]
- Fengjiao, J.; Zhaozhen, W.; Xiao, H.; Jiahui, Z.; Zihe, G.; Xiao, H.; Junfang, Q.; Chen, L.; Yue, W. The PI3K/Akt/GSK-3β/ROS/EIF2B Pathway Promotes Breast Cancer Growth and Metastasis via Suppression of NK Cell Cytotoxicity and Tumor Cell Susceptibility. Cancer Biol. Med. 2019, 16, 38. [Google Scholar] [CrossRef] [Green Version]
- Paplomata, E.; O’Regan, R. The PI3K/AKT/MTOR Pathway in Breast Cancer: Targets, Trials and Biomarkers. Ther. Adv. Med. Oncol. 2014, 6, 154–166. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Massihnia, D.; Galvano, A.; Fanale, D.; Perez, A.; Castiglia, M.; Incorvaia, L.; Listì, A.; Rizzo, S.; Cicero, G.; Bazan, V.; et al. Triple Negative Breast Cancer: Shedding Light onto the Role of Pi3k/Akt/Mtor Pathway. Oncotarget 2016, 7, 60712–60722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Woo, S.-U.; Sangai, T.; Akcakanat, A.; Chen, H.; Wei, C.; Meric-Bernstam, F. Vertical Inhibition of the PI3K/Akt/MTOR Pathway Is Synergistic in Breast Cancer. Oncogenesis 2017, 6, e385. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wen, M.; Zhang, Z.; Niu, S.; Sha, H.; Yang, R.; Yun, Y.; Lu, H. Deep-Learning-Based Drug–Target Interaction Prediction. J. Proteome Res. 2017, 16, 1401–1409. [Google Scholar] [CrossRef]
- Ding, R.; Shi, J.; Pabon, K.; Scotto, K.W. Xanthines Down-Regulate the Drug Transporter ABCG2 and Reverse Multidrug Resistance. Mol. Pharmacol. 2012, 81, 328–337. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Models | NR | GPCR | IC | E |
|---|---|---|---|---|
| NetCBP [26] | 0.8394 | 0.8235 | 0.8034 | 0.8251 |
| Huang et al. [27] | 0.9634 | 0.9053 | 0.9382 | 0.9601 |
| Bigram-PSSM [21] | 0.8690 | 0.8720 | 0.8890 | 0.9480 |
| iDTI-ESBoost [20] | 0.9285 | 0.9322 | 0.9369 | 0.9689 |
| Li et al. [28] | 0.9300 | 0.9171 | 0.8856 | 0.9288 |
| KBMF2K [29] | 0.8240 | 0.8570 | 0.7990 | 0.8320 |
| NRLMF [30] | 0.9500 | 0.9690 | 0.9890 | 0.9870 |
| PsePDC-DTIs | 0.9886 | 0.9923 | 0.9956 | 0.9983 |
| Models | NR | GPCR | IC | E |
|---|---|---|---|---|
| Bigram-PSSM [21] | 0.4110 | 0.2820 | 0.3900 | 0.5460 |
| iDTI-ESBoost [20] | 0.7900 | 0.5000 | 0.4800 | 0.6800 |
| NRLMF [30] | 0.7280 | 0.7490 | 0.9060 | 0.8920 |
| PsePDC-DTIs | 0.9875 | 0.9923 | 0.9958 | 0.9984 |
| Drug | Drug_Name | Target | Target_Name | Prob |
|---|---|---|---|---|
| DB00201 | Caffeine | hsa3783 | KCNN4 | 0.988 |
| DB00277 | Theophylline | hsa3783 | KCNN4 | 0.982 |
| DB01412 | Theobromine | hsa3783 | KCNN4 | 0.93 |
| DB00530 | Erlotinib | hsa238 | ALK | 0.886 |
| DB00806 | Pentoxifylline | hsa3783 | KCNN4 | 0.884 |
| DB00824 | Enprofylline | hsa3783 | KCNN4 | 0.866 |
| DB00530 | Erlotinib | hsa2263 | FGFR2 | 0.864 |
| DB00661 | Verapamil | hsa57719 | ANO8 | 0.846 |
| DB01303 | Oxtriphylline | hsa3783 | KCNN4 | 0.844 |
| DB08916 | Afatinib | hsa2263 | FGFR2 | 0.806 |
| Datasets | Drugs | Targets | Interactions | Positive Samples | Negative Samples | Sample Ratio |
|---|---|---|---|---|---|---|
| Enzyme | 445 | 664 | 2926 | 2926 | 292,554 | 99.98 |
| IC | 210 | 204 | 1476 | 1476 | 41,364 | 28.02 |
| GPCR | 223 | 95 | 635 | 635 | 20,550 | 32.36 |
| NR | 54 | 26 | 90 | 90 | 1314 | 14.60 |
| Total | 932 | 989 | 5127 | 5127 | 355,782 | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Song, J.; Xu, Z.; Cao, L.; Wang, M.; Hou, Y.; Li, K. The Discovery of New Drug-Target Interactions for Breast Cancer Treatment. Molecules 2021, 26, 7474. https://doi.org/10.3390/molecules26247474
Song J, Xu Z, Cao L, Wang M, Hou Y, Li K. The Discovery of New Drug-Target Interactions for Breast Cancer Treatment. Molecules. 2021; 26(24):7474. https://doi.org/10.3390/molecules26247474
Chicago/Turabian StyleSong, Jiali, Zhenyi Xu, Lei Cao, Meng Wang, Yan Hou, and Kang Li. 2021. "The Discovery of New Drug-Target Interactions for Breast Cancer Treatment" Molecules 26, no. 24: 7474. https://doi.org/10.3390/molecules26247474





