Computational Design of Macrocyclic Binders of S100B(ββ): Novel Peptide Theranostics
Abstract
:1. Introduction
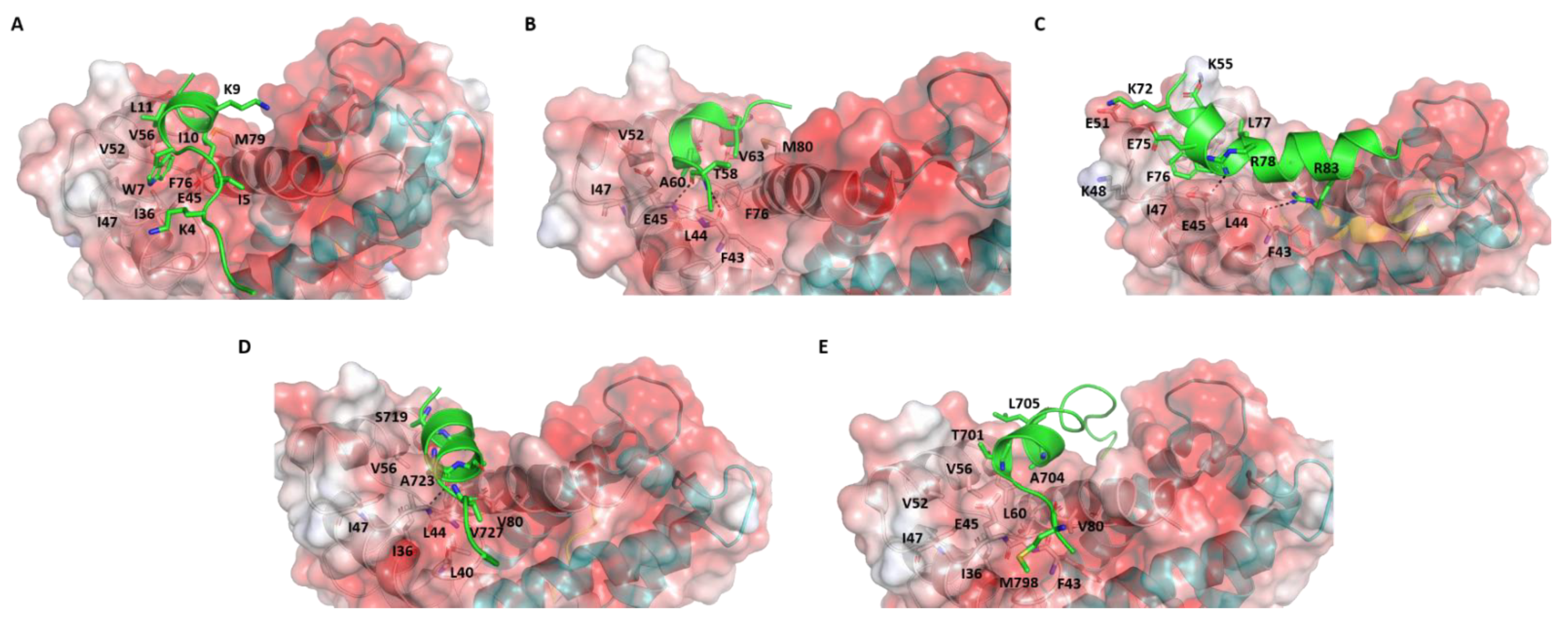
2. Comparative Analysis of S100B(ββ)-Peptide Complexes
3. Methods
Molecular Dynamics (MD) Simulations
4. Results and Discussion
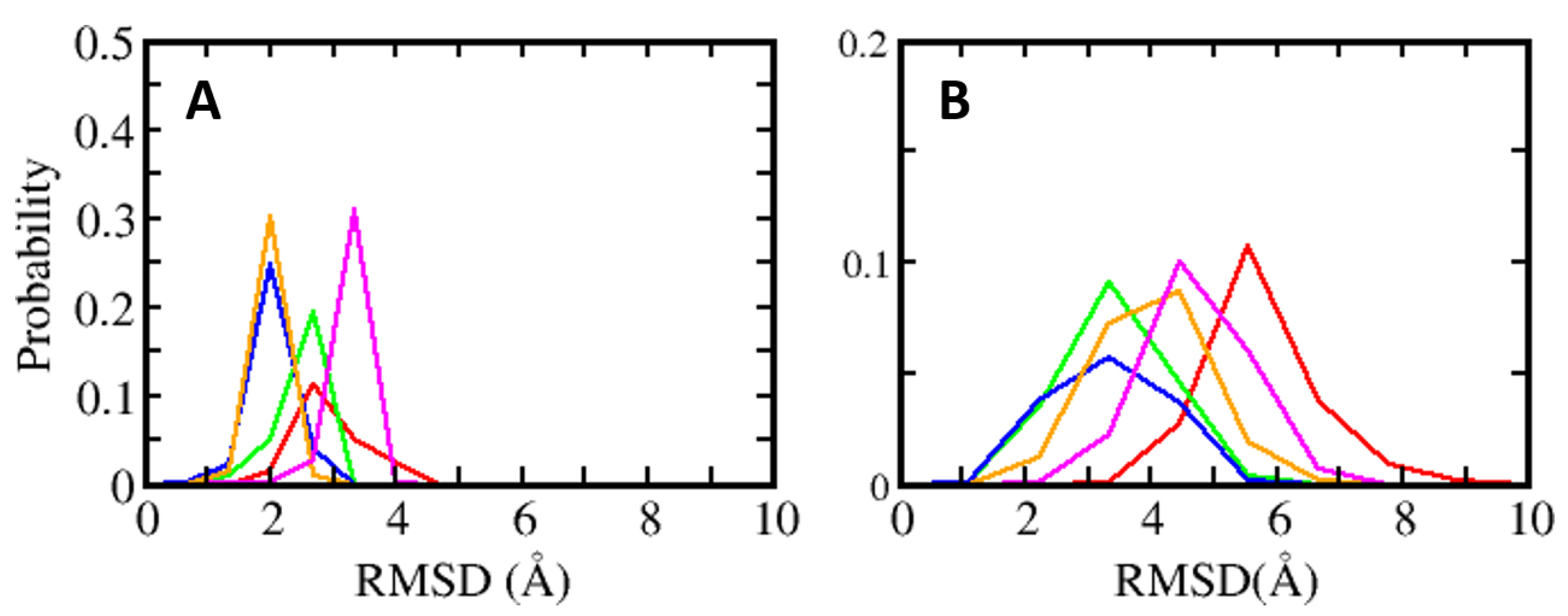
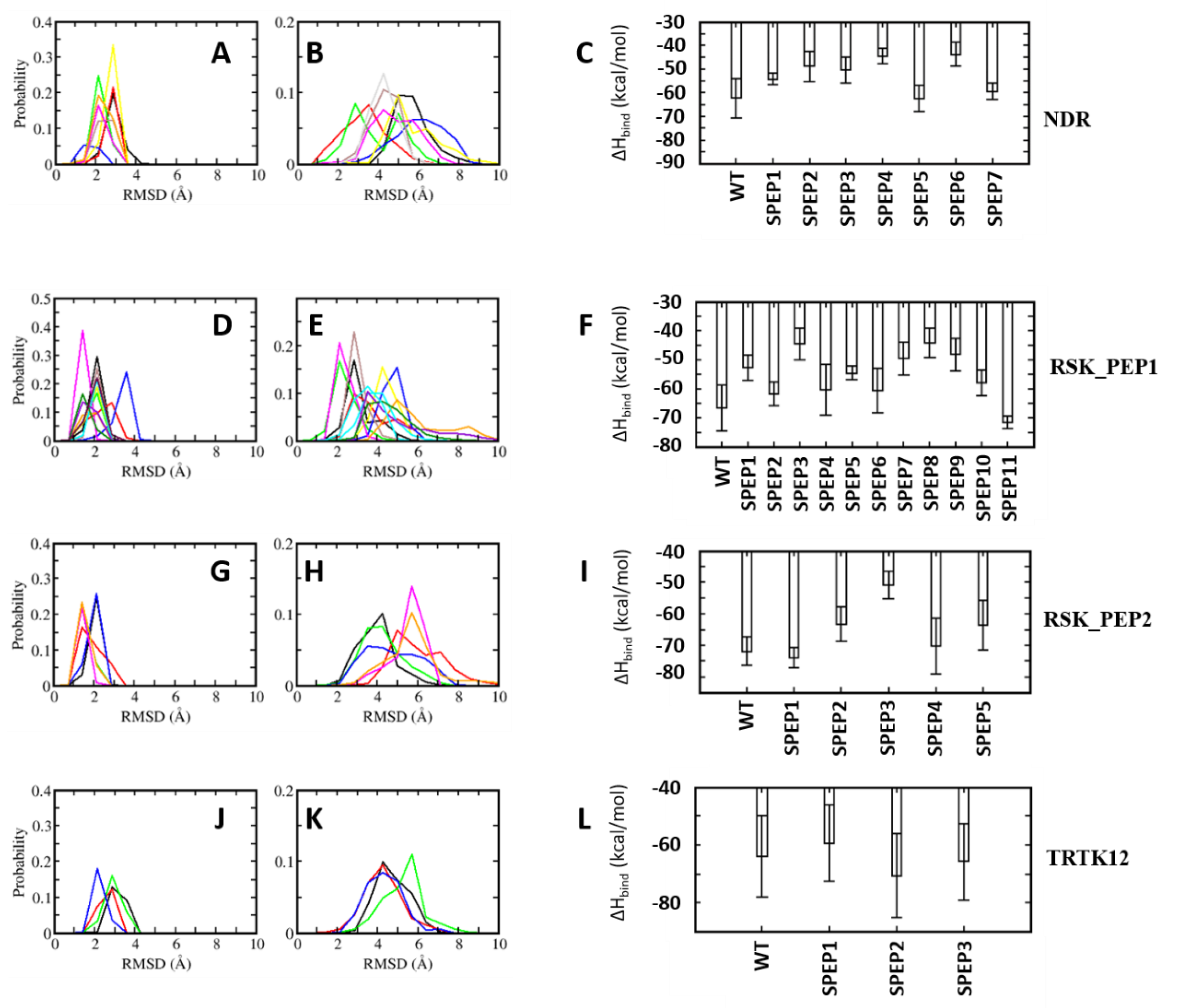
4.1. Conformational Dynamics of S100B(ββ)-Peptide Complexes
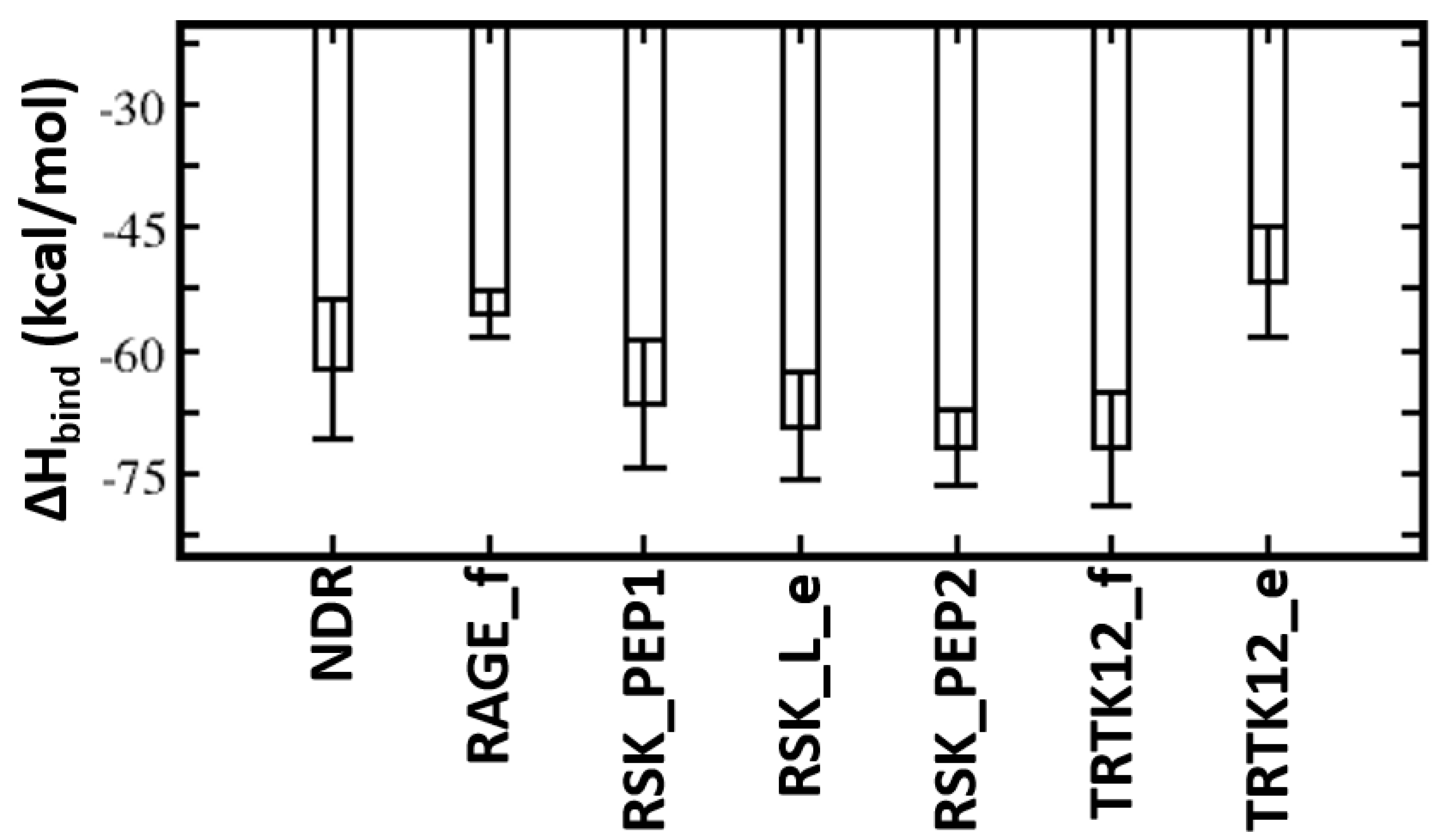
4.2. Energetics of S100B(ββ)–Peptide Complexes
4.3. In Silico Optimization of S100B(ββ) Binding Peptides/Design of Consensus Peptide
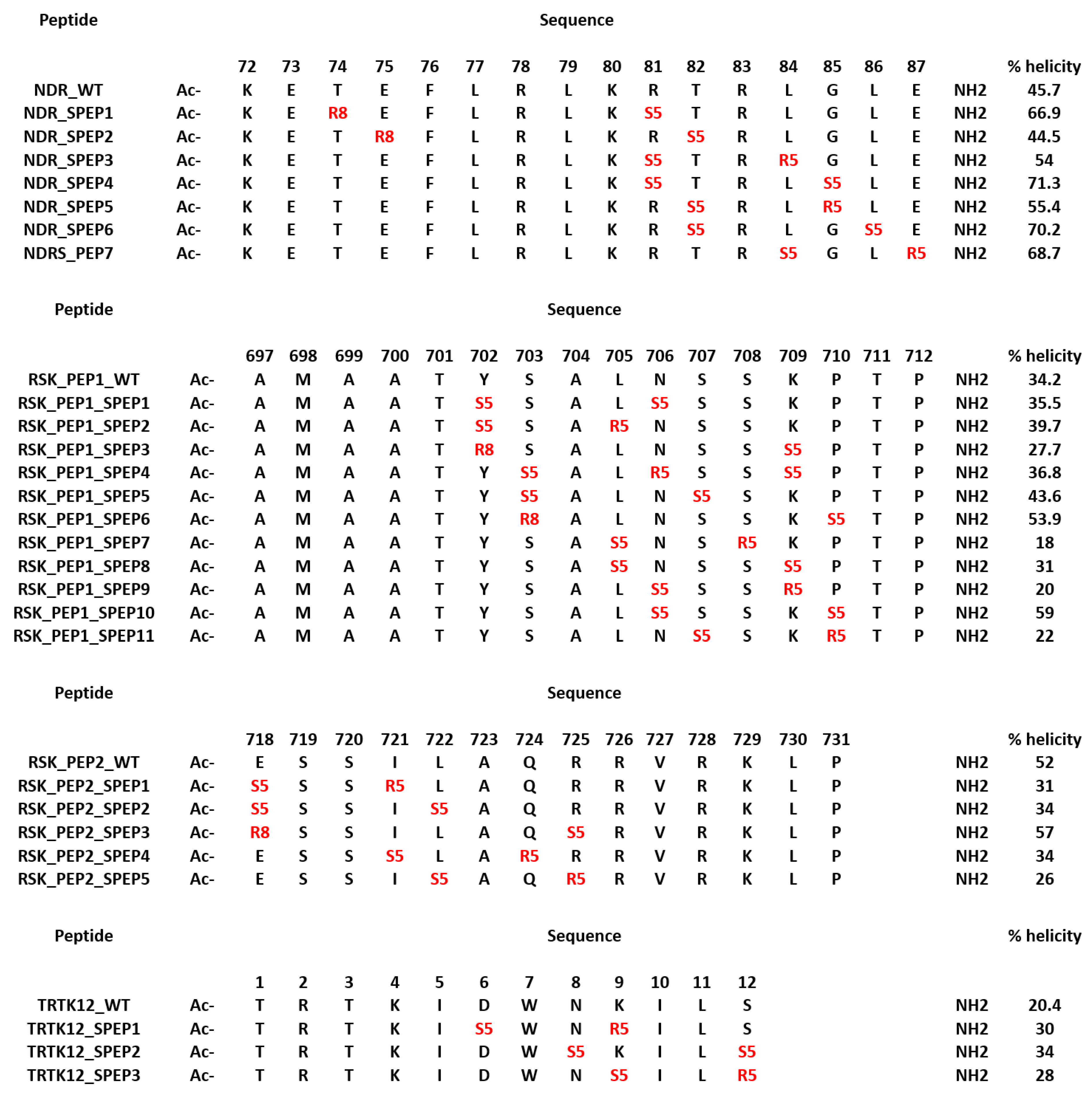
4.4. Design of Stapled Peptide Binders of S100B(ββ)
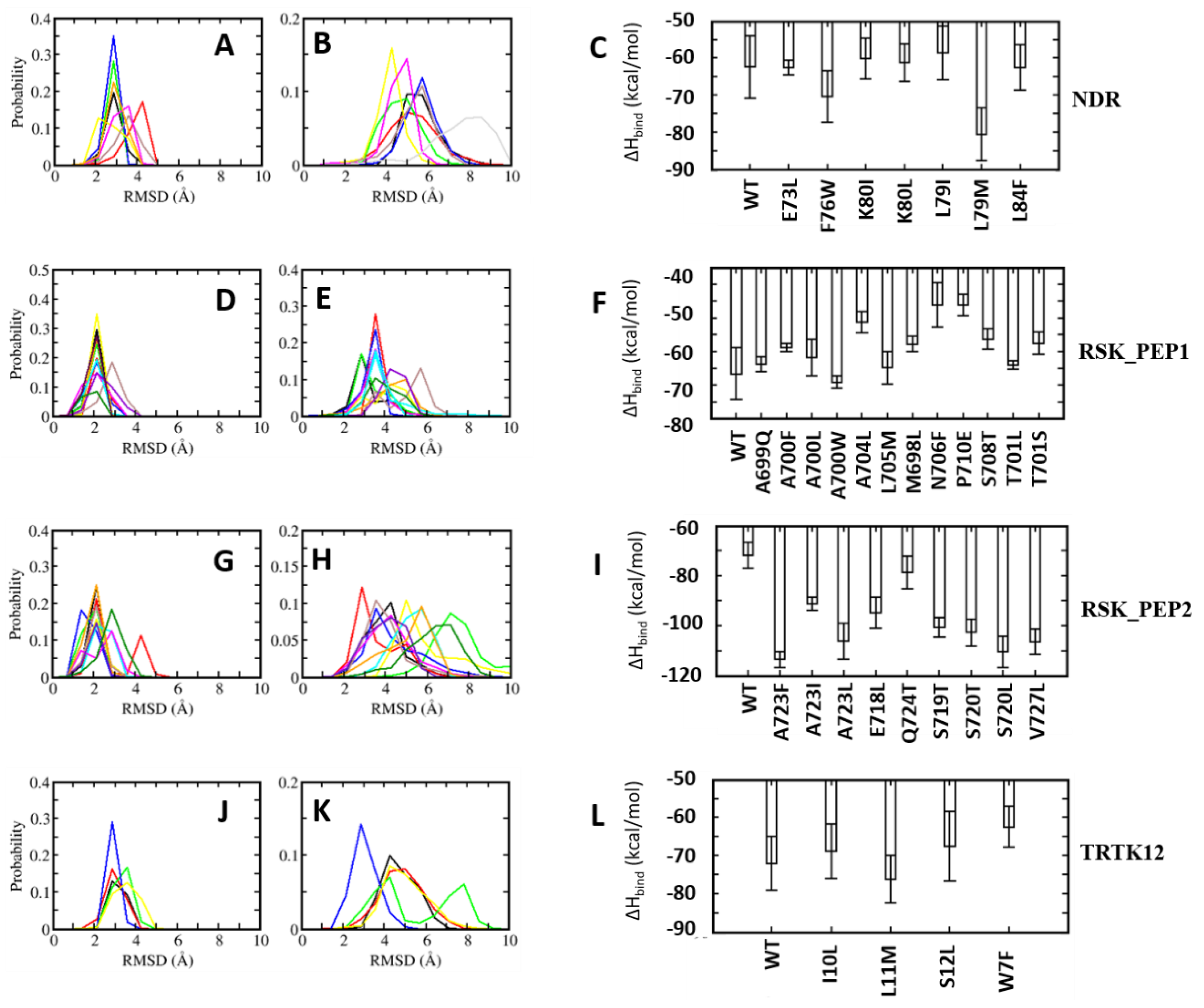
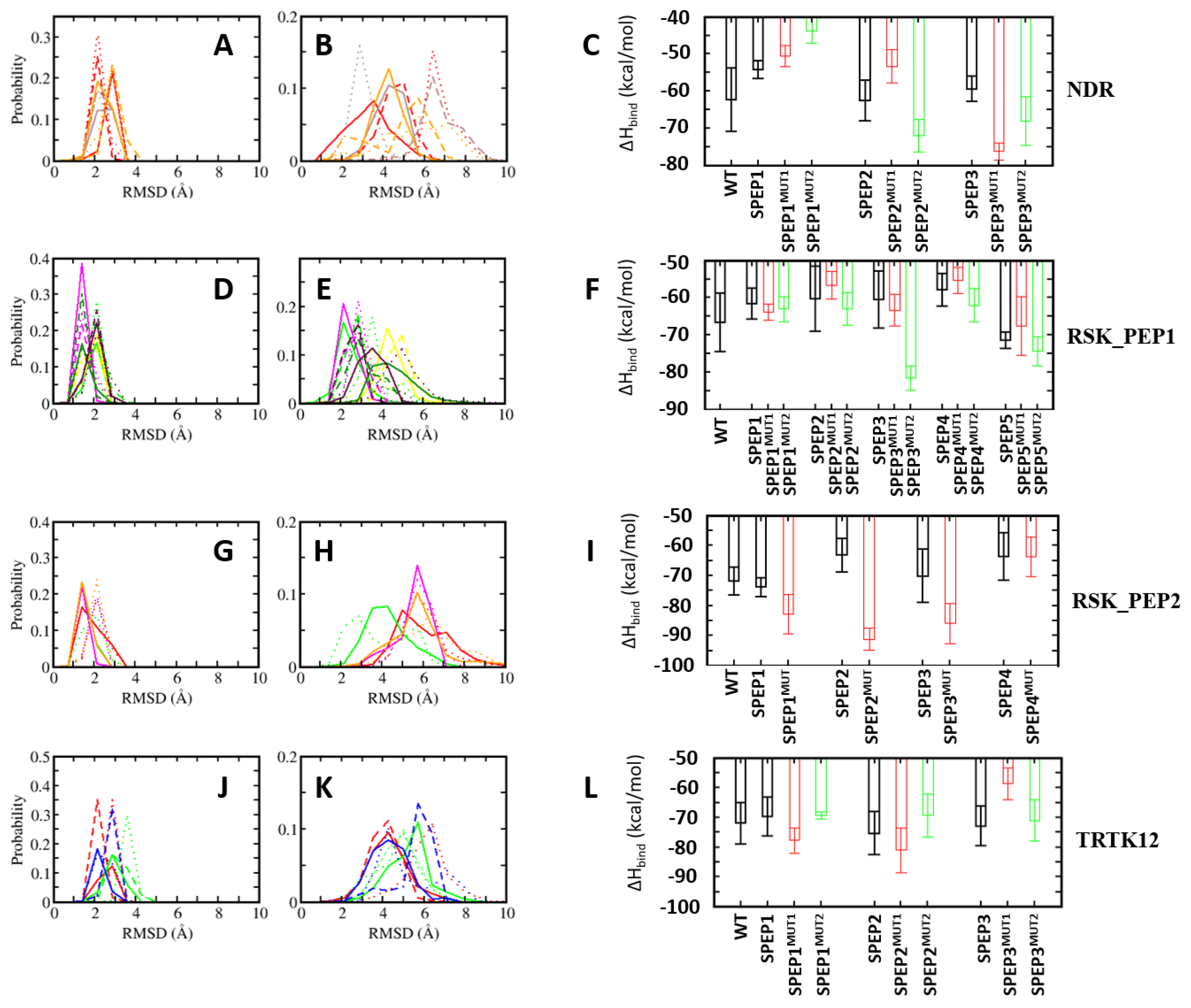
4.5. Optimization of Stapled Peptides through In Silico Mutagenesis
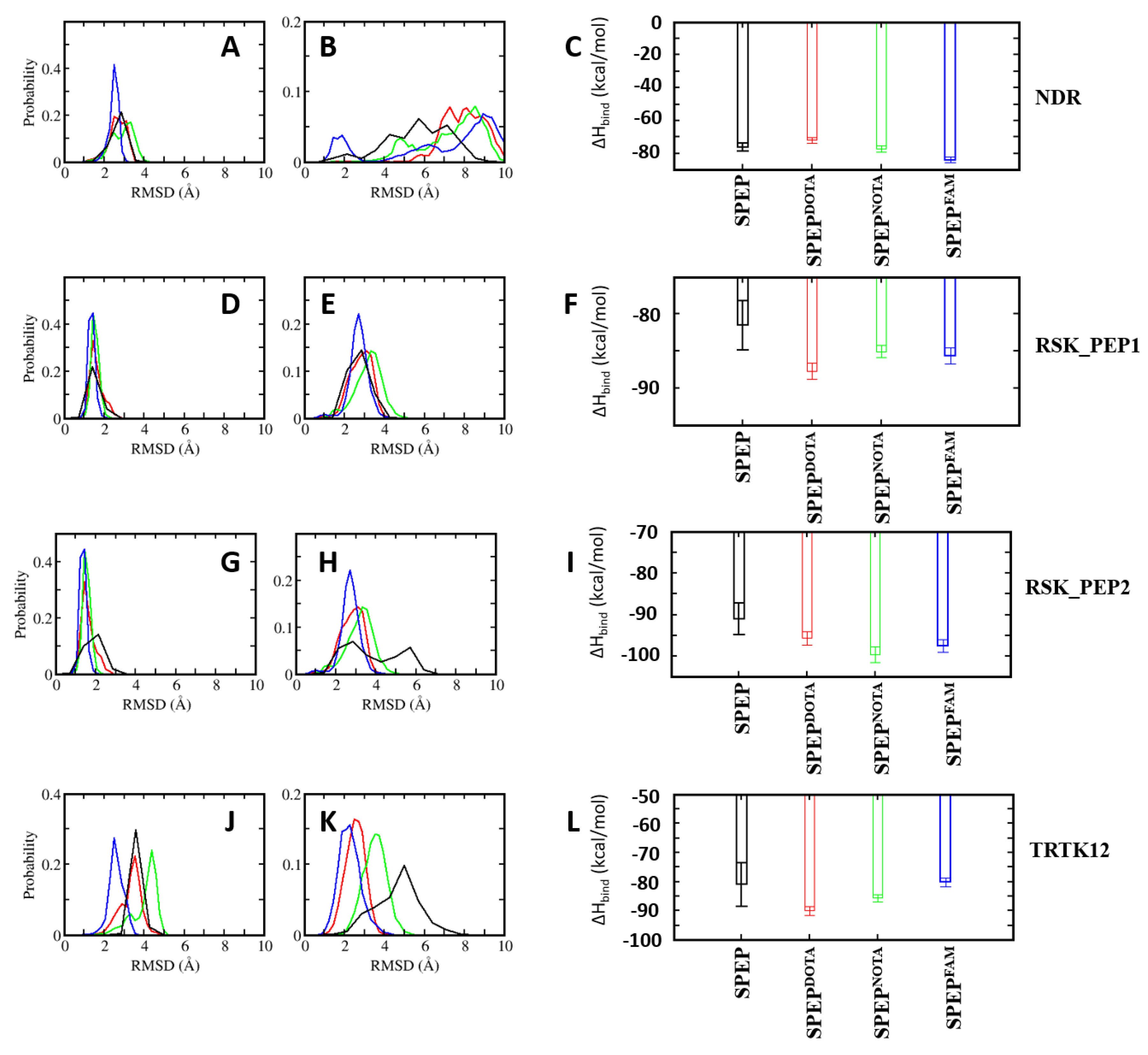
4.6. Design of Stapled Peptide-Based Imaging Probes for Detection of Biomarker S100B(ββ)
5. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Donato, R.; Sorci, G.; Riuzzi, F.; Arcuri, C.; Bianchi, R.; Brozzi, F.; Tubaro, C.; Giambanco, I. S100B’s double life: Intracellular regulator and extracellular signal. Biochi. Biophys. Acta. 2009, 1793, 1008–1022. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Riuzzi, F.; Sorci, G.; Donato, R. S100B stimulates myoblast proliferation and inhibits myoblast differentiation by independently stimulating ERK1/2 and inhibiting p38 MAPK. J. Cell Physiol. 2006, 207, 461–470. [Google Scholar] [CrossRef] [PubMed]
- Arcuri, C.; Bianchi, R.; Brozzi, F.; Donato, R. S100B increases proliferation in PC12 neuronal cells and reduces their responsiveness to nerve growth factor via Akt activation. J. Biol. Chem. 2005, 280, 4402–4414. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kligman, D.; Hilt, D.C. The S100 protein family. Trends Biochem. Sci. 1988, 13, 437–443. [Google Scholar] [CrossRef]
- Marenholz, I.; Heizmann, C.W.; Fritz, G. S100 proteins in mouse and man: From evolution to function and pathology (including an update of the nomenclature). Biochem. Biophys. Res. Commun. 2004, 322, 1111–1122. [Google Scholar] [CrossRef]
- Donato, R. S100: A multigenic family of calcium-modulated proteins of the EF-hand type with intracellular and extracellular functional roles. Int. J. Biochem. Cell Biol. 2001, 33, 637–668. [Google Scholar] [CrossRef]
- Donato, R. Perspectives in S-100 protein biology. Cell Calcium 1991, 12, 713–726. [Google Scholar] [CrossRef]
- Zimmer, D.B.; Cornwall, E.H.; Landar, A.; Song, W. The S100 protein family: History, function, and expression. Brain Res. Bull. 1995, 37, 417–429. [Google Scholar] [CrossRef]
- Castets, F.; Griffin, W.S.; Marks, A.; Van Eldik, L.J. Transcriptional regulation of the human S100 beta gene. Brain Res. Mol. Brain Res. 1997, 46, 208–216. [Google Scholar] [CrossRef]
- Drohat, A.C.; Nenortas, E.; Beckett, D.; Weber, D.J. Oligomerization state of S100B(ββ) at nanomolar concentration determined by large-zone analytical gel filtration chromatography. Protein Sci. 1997, 6, 1577–1582. [Google Scholar] [CrossRef] [Green Version]
- Amburgey, J.C.; Abildgaard, F.; Starich, M.R.; Shah, S.; Hilt, D.C.; Weber, D.J. 1H, 13C, 15N NMR assignments and solution secondary structure of rat Apo-S100β. J. Biomol. NMR. 1995, 6, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Drohat, A.C.; Amburgey, J.C.; Abildgaard, F.; Starich, M.R.; Baldisseri, D.; Weber, D.J. Solution structure of rat apo-S100(ββ) as determined by NMR spectroscopy. Biochemistry 1996, 35, 11577–11588. [Google Scholar] [CrossRef] [PubMed]
- Drohat, A.C.; Tjandra, N.; Baldisseri, D.M.; Weber, D.J. The use of dipolar couplings for determining the solution structure of rat apo-S100B(ββ). Protein Sci. 1999, 8, 800–809. [Google Scholar] [CrossRef] [PubMed]
- Kilby, P.M.; Van Eldik, L.J.; Roberts, G.C.K. The solution structure of the bovine S100B protein dimer in the calcium-free state. Structure 1996, 4, 1041–1052. [Google Scholar] [CrossRef] [Green Version]
- Thulin, E.; Kesvatera, T.; Linse, S. Molecular determinants of S100B oligomer formation. PLoS ONE 2011, 6, e14768. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sorci, G.; Riuzzi, F.; Arcuri, C.; Tubaro, C.; Bianchi, R.; Giambanco, I.; Donato, R. S100B protein in tissue development, repair and regeneration. World J. Boil. Chem. 2013, 4, 1–12. [Google Scholar] [CrossRef]
- Kretz, A.L.; von Karstedt, S.; Wittau, M.; Henne-Bruns, D.; Lemke, J. Friend or Foe: S100 Proteins in Cancer. Cancers 2020, 12, 2037. [Google Scholar]
- Baudier, J.; Gentil, B.J. The S100B Protein and Partners in Adipocyte Response to Cold Stress and Adaptive Thermogenesis: Facts, Hypotheses, and Perspectives. Biomolecules 2020, 10, 843. [Google Scholar] [CrossRef]
- Xiong, T.F.; Pan, F.Q.; Li, D. Expression and clinical significance of S100 family genes in patients with melanoma. Melanoma Res. 2019, 29, 23–29. [Google Scholar] [CrossRef]
- Frauchiger, A.L.; Dummer, R.; Mangana, J. Serum S100B levels in melanoma. Methods Mol. Biol. 2019, 1929, 691–700. [Google Scholar]
- Undén, J.; Ingebrigtsen, T.; Romner, B. Scandinavian guidelines for initial management of minimal, mild and moderate head injuries in adults: An evidence and consensus-based update. BMC Med. 2013, 11, 50. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jones, C.M.C.; Harmon, C.; McCann, M.; Gunyan, H.; Bazarian, J.J. S100B outperforms clinical decision rules for the identification of intracranial injury on head CT scan after mild traumatic brain injury. Brain Inj. 2020, 34, 407–414. [Google Scholar] [CrossRef] [PubMed]
- Vedin, T.; Svensson, S.; Edelhamre, M.; Karlsson, M.; Bergenheim, M.; Larsson, P.A. Management of mild traumatic brain injury-trauma energy level and medical history as possible predictors for intracranial hemorrhage. Eur. J. Trauma Emerg. Surg. 2019, 45, 901–907. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bai, Y.; Li, L.D.; Li, J.; Lu, X. Prognostic values of S100 family members in ovarian cancer patients. BMC Cancer. 2018, 18, 1256. [Google Scholar] [CrossRef] [PubMed]
- Yen, M.C.; Huang, Y.C.; Kan, J.Y.; Kuo, P.L.; Hou, M.F.; Hsu, Y.L. S100B expression in breast cancer as a predictive marker for cancer metastasis. Int. J. Oncol. 2018, 52, 433–440. [Google Scholar] [CrossRef] [Green Version]
- Wang, T.; Huo, X.; Chong, Z.; Khan, H.; Liu, R. A review of S100 protein family in lung cancer. Clin. Chim. Acta 2018, 476, 54–59. [Google Scholar] [CrossRef]
- Kondrup, M.; Nygaard, A.D.; Madsen, J.S.; Bechmann, T. S100B as a biomarker for brain metastases in patients with non-small cell lung cancer. Biomed. Rep. 2020, 12, 204–208. [Google Scholar] [CrossRef] [Green Version]
- Bresnick, A.R.; Weber, D.J.; Zimmer, D.B. S100 proteins in cancer. Nat. Rev. Cancer. 2015, 15, 96–109. [Google Scholar] [CrossRef] [Green Version]
- Rustandi, R.R.; Drohat, A.C.; Baldisseri, D.M.; Wilder, P.T.; Weber, D.J. The Ca2+-dependent interaction of S100B(ββ) with a peptide derived from p53. Biochemistry 1998, 37, 1951–1960. [Google Scholar] [CrossRef]
- Lin, J.; Blake, M.; Tang, C.; Zimmer, D.; Rustandi, R.R.; Weber, D.J.; Carrier, F. Inhibition of p53 transcriptional activity by the S100B calcium-binding protein. J. Biol. Chem. 2001, 276, 35037–35041. [Google Scholar] [CrossRef] [Green Version]
- Wilder, P.T.; Rustandi, R.R.; Drohat, A.C.; Weber, D.J. S100B (ββ) inhibits the protein kinase C-dependent phosphorylation of a peptide derived from p53 in a Ca2+-dependent manner. Protein Sci. 1998, 7, 794–798. [Google Scholar] [CrossRef] [PubMed]
- Baudier, J.; Delphin, C.; Grunwald, D.; Khochbin, S.; Lawrence, J.J. Characterization of the tumor suppressor protein p53 as a protein kinase C substrate and a S100b-binding protein. Proc. Natl. Acad. Sci. USA 1992, 89, 11627–11631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, J.; Yang, Q.; Wilder, P.T.; Carrier, F.; Weber, D.J. The calcium-binding protein S100B down-regulates p53 and apoptosis in malignant melanoma. J. Biol. Chem. 2010, 285, 27487–27498. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, H.; Zhang, L.; Zhang, I.Y.; Chen, X.; Da Fonseca, A.; Wu, S.; Ren, H.; Badie, S.; Sadeghi, S.; Ouyang, M.; et al. S100B Promotes Glioma Growth through Chemoattraction of Myeloid-Derived Macrophages. Clin. Cancer Res. 2013, 19, 3764–3775. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, J.; Yang, Q.; Yan, Z.; Markowitz, J.; Wilder, P.T.; Carrier, F.; Weber, D.J. Inhibiting S100B restores p53 levels in primary malignant melanoma cancer cells. J. Biol. Chem. 2004, 279, 34071–34077. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Webster, M.R.; Fane, M.E.; Alicea, G.M.; Basu, S.; Kossenkov, A.V.; Marino, G.E.; Douglass, S.M.; Kaur, A.; Ecker, B.L.; Gnanapradeepan, K.; et al. Paradoxical role for wild-type p53 in driving therapy resistance in melanoma. Mol. Cell. 2020, 77, 633–644. [Google Scholar] [CrossRef]
- Hartman, K.G.; Vitolo, M.I.; Pierce, A.D.; Fox, J.M.; Shapiro, P.; Martin, S.S.; Wilder, P.T.; Weber, D.J. Complex formation between S100B protein and the p90 ribosomal S6 kinase (RSK) in malignant melanoma is calcium-dependent and inhibits extracellular signal-regulated kinase (ERK)-mediated phosphorylation of RSK. J. Biol. Chem. 2014, 289, 12886–12895. [Google Scholar] [CrossRef] [Green Version]
- Bianchi, R.; Kastrisianaki, E.; Giambanco, I.; Donato, R. S100B protein stimulates microglia migration via RAGE-dependent up-regulation of chemokine expression and release. J. Biol. Chem. 2011, 286, 7214–7226. [Google Scholar] [CrossRef] [Green Version]
- Zimmer, D.B.; Lapidus, R.G.; Weber, D.J. In vivo screening of S100B inhibitors for melanoma therapy. Methods Mol. Biol. 2013, 963, 303–317. [Google Scholar]
- McKnight, L.E.; Raman, E.P.; Bezawada, P.; Kudrimoti, S.; Wilder, P.T.; Hartman, K.G.; Godoy-Ruiz, R.; Toth, E.A.; Coop, A.; MacKerell, A.D., Jr.; et al. Structure-Based Discovery of a Novel Pentamidine-Related Inhibitor of the Calcium-Binding Protein S100B. ACS Med. Chem. Lett. 2012, 3, 975–979. [Google Scholar] [CrossRef]
- Cavalier, M.C. Small Molecule Inhibitors of Ca2+-S100B Reveal Two Protein Conformations. J. Med. Chem. 2016, 59, 592–608. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cavalier, M.C.; Pierce, A.D.; Wilder, P.T.; Alasady, M.J.; Hartman, K.G.; Neau, D.B.; Foley, T.L.; Jadhav, A.; Maloney, D.J.; Simeonov, A.; et al. Covalent small molecule inhibitors of Ca2+-bound S100B. Biochemistry 2014, 53, 6628–6640. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bakail, M.; Ocshsenbein, F. Targeting protein–protein interactions, a wide open field for drug design. Comptes Rendus Chim. 2016, 19, 19–27. [Google Scholar] [CrossRef] [Green Version]
- Arkin, M.R.; Tang, Y.; Wells, J.A. Small-molecule inhibitors of protein-protein interactions: Progressing towards the reality. Chem. Biol. 2014, 21, 1102–1114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wojcik, P.; Berlick, L. Peptide-based inhibitors of protein-protein interactions. Bioorg. Med. Chem. Lett. 2016, 26, 707–710. [Google Scholar] [CrossRef]
- Lau, J.L.; Dunn, M.K. Therapeutic peptides: Historical perspectives, current development trends, and future directions. Bioorg. Med. Chem. 2018, 26, 2700–2707. [Google Scholar] [CrossRef]
- Fosgerau, K.; Hoffmann, T. Peptide therapeutics: Current status and future directions. Drug Discov. Today 2015, 20, 122–128. [Google Scholar] [CrossRef] [Green Version]
- Walensky, L.D.; Bird, G.H. Hydrocarbon-stapled peptides: Principles, practice, and progress. J. Med. Chem. 2014, 57, 6275–6288. [Google Scholar] [CrossRef] [Green Version]
- Henninot, A.; Collins, J.C.; Nuss, J.M. The current state of peptide drug discovery: Back to the future? J. Med. Chem. 2018, 61, 1382–1414. [Google Scholar] [CrossRef]
- Klein, M. Stabilized helical peptides: Overview of the technologies and its impact on drug discovery. Expert Opin. Drug Discov. 2017, 12, 1117–1125. [Google Scholar] [CrossRef]
- Sawyer, T.K.; Partridge, A.W.; Kaan, H.Y.; Juang, Y.C.; Lim, S.; Johannes, C.; Yuen, T.Y.; Verma, C.; Kannan, S.; Aronica, P.; et al. Macrocyclic α helical peptide therapeutic modality: A perspective of learnings and challenges. Bioorg. Med. Chem. 2018, 26, 2807–2815. [Google Scholar] [CrossRef] [PubMed]
- Wiedmann, M.M.; Tan, Y.S.; Wu, Y.; Aibara, S.; Xu, W.; Sore, H.F.; Verma, C.S.; Itzhaki, L.; Stewart, M.; Brenton, J.D.; et al. Development of Cell-Permeable, Non-Helical Constrained Peptides to Target a Key Protein–Protein Interaction in Ovarian Cancer. Angew. Chem. Int. Ed. 2017, 56, 524–529. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bluntzer, M.T.J.; O’Connell, J.; Baker, T.S.; Michel, J.; Hulme, A.N. Designing stapled peptides to inhibit protein-protein interactions. Pept. Sci. 2020, 113, e24191. [Google Scholar]
- Rustandi, R.R.; Baldisseri, D.M.; Weber, D.J. Structure of the negative regulatory domain of p53 bound to S100B(ββ). Nat. Struct. Biol. 2000, 7, 570–574. [Google Scholar]
- Bhattacharya, S.; Large, E.; Heizmann, C.W.; Hemmings, B.A.; Chazin, W.J. Structure of the Ca2+/S100B/NDR Kinase Peptide Complex: Insights into S100 Target Specificity and Activation of the Kinase. Biochemistry 2003, 42, 14416–14426. [Google Scholar] [CrossRef]
- Ivanenkov, V.V.; Jamieson, G.A., Jr.; Gruenstein, E.; Dimlich, R.V. Characterization of S-100b binding epitopes. Identification of a novel target, the actin capping protein, CapZ. J. Biol. Chem. 1995, 270, 14651–14658. [Google Scholar] [CrossRef] [Green Version]
- Charpentier, T.H.; Thompson, L.E.; Liriano, M.A.; Varney, K.M.; Wilder, P.T.; Pozharski, E.; Toth, E.A.; Weber, D.J. The Effects of CapZ Peptide (TRTK-12) Binding to S100B–Ca2+ as Examined by NMR and X-ray Crystallography. J. Mol. Biol. 2010, 396, 1227–1243. [Google Scholar] [CrossRef] [Green Version]
- Jensen, J.; Indurthi, V.S.; Neau, D.B.; Vetter, S.W.; Colbert, C.L. Structural insights into the binding of the human receptor for advanced glycation end products (RAGE) by S100B, as revealed by an S100B–RAGE-derived peptide complex. Acta Crystallogr. Sect. D Biol. Crystallogr. 2015, 71, 1176–1183. [Google Scholar] [CrossRef] [Green Version]
- Gogl, G.; Alexa, A.; Kiss, B.; Katona, G.; Kovács, M.; Bodor, A.; Reményi, A.; Nyitray, L. Structural Basis of Ribosomal S6 Kinase 1 (RSK1) Inhibition by S100B Protein. J. Biol. Chem. 2016, 291, 11–27. [Google Scholar] [CrossRef] [Green Version]
- Inman, K.G.; Yang, R.; Rustandi, R.R.; Miller, K.E.; Baldisseri, D.M.; Weber, D.J. Solution NMR structure of S100B bound to the high-affinity target peptide TRTK-12. J. Mol. Biol. 2002, 324, 1003–1014. [Google Scholar] [CrossRef]
- Kannan, S.; Aronica, P.G.; Tan, Y.S.; Verma, C.S. ACS Omega, Inhibiting S100B(ββ) for Activating Wild-Type p53: Design of Stapled Peptides. ACS Omega 2019, 4, 5335–5344. [Google Scholar] [CrossRef]
- 62. Case, D.A.; Ben-Shalom, I.Y.; Brozell, M.S.R.; Cerutti, D.S.; Cheatham, T.E., III; Cruzeiro, V.W.D.; Darden, T.A.; Duke, R.E.; Ghoreishi, D.; Gilson, M.K.; et al. AMBER 18; University of California: San Francisco, CA, USA, 2018. [Google Scholar]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the Accuracy of Protein Side Chain Backbone Parameters from, ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Valiev, M.; Bylaska, E.J.; Govind, N.; Kowalski, K.; Straatsma, T.P.; Van Dam, H.J.; Wang, D.; Nieplocha, J.; Apra, E.; Windus, T.L.; et al. NWChem: A comprehensive and scalable open-source solution for large scale molecular simulations. Comput. Phys. Commun. 2010, 181, 1477–1489. [Google Scholar] [CrossRef] [Green Version]
- Tan, Y.S.; Reeks, J.; Brown, C.J.; Thean, D.; Ferrer Gago, F.J.; Yuen, T.Y.; Goh, E.T.; Lee, X.E.; Jennings, C.E.; Joseph, T.L.; et al. Benzene Probes in Molecular Dynamics Simulations Reveal Novel Binding Sites for Ligand Design. J. Phys. Chem. Lett. 2016, 7, 3452–3457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD—visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- De Lano, W. The PyMOL Molecular Graphics System; De Lano Scientific: San Carlos, CA, USA, 2002. [Google Scholar]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kannan, S.; Aronica, P.G.A.; Nguyen, T.B.; Li, J.; Verma, C.S. Computational Design of Macrocyclic Binders of S100B(ββ): Novel Peptide Theranostics. Molecules 2021, 26, 721. https://doi.org/10.3390/molecules26030721
Kannan S, Aronica PGA, Nguyen TB, Li J, Verma CS. Computational Design of Macrocyclic Binders of S100B(ββ): Novel Peptide Theranostics. Molecules. 2021; 26(3):721. https://doi.org/10.3390/molecules26030721
Chicago/Turabian StyleKannan, Srinivasaraghavan, Pietro G. A. Aronica, Thanh Binh Nguyen, Jianguo Li, and Chandra S. Verma. 2021. "Computational Design of Macrocyclic Binders of S100B(ββ): Novel Peptide Theranostics" Molecules 26, no. 3: 721. https://doi.org/10.3390/molecules26030721






