Structural Refinement of 2,4-Thiazolidinedione Derivatives as New Anticancer Agents Able to Modulate the BAG3 Protein
Abstract
:1. Introduction
2. Results and Discussion
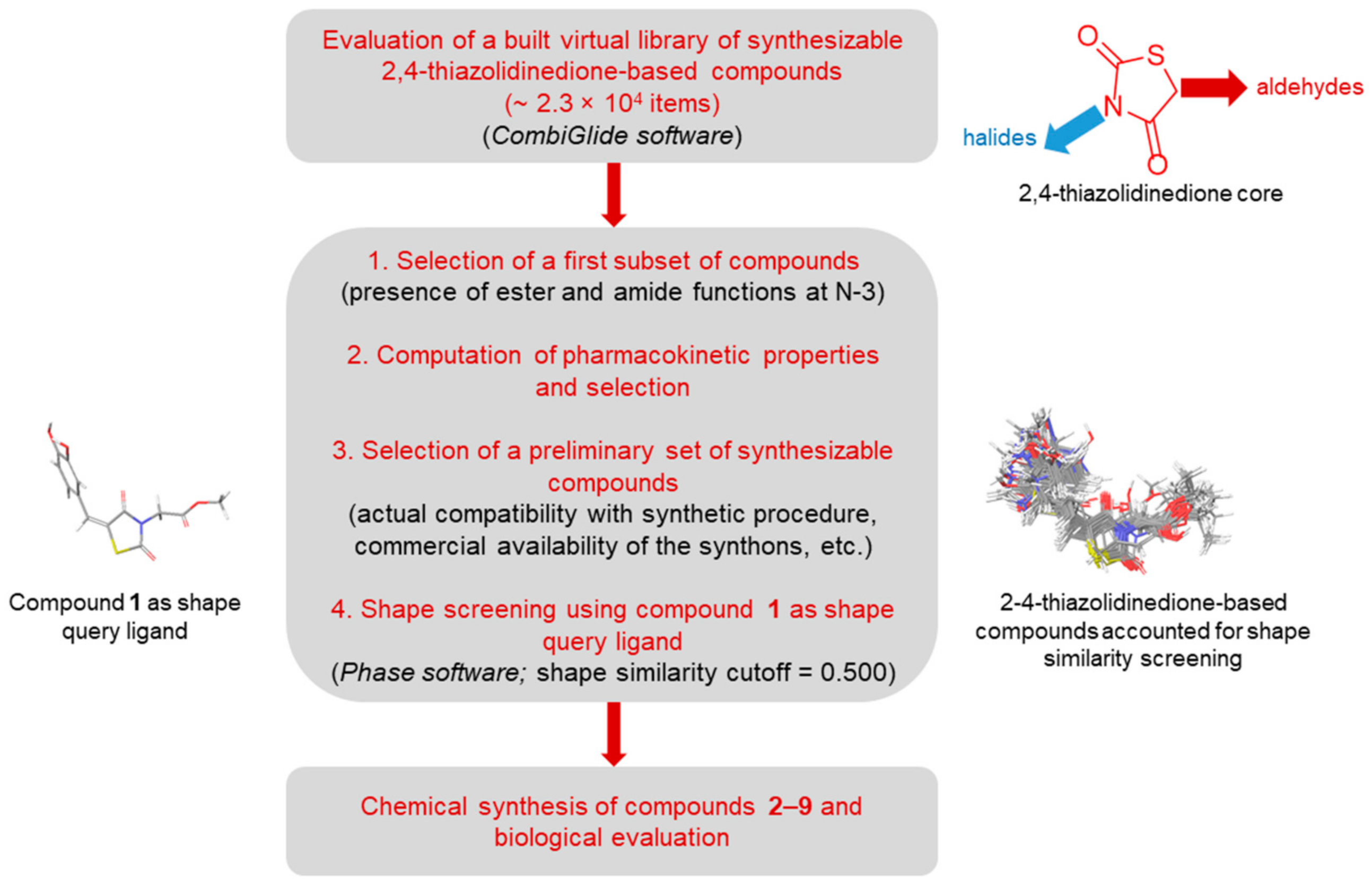
2.1. In-Silico Screening
2.2. Synthesis of 2,4-Thiazolidinedione Derivatives 2–9
2.3. Biophysical Assays
2.4. Biological Assays
3. Materials and Methods
3.1. In-Silico Screening
3.2. Synthesis of 2,4-Thiazolidinedione Molecules
- General procedure (A) for the synthesis of 10a–10d
- Thiazolidine-2,4-dione 10 (1.45 mmol; 1.0 equiv), the appropriate aromatic aldehydes a–d (1.45 mmol; 1.0 equiv), piperidine (1.17 mmol; 0.8 equiv), and ethanol (5 mL) were placed in a 25-mL round-bottom flask. The reaction mixture was continuously stirred and refluxed for 16–24 h (Scheme 1) [22]. The crude product was poured into water and extracted with AcOEt (3 × 25 mL). The combined organic phases were washed with water (3 × 25.0 mL), dried over anhydrous Na2SO4, filtered, and concentrated in vacuum. The desired compounds 10a–10d were confirmed by analytical RP-HPLC (Reversed-Phase-High Performance Liquid Chromatography) (Nucleodur© C8 reversed-phase column: 100 × 2 mm, 4 μM, 80 Å, flow rate = 1 mL/min) (Phenomenex, Castel Maggiore, BO, Italy) and 1H NMR spectra. 10a–10d were used without any further purification in the next step.
- (Z)-5-(3,4-dihydroxybenzylidene)thiazolidine-2,4-dione (10a)
- Compound 10a was obtained by following the general procedure (A), from the reaction between 10 and 3,4-dihydroxybenzaldehyde (a), as a brown solid (310 mg, 85% yield by HPLC analysis). RP-HPLC tR = 21.5, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.55 (s, 1H), 6.91 (d, J = 2.2 Hz, 1H), 6.86 (dd, J = 8.3, 2.2 Hz, 1H), 6.77 (d, J = 8.3 Hz, 1H). ESI-MS: calculated for C10H7NO4S 237.01 found m/z = 238.00 [M + H]+.
- (Z)-5-(3,5-dihydroxybenzylidene)thiazolidine-2,4-dione (10b)
- Compound 10b was obtained by following the general procedure (A), from the reaction between 10 and 3,5-dihydroxybenzaldehyde (b), as a brown solid (340 mg, 80% yield by HPLC analysis). RP-HPLC tR = 21.5, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.51 (s, 1H), 6.39 (d, J = 2.1 Hz, 2H), 6.25 (t, J = 2.1 Hz, 1H). ESI-MS: calculated for C10H7NO4S 237.01 found m/z = 238.02 [M + H]+.
- (Z)-5-(4-(bis(2-hydroxyethyl)amino)benzylidene)thiazolidine-2,4-dione (10c)
- Compound 10c was prepared in the same manner described for the preparation of 10a following the general procedure (A). From the reaction between 10 and 4-[N,N-Bis(2-hydroxyethyl)amino]benzaldehyde (c), 10c was obtained as a brown solid (420 mg, 80% yield by HPLC analysis). RP-HPLC tR = 20.4, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.53 (s, 1H), 7.30 (d, J = 8.9 Hz, 2H), 6.76 (dd, J = 7.4, 5.6 Hz, 2H), 3.66 (t, J = 5.7 Hz, 4H), 3.53 (t, J = 6.0 Hz, 4H). ESI-MS (Electrospray Ionization-Mass Spectrometry): calculated for C14H16N2O4S 308.08 found m/z = 309.13 [M + H]+.
- (Z)-4-(6-((2,4-dioxothiazolidin-5-ylidene)methyl)pyridin-3-yl)benzonitrile (10d)
- Compound 10d was obtained by following the general procedure (A), from the reaction between 10 and 4-(6-formylpyridin-2-yl)benzonitrile (d), as a brown solid (430 mg, 85% yield by HPLC analysis). RP-HPLC tR = 32.5, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 8.43 (d, J = 8.1 Hz, 2H), 7.98–7.87 (m, 4H), 7.66 (s, 1H). 7.62 (d, J = 7.2 Hz, 1H). ESI-MS: calculated for C16H9N3O2S 307.04 found m/z = 308.11 [M + H]+.
- General procedure (B) for the synthesis of 11,2–4, and6–9
- In a two-neck round-bottom flask, sodium hydride NaH (1.2 equiv) was added portion-wise to a solution of 10a–10d (0.07 g; 1.0 equiv) in dry DMF (3.5 mL). The reaction mixture was stirred for 1.5 h at 80 °C and it was allowed to return to room temperature. Then, ethyl bromoacetate (e) (1.5 equiv) for the compounds 3, 6, 8, methyl bromoacetate (f) (1.5 equiv) for 2, 4, 7, 9, and bromoacetic acid (1.5 equiv) (g) for the synthesis of 11 was added and the mixture was stirred at 80°C for 16–20 h. The reaction was monitored by TLC (Thin-layer chromatography) (Synthetic Scheme 1) [22,23]. The suspension was poured into ice-cold water and the solid product was filtered and recovered. HPLC purification was performed by semi-preparative reversed-phase HPLC (Nucleodur© C8 reversed-phase column: 250 × 10.00 mm, 4 μM, 80 Å, flow rate = 4 mL/min) and the final products were characterized by ESI-MS and NMR spectra.
- (Z)-methyl 2-(5-(3,4-dihydroxybenzylidene)-2,4-dioxothiazolidin-3-yl) acetate (2)
- Compound 2 was obtained by following the general procedure (B), from the reaction between 10a and methyl bromoacetate (f), as a brown solid (75 mg, 50% yield after HPLC purification). RP-HPLC tR = 28.9 min, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.82 (s, 1H), 7.07 (d, J = 2.2 Hz, 1H), 7.02 (dd, J = 8.4, 2.2, 1H), 6.91 (d, J = 8.2 Hz, 1H), 4.52 (s, 2H), 3,79 (s, 3H). 13C NMR (100 MHz, CD3OD): δ = 168.79, 168.71, 166.95, 150.05, 146.97, 136.02, 126.10, 125.51, 117.55, 117.33, 116.74, 53.00, 42.52. ESI-MS: calculated for C13H11NO6S, 309.03; found m/z = 332.02 [M + Na]+.
- (Z)-ethyl 2-(5-(3,5-dihydroxybenzylidene)-2,4-dioxothiazolidin-3-yl) acetate (3)
- Compound 3 was obtained by following the general procedure (B), from the reaction between 10b and ethyl bromoacetate (e), as a brown solid (80 mg, 60% yield after HPLC purification). RP-HPLC tR = 28.1 min, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, (CD3)2SO): δH = 9.72 (s, 2H), 7.80 (s, 1H), 6.52 (d, J = 6.9 Hz, 2H), 6.38 (d, J = 7.2 Hz, 1H), 4.49 (s, 2H), 4.18 (q, J = 7.3 Hz, 2H), 1.23 (t, J = 7.3 Hz, 3H). 13C NMR (100 MHz, CD3OD): δ = 167.38, 166.89, 165.54, 159.09 (2C), 134.67, 134.44, 120.59, 108.13 (2C), 105.00, 61.69, 41.60, 13.04. ESI-MS: calculated for C14H13NO6S, 323.05; found m/z = 346.03 [M + Na]+.
- (Z)-methyl 2-(5-(3,5-dihydroxybenzylidene)-2,4-dioxothiazolidin-3-yl) acetate (4)
- Compound 4 was obtained by following the general procedure (B), from the reaction between 10b and methyl bromoacetate (f), as a brown solid (75 mg, 75% yield after HPLC purification). RP-HPLC tR = 29.5 min, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.78 (s, 1H), 6.54 (d, J = 2.1 Hz, 2H), 6.39 (t, J = 2.1 Hz, 1H), 4.53 (s, 2H), 3.80 (s, 3H). 13C NMR (100 MHz, CD3OD): δ = 167.42, 167.38, 165.50, 159.07 (2C), 134.66, 134.45, 120.52, 108.24 (2C), 104.95, 51.85, 41.37. ESI-MS: calculated for C13H11NO6S, 309.03; found m/z = 332.02 [M + Na]+.
- (Z)-ethyl 2-(5-(4-(bis(2-hydroxyethyl) amino) benzylidene)-2,4-dioxothiazolidin-3-yl) acetate (6)
- Compound 6 was obtained by following the general procedure (B), from the reaction between 10c and ethyl bromoacetate (e), as a yellow solid (60 mg, 45% yield after HPLC purification). RP-HPLC tR = 26.2 min, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.84 (s, 1H), 7.49–7.45 (m, 2H), 6.93–6.90 (m, 2H), 4.49 (s, 2H), 4.28–4.22 (m, 2H), 3.79 (t, J = 6.0, 4H), 3.67 (t, J = 6.0 Hz, 4H), 1.33–1.28 (m, 3H). 13C NMR (100 MHz, CD3OD): δ = 167.8, 167.1, 166.0, 150.4, 134.9, 132.5 (2C), 120.3, 113.0, 111.8 (2C), 61.6, 58.8 (2C), 53.3 (2C), 41.4, 12.88. ESI-MS: calculated for C18H22N2O6S, 394.12; found m/z = 395.23 [M + H]+.
- (Z)-methyl 2-(5-(4-(bis(2-hydroxyethyl) amino) benzylidene)-2,4-dioxothiazolidin-3-yl) acetate (7)
- Compound 7 was obtained by following the general procedure (B), from the reaction between 10c and methyl bromoacetate (f), as a yellow solid (70 mg, 40% yield after HPLC purification). RP-HPLC tR = 24.2 min, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.84 (s, 1H), 7.47 (d, J = 8.6 Hz, 2H), 6.91 (d, J = 8.6 Hz, 2H), 4.51 (s, 2H), 3.81–3.77 (m, 7H), 3.67 (t, J = 5.7 Hz, 4H), 1.31 (s, 2H). 13C NMR (100 MHz, CD3OD): δ = 169.0, 168.8, 167.1, 151.6, 136.2, 133.7 (2C), 121.6, 114.2, 113.1 (2C), 60.0 (2C), 54.5 (2C), 53.0, 42.5. ESI-MS: calculated for C17H20N2O6S, 380.10; found m/z = 381.20 [M + H]+.
- (Z)-ethyl 2-(5-((5-(4-cyanophenyl) pyridin-2-yl) methylene)-2,4-dioxothiazolidin-3-yl) acetate (8)
- Compound 8 was obtained by following the general procedure (B), from the reaction between 10d and ethyl bromoacetate (e), as a yellow solid (100 mg, 50% yield after HPLC purification). RP-HPLC tR = 42.3 min, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CDCl3): δH = 8.24 (d, J = 8.3 Hz, 2H), 7.97–7.92 (m, 2H), 7.87 (d, J = 8.2 Hz, 2H), 7.80 (d, J = 8.0 Hz, 1H), 7.58 (d, J = 7.6 Hz, 1H), 4.52 (s, 2H), 4.27 (q, J = 7.1 Hz, 2H), 1.32 (t, J = 7.1 Hz, 3H). 13C NMR (100 MHz, CDCl3): δ = 170.8, 166.3, 165.8, 155.9, 151.8, 142.3, 138.3, 132.8 (2C), 129.3, 127.8 (2C), 126.9, 126.6, 121.1, 118.6, 113.3, 62.2, 41.7, 14.06. ESI-MS: calculated for C20H15N3O4S, 393.08; found m/z = 416.03 [M + Na]+.
- (Z)-methyl 2-(5-((5-(4-cyanophenyl) pyridin-2-yl) methylene)-2,4-dioxothiazolidin-3-yl) acetate (9)
- Compound 9 was obtained by following the general procedure (B), from the reaction between 10d and methyl bromoacetate (f), as a dark brown solid (75 mg, 45% yield after HPLC purification). RP-HPLC tR = 43.2 min, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CDCl3): δH = 8.17–8.14 (m, 2H), 7.89–7.84 (m, 2H), 7.80–7.76 (m, 2H), 7.73 (dd, J = 8.0, 0.9 Hz, 1H), 7.50 (dd, J = 7.7, 0.9 Hz, 1H), 4.45 (s, 2H), 3.73 (s, 3H). 13C NMR (100 MHz, (CD3)2SO): δ = 170.5, 167.7, 165.5, 155.2, 151.5, 142.2, 139.5, 133.3 (2C), 130.4, 128.5, 128.1 (2C), 125.3, 122.6, 119.1, 112.6, 53.1, 42.0. ESI-MS: calculated for C19H13N3O4S, 379.06; found m/z = 380.21 [M + H]+.
- (Z)-2-(5-(3,5-dihydroxybenzylidene)-2,4-dioxothiazolidin-3-yl) acetic acid (11)
- Compound 11 was obtained by following the general procedure (B), from the reaction between 10a and bromoacetic acid (g), as a brown solid (90 mg, 70% yield after HPLC purification). RP-HPLC tR = 16.3 gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.79 (s, 1H), 6.62 (d, J = 2.0 Hz, 2H), 6.26 (t, J = 1.9 Hz, 1H), 4.63 (s, 2H). ESI-MS: calculated for C12H9NO6S, 295.02; found m/z = 296.00 [M + Na]+. The desired compound was directly used for the following step.
- (Z)-5-(3,5-dihydroxybenzylidene)-3-(2-morpholino-2-oxoethyl) thiazolidine-2,4-dione (5)
- Compounds 1-Hydroxybenzotriazole (0.48 mmol; 2.0 equiv) and N,N′-diisopropylcarbodiimide (0.36 mmol; 1.5 equiv) were added to a solution of 11 (0.24 mmol; 1.0 equiv) in DMF (3 mL). The mixture was stirred for 1h at room temperature and then morpholine (0.48 mmol; 2.0 equiv) was added. The reaction mixture was stirred overnight at room temperature to obtain 5 (Scheme 1) [30]. After completion, the suspension was diluted with 10 mL of water and extracted with AcOEt (3 × 20 mL). The combined organic layers were dried over Na2SO4 and evaporated under vacuum. A portion of the crude product was purified by semi-preparative reversed-phase HPLC (Nucleodur© C8 reversed-phase column: 250 × 10.00 mm, 4 μM, 80 Å, flow rate = 4 mL/min). The desired compound 5 was characterized by ESI-MS and NMR spectra.
- Compound 5 was obtained as a pale brown solid (85 mg, 55% yield after HPLC purification). RP-HPLC tR = 17.5 min, gradient condition: from 5% B ending to 100% B 50 min, flow rate of 4 mL/min, λ = 240 nm. 1H NMR (400 MHz, CD3OD): δH = 7.71 (s, 1H), 6.57 (s, 2H), 6.40–6.37 (m, 2H), 6.26 (dt, J = 6.1, 2.2 Hz, 1H), 5.34 (s, 2H), 3.61–3.54 (m, 4H), 3.52–3.45 (m, 4H). 13C NMR (100 MHz, CD3OD): δ = 171.9, 170.7, 169.3, 159.0 (2C), 132.9, 132.2, 120.5, 108.2 (2C), 104.9, 75.9 (2C), 66.2 (2C), 29.4. ESI-MS: calculated for C16H16N2O6S, 364.07; found m/z = 362.90 [M − H]+.
3.3. Biophysical Assay
3.4. Biological Assays
3.4.1. Cell Culture
3.4.2. Cell Viability Assay
3.4.3. Apoptosis and Cell Cycle Analysis
3.4.4. Measurement of Caspase 3 and Caspase 9 Levels
3.4.5. Data Analysis
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
References
- Kögel, D.; Linder, B.; Brunschweiger, A.; Chines, S.; Behl, C. At the crossroads of apoptosis and autophagy: Multiple roles of the co-chaperone BAG3 in stress and therapy resistance of cancer. Cells 2020, 9, 574. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kabbage, M.; Dickman, M.B. The BAG proteins: A ubiquitous family of chaperone regulators. Cell. Mol. Life Sci. 2008, 65, 1390–1402. [Google Scholar] [CrossRef] [PubMed]
- Behl, C. Breaking BAG: The co-chaperone BAG3 in health and disease. Trends Pharmacol. Sci. 2016, 37, 672–688. [Google Scholar] [CrossRef] [PubMed]
- Sondermann, H.; Scheufler, C.; Schneider, C.; Höhfeld, J.; Hartl, F.U.; Moarefi, I. Structure of a Bag/Hsc70 complex: Convergent functional evolution of Hsp70 nucleotide exchange factors. Science 2001, 291, 1553–1557. [Google Scholar] [CrossRef]
- Doong, H.; Vrailas, A.; Kohn, E.C. What’s in the ‘BAG’?—A functional domain analysis of the BAG-family proteins. Cancer Lett. 2002, 188, 25–32. [Google Scholar] [CrossRef] [Green Version]
- Gamerdinger, M.; Hajieva, P.; Kaya, A.M.; Wolfrum, U.; Hartl, F.U.; Behl, C. Protein quality control during aging involves recruitment of the macroautophagy pathway by BAG3. EMBO J. 2009, 28, 889–901. [Google Scholar] [CrossRef] [Green Version]
- Rosati, A.; Graziano, V.; De Laurenzi, V.; Pascale, M.; Turco, M.C. BAG3: A multifaceted protein that regulates major cell pathways. Cell Death Discov. 2011, 2, e141. [Google Scholar] [CrossRef]
- Chiappetta, G.; Ammirante, M.; Basile, A.; Rosati, A.; Festa, M.; Monaco, M.; Vuttariello, E.; Pasquinelli, R.; Arra, C.; Zerilli, M.; et al. The antiapoptotic protein BAG3 is expressed in thyroid carcinomas and modulates apoptosis mediated by tumor necrosis factor-related apoptosis-inducing ligand. J. Clin. Endocrinol. Metab. 2007, 92, 1159–1163. [Google Scholar] [CrossRef]
- Staibano, S.; Mascolo, M.; Di Benedetto, M.; Vecchione, M.L.; Ilardi, G.; Di Lorenzo, G.; Autorino, R.; Salerno, V.; Morena, A.; Rocco, A.; et al. BAG3 protein delocalisation in prostate carcinoma. Tumor Biol. 2010, 31, 461–469. [Google Scholar] [CrossRef] [Green Version]
- Franco, R.; Scognamiglio, G.; Salerno, V.; Sebastiani, A.; Cennamo, G.; Ascierto, P.A.; Botti, G.; Turco, M.C.; Rosati, A. Expression of the anti-apoptotic protein BAG3 in human melanomas. J. Investig. Dermatol. 2012, 132, 252–254. [Google Scholar] [CrossRef] [Green Version]
- Yang, X.; Tian, Z.; Gou, W.; Takahashi, H.; Xing, Y.; Takano, Y.; Zheng, H.-C. Bag-3 expression is involved in pathogenesis and progression of colorectal carcinomas. Histol. Histopathol. 2013, 28, 1147–1156. [Google Scholar] [CrossRef]
- Nymoen, D.A.; Falkenthal, T.E.H.; Holth, A.; Ow, G.S.; Ivshina, A.V.; Tropé, C.G.; Kuznetsov, V.A.; Staff, A.C.; Davidson, B. Expression and clinical role of chemoresponse-associated genes in ovarian serous carcinoma. Gynecol. Oncol. 2015, 139, 30–39. [Google Scholar] [CrossRef]
- Chiappetta, G.; Basile, A.; Barbieri, A.; Falco, A.; Rosati, A.; Festa, M.; Pasquinelli, R.; Califano, D.; Palma, G.; Costanzo, R.; et al. The anti-apoptotic BAG3 protein is expressed in lung carcinomas and regulates small cell lung carcinoma (SCLC) tumor growth. Oncotarget 2014, 5, 6846–6853. [Google Scholar] [CrossRef] [Green Version]
- Nourashrafeddin, S.; Aarabi, M.; Modarressi, M.H.; Rahmati, M.; Nouri, M. The evaluation of WBP2NL-related genes expression in breast cancer. Pathol. Oncol. Res. 2015, 21, 293–300. [Google Scholar] [CrossRef]
- Esposito, V.; Baldi, C.; Zeppa, P.; Festa, M.; Guerriero, L.; D’Avenia, M.; Chetta, M.; Zullo, F.; De Laurenzi, V.; Turco, M.C.; et al. BAG3 protein is over-expressed in endometrioid endometrial adenocarcinomas. J. Cell. Physiol. 2017, 232, 309–311. [Google Scholar] [CrossRef]
- Rosati, A.; Basile, A.; Falco, A.; D’Avenia, M.; Festa, M.; Graziano, V.; De Laurenzi, V.; Arra, C.; Pascale, M.; Turco, M.C. Role of BAG3 protein in leukemia cell survival and response to therapy. Biochim. Biophys. Acta Rev. Cancer 2012, 1826, 365–369. [Google Scholar] [CrossRef]
- Rosati, A.; Basile, A.; D’Auria, R.; D’Avenia, M.; De Marco, M.; Falco, A.; Festa, M.; Guerriero, L.; Iorio, V.; Parente, R.; et al. BAG3 promotes pancreatic ductal adenocarcinoma growth by activating stromal macrophages. Nat. Commun. 2015, 6, 8965. [Google Scholar] [CrossRef]
- Terracciano, S.; Lauro, G.; Russo, A.; Vaccaro, M.C.; Vassallo, A.; De Marco, M.; Ranieri, B.; Rosati, A.; Turco, M.C.; Riccio, R. Discovery and synthesis of the first selective BAG domain modulator of BAG3 as an attractive candidate for the development of a new class of chemotherapeutics. Chem. Commun. 2018, 54, 7613–7616. [Google Scholar] [CrossRef]
- Schrödinger Release 2020-1. In Phase; Schrödinger, LLC: New York, NY, USA, 2020.
- Sastry, G.M.; Inakollu, V.S.; Sherman, W. Boosting virtual screening enrichments with data fusion: Coalescing hits from two-dimensional fingerprints, shape, and docking. J. Chem. Inf. Model. 2013, 53, 1531–1542. [Google Scholar] [CrossRef]
- Pala, D.; Beuming, T.; Sherman, W.; Lodola, A.; Rivara, S.; Mor, M. Structure-based virtual screening of MT2 melatonin receptor: Influence of template choice and structural refinement. J. Chem. Inf. Model. 2013, 53, 821–835. [Google Scholar] [CrossRef]
- Bruno, G.; Costantino, L.; Curinga, C.; Maccari, R.; Monforte, F.; Nicolo, F.; Ottana, R.; Vigorita, M.G. Synthesis and aldose reductase inhibitory activity of 5-arylidene-2, 4-thiazolidinediones. Bioorg. Med. Chem. 2002, 10, 1077–1084. [Google Scholar] [CrossRef]
- Leonova, E.S.; Makarov, M.V.; Rybalkina, E.Y.; Nayani, S.L.; Tongwa, P.; Fonari, A.; Timofeeva, T.V.; Odinets, I.L. Structure–cytotoxicity relationship in a series of N-phosphorus substituted E, E-3, 5-bis (3-pyridinylmethylene)-and E, E-3, 5-bis (4-pyridinylmethylene) piperid-4-ones. Eur. J. Med. Chem. 2010, 45, 5926–5934. [Google Scholar] [CrossRef]
- Williams, A.; Ibrahim, I.T. Carbodiimide chemistry: Recent advances. Chem. Rev. 1981, 81, 589–636. [Google Scholar] [CrossRef]
- Carpino, L.A.; El-Faham, A. The diisopropylcarbodiimide/1-hydroxy-7-azabenzotriazole system: Segment coupling and stepwise peptide assembly. Tetrahedron 1999, 55, 6813–6830. [Google Scholar] [CrossRef]
- De Marco, M.; Basile, A.; Iorio, V.; Festa, M.; Falco, A.; Ranieri, B.; Pascale, M.; Sala, G.; Remondelli, P.; Capunzo, M.; et al. Role of BAG3 in cancer progression: A therapeutic opportunity. Semin. Cell Dev. Biol. 2018, 78, 85–92. [Google Scholar] [CrossRef]
- Porter, A.G.; Jänicke, R.U. Emerging roles of caspase-3 in apoptosis. Cell Death Differ. 1999, 6, 99–104. [Google Scholar] [CrossRef]
- Kuida, K. Caspase-9. Int. J. Biochem. Cell Biol. 2000, 32, 121–124. [Google Scholar] [CrossRef]
- Gentilella, A.; Khalili, K. Autoregulation of co-chaperone BAG3 gene transcription. J. Cell. Biochem. 2009, 108, 1117–1124. [Google Scholar] [CrossRef] [Green Version]
- Vaisburg, A.; Bernstein, N.; Frechette, S.; Allan, M.; Abou-Khalil, E.; Leit, S.; Moradei, O.; Bouchain, G.; Wang, J.; Woo, S.H. (2-Amino-phenyl)-amides of ω-substituted alkanoic acids as new histone deacetylase inhibitors. Bioorg. Med. Chem. Lett. 2004, 14, 283–287. [Google Scholar] [CrossRef]
- Persico, M.; Ramunno, A.; Maglio, V.; Franceschelli, S.; Esposito, C.; Carotenuto, A.; Brancaccio, D.; De Pasquale, V.; Pavone, L.M.; Varra, M.; et al. New anticancer agents mimicking protein recognition motifs. J. Med. Chem. 2013, 56, 6666–6680. [Google Scholar] [CrossRef]
- Pecoraro, M.; Pinto, A.; Popolo, A. Inhibition of Connexin 43 translocation on mitochondria accelerates CoCl2-induced apoptotic response in a chemical model of hypoxia. Toxicol. In Vitro 2018, 47, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Franceschelli, S.; Bruno, A.P.; Festa, M.; Falco, A.; Gionti, E.; D’Avenia, M.; De Marco, M.; Basile, A.; Iorio, V.; Marzullo, L.; et al. BAG3 protein is involved in endothelial cell response to phenethyl isothiocyanate. Oxid. Med. Cell Longev. 2018, 2018, 5967890. [Google Scholar] [CrossRef] [PubMed] [Green Version]






| Compound | rBAG3 KD ± SD (nM) | rBAG3BD (KD ± SD (nM) | rBAG4 KD ± SD (nM) |
|---|---|---|---|
| 1 (lead) | 11.1 ± 3.9 | 6.4 ± 2.2 | n.b.a |
| 2 | 115.2 ± 1.4 | n.b. | n.b. |
| 3 | n.b. | n.d.b | n.d. |
| 4 | n.b. | n.d. | n.d. |
| 5 | 12.4 ± 1.2 | n.b. | n.b. |
| 6 | 6.3 ± 0.3 | 27.6 ± 1.9 | n.b. |
| 7 | 53.1 ± 9.1 | 45.7 ± 1.7 | n.b. |
| 8 | 143.5 ± 1.5 | n.b. | n.b. |
| 9 | 24.7 ± 1.8 | 22.4 ± 0.7 | n.b. |
| Compound | A375 Cell Line IC50 ± SD (µM) | HeLa Cell Line IC50 ± SD (µM) |
|---|---|---|
| 1 (lead) | 15.08 ± 0.9 | >50 |
| 2 | >50 | 27.30 ± 1.2 |
| 5 | 25.46 ± 1.1 | 30.02 ± 1.3 |
| 6 | 19.36 ± 1.2 | 18.67 ± 0.9 |
| 7 | >50 | 49.37 ± 1.4 |
| 8 | 23.15 ± 1.0 | 25.90 ± 0.8 |
| 9 | 28.40 ± 0.9 | 31.28 ± 1.0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruggiero, D.; Terracciano, S.; Lauro, G.; Pecoraro, M.; Franceschelli, S.; Bifulco, G.; Bruno, I. Structural Refinement of 2,4-Thiazolidinedione Derivatives as New Anticancer Agents Able to Modulate the BAG3 Protein. Molecules 2022, 27, 665. https://doi.org/10.3390/molecules27030665
Ruggiero D, Terracciano S, Lauro G, Pecoraro M, Franceschelli S, Bifulco G, Bruno I. Structural Refinement of 2,4-Thiazolidinedione Derivatives as New Anticancer Agents Able to Modulate the BAG3 Protein. Molecules. 2022; 27(3):665. https://doi.org/10.3390/molecules27030665
Chicago/Turabian StyleRuggiero, Dafne, Stefania Terracciano, Gianluigi Lauro, Michela Pecoraro, Silvia Franceschelli, Giuseppe Bifulco, and Ines Bruno. 2022. "Structural Refinement of 2,4-Thiazolidinedione Derivatives as New Anticancer Agents Able to Modulate the BAG3 Protein" Molecules 27, no. 3: 665. https://doi.org/10.3390/molecules27030665










