Mintaimycins, a Group of Novel Peptide Metabolites from Micromonospora sp. C-3509
Abstract
:1. Introduction
2. Results and Discussion
2.1. Discovery of New Secondary Metaboltes (Mintaimycins) from Micromonospora sp. C-3509
2.2. Structural Elucidation of Mintaimycins (1–4)
2.3. Stereochemistry of Mintaimycins
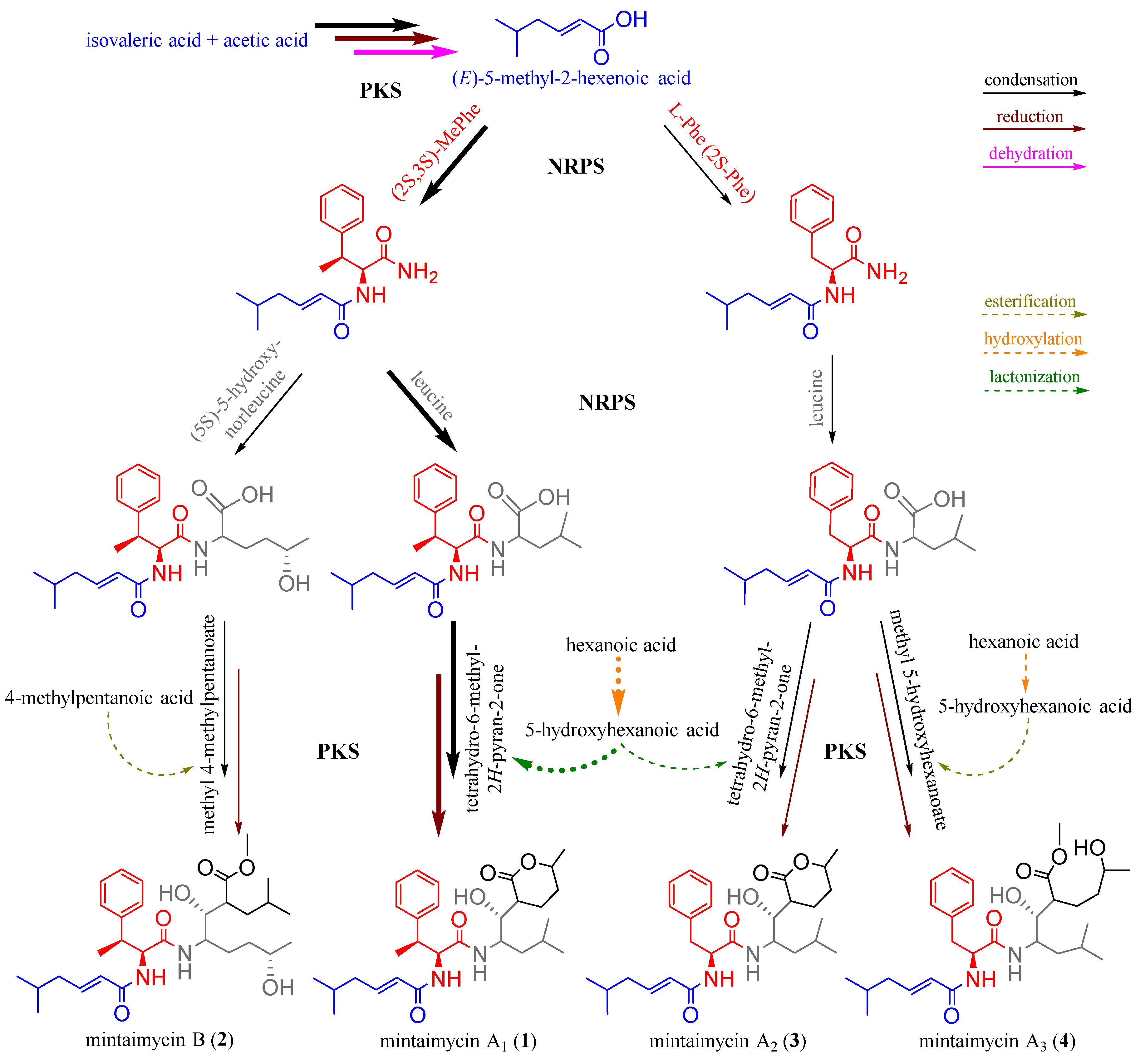
2.4. Speculation of Pathway for Mintaimycins (1–4) Biosynthesis
2.5. Biological Activities of Mintaimycins
3. Materials and Methods
3.1. General Procedures
3.2. Solid State Fermentation of Micromonospora sp. C-3509
3.3. Extraction and Isolation of Mintaimycins (1–4)
3.4. Stereochemistry of β-Methylphenylalanine (β-MePhe) and Phenylalanine (Phe) Residue in Mintaimycins by Marfey’s Method
3.5. Preparation of (R)- and (S)-MTPA Ester Derivatives of Mintaimycins A1 (1), B (2) and A2 (3) by Mosher’s Method
3.6. Pre-Adipocyte Differentiation Assay
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Maiese, W.M.; Lechevalier, M.P.; Lechevalier, H.A.; Korshalla, J.; Kuck, N.; Fantini, A.; Wildey, M.J.; Thomas, J.; Greenstein, M. Calicheamicins, a novel family of antitumor antibiotics: Taxonomy, fermentation and biological properties. J. Antibiot. 1989, 42, 558–563. [Google Scholar] [CrossRef] [PubMed]
- Skellam, E.J.; Stewart, A.K.; Strangman, W.K.; Wright, J.L. Identification of micromonolactam, a new polyene macro cyclic lactam from two Marine Micromonospora strains using chemical and molecular methods: Clarification of the biosynthetic pathway from a glutamate starter unit. J. Antibiot. 2013, 66, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Nie, Y.L.; Wu, Y.D.; Wang, C.X.; Lin, R.; Xie, Y.; Fang, D.S.; Jiang, H.; Lian, Y.Y. Structure elucidation and antitumour activity of a new macrolactam produced by marine-derived actinomycete Micromonospora sp. FIM05328. Nat. Prod. Res. 2018, 32, 2133–2138. [Google Scholar] [CrossRef] [PubMed]
- Antal, N.; Fiedler, H.P.; Stackebrandt, E.; Beil, W.; Stroch, K.; Zeeck, A. Retymicin, galtamycin B, saquayamycin Z and ribofuranosyllumichrome, novel secondary metabolites from Micromonospora sp. Tü 6368. I. Taxonomy, fermentation, isolation and biological activities. J. Antibiot. 2005, 58, 95–102. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Furumai, T.; Igarashi, Y.; Higuchi, H.; Saito, N.; Oki, T. Kosinostatin, a quinocycline antibiotic with antitumor activity from Micromonospora sp. TP-A0468. J. Antibiot. 2002, 55, 128–133. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pang, X.; Zhao, J.; Fang, X.; Liu, H.; Zhang, Y.; Cen, S.; Yu, L. Surfactin derivatives from Micromonospora sp. CPCC 202787 and their anti-HIV activities. J. Antibiot. 2017, 70, 105–108. [Google Scholar] [CrossRef] [PubMed]
- McBrien, K.D.; Berry, R.L.; Lowe, S.E.; Neddermann, K.M.; Bursuker, I.; Huang, S.; Klohr, S.E.; Leet, J.E. Rakicidins, new cytotoxic lipopeptides from Micromonospora sp. fermentation, isolation and characterization. J. Antibiot. 1995, 48, 1446–1452. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qi, S.; Gui, M.; Li, H.; Yu, C.; Li, H.; Zeng, Z.; Sun, P. Secondary Metabolites from Marine Micromonospora: Chemistry and Bioactivities. Chem. Biodivers. 2020, 17, e2000024. [Google Scholar] [CrossRef]
- Hu, X.; Hu, X.; Hu, X.; Li, S.; Li, L.; Yu, L.; Liu, H.; You, X.; Wang, Z.; Li, L.; et al. Cytotoxic and Antibacterial Cervinomycins B1-4 from a Streptomyces Species. J. Nat. Prod. 2019, 82, 2337–2342. [Google Scholar] [CrossRef]
- Li, L.; Li, S.; Jiang, B.; Zhang, M.; Zhang, J.; Yang, B.; Li, L.; Yu, L.; Liu, H.; You, X.; et al. Isarubrolones Containing a Pyridooxazinium Unit from Streptomyces as Autophagy Activators. J. Nat. Prod. 2019, 82, 1149–1154. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Hu, X.; Sun, G.; Li, L.; Jiang, B.; Li, S.; Bai, L.; Liu, H.; Yu, L.; Wu, L. Genome-Guided Discovery of Pretilactam from Actinosynnema pretiosum ATCC 31565. Molecules 2019, 24, 2281. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, C.; Zhang, W.; Gao, R.; Jin, L.; Li, D.; Shao, R. Separation, purification and structure identification of the antitumor antibiotic component from fermentation broth of a new Micromonospora sp. C-3509. Chin. J. New Drugs 2010, 19, 564–568. [Google Scholar]
- Pirali, G.; Pagani, H.; Grandi, M.; Paternoster, M.; Tamoni, G.; Cassani, G. Antimicrobial and Antitumor Antibiotic M 9026 and Its Pure Individual Factors 1, 2 and 3, and Microbial Process for Production thereof. U.S. Patent 4692333, 8 September 1987. [Google Scholar]
- Igarashi, Y.; Yu, L.; Ikeda, M.; Oikawa, T.; Kitani, S.; Nihira, T.; Bayanmunkh, B.; Panbangred, W. Jomthonic acid, a modified amino acid from a soil-derived Streptomyces. J. Nat. Prod. 2012, 75, 986–990. [Google Scholar] [CrossRef] [PubMed]
- Jiao, W.H.; Khalil, Z.; Dewapriya, P.; Salim, A.A.; Lin, H.W.; Capon, R.J. Trichodermides A–E: New Peptaibols Isolated from the Australian Termite Nest-Derived Fungus Trichoderma virens CMB-TN16. J. Nat. Prod. 2018, 81, 976–984. [Google Scholar] [CrossRef] [PubMed]
- Annang, F.; Perez-Victoria, I.; Perez-Moreno, G.; Domingo, E.; Gonzalez, I.; Tormo, J.R.; Martin, J.; Ruiz-Perez, L.M.; Genilloud, O.; Gonzalez-Pacanowska, D.; et al. MDN-0185, an Antiplasmodial Polycyclic Xanthone Isolated from Micromonospora sp. CA-256353. J. Nat. Prod. 2018, 81, 1687–1691. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Salcedo, R.; Alvarez-Alvarez, R.; Olano, C.; Canedo, L.; Brana, A.F.; Mendez, C.; de la Calle, F.; Salas, J.A. Characterization of the Jomthonic Acids Biosynthesis Pathway and Isolation of Novel Analogues in Streptomyces caniferus GUA-06-05-006A. Mar. Drugs 2018, 16, 259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, L.; Oikawa, T.; Kitani, S.; Nihira, T.; Bayanmunkh, B.; Panbangred, W.; Igarashi, Y. Jomthonic acids B and C, two new modified amino acids from Streptomyces sp. J. Antibiot. 2014, 67, 345–347. [Google Scholar] [CrossRef] [PubMed]
- Mehlem, A.; Hagberg, C.E.; Muhl, L.; Eriksson, U.; Falkevall, A. Imaging of neutral lipids by oil red O for analyzing the metabolic status in health and disease. Nat. Protoc. 2013, 8, 1149–1154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
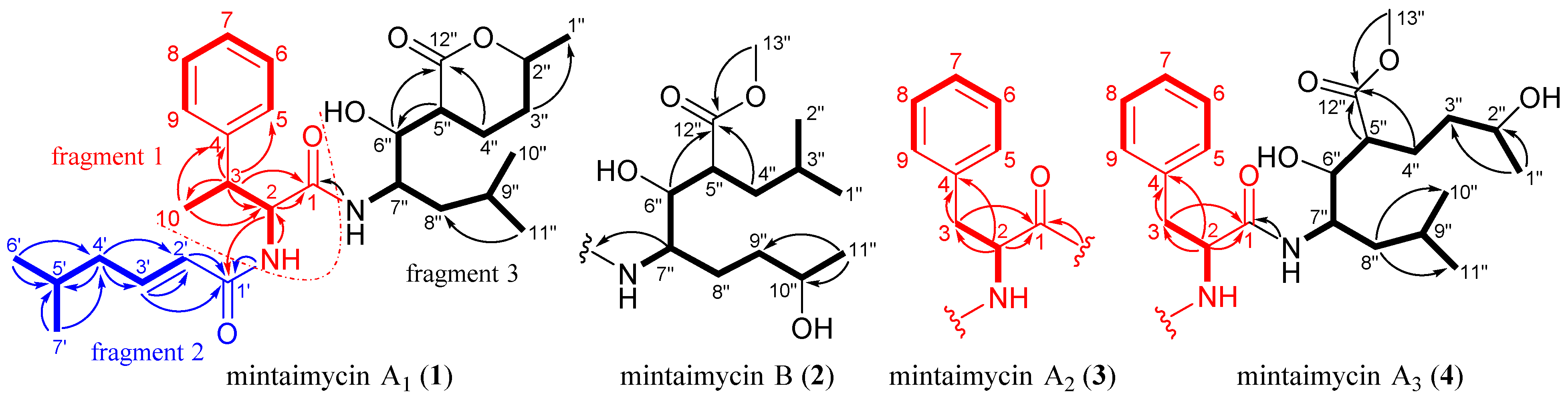

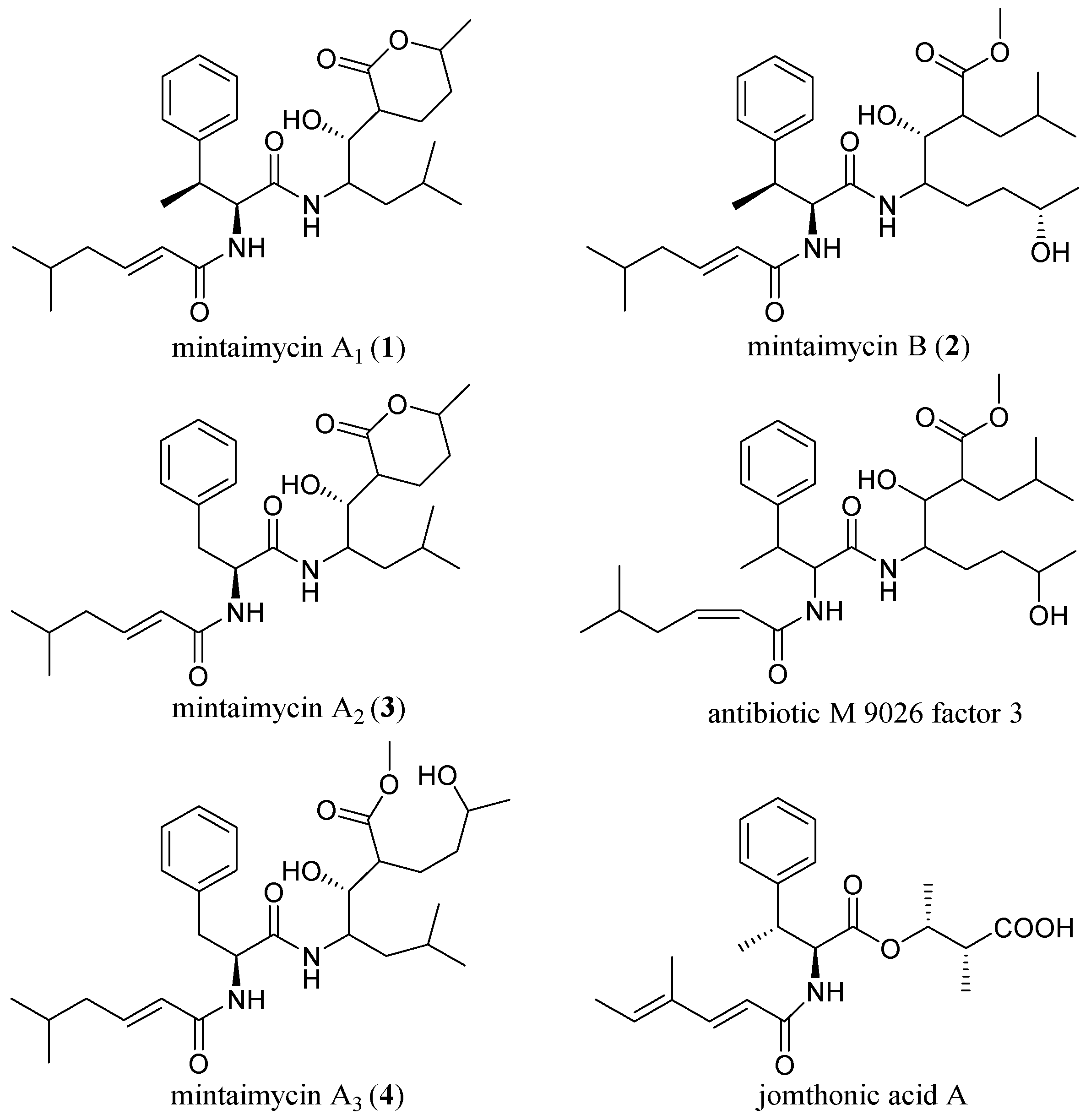

| Mintaimycin A1 (1) | Mintaimycin B (2) | Mintaimycin A2 (3) | Mintaimycin A3 (4) | |||||
|---|---|---|---|---|---|---|---|---|
| Position | δC, Type | δH (J in Hz) | δC, Type | δH (J in Hz) | δC, Type | δH (J in Hz) | δC, Type | δH (J in Hz) |
| 1 | 172.5, C | 170.7, C | 171.0, C | 172.5, C | ||||
| 1-NH | 7.22, d (10.2) | 6.15, d (8.0) | 7.88, d (9.6) | 7.12, d (8.8) | ||||
| 2 | 60.8, CH | 4.50, dd (10.2, 8.4) | 59.1, CH | 4.50, t (7.2) | 55.0, CH | 4.44, m | 56.6, CH | 4.67, m |
| 3 | 42.4, CH | 3.19, dq (10.2, 6.6) | 40.5, CH | 3.43, m | 37.5, CH2 | 2.88, dd (15.8, 6.6); 2.80, dd (15.8, 9.0) | 39.6, CH2 | 2.93, dd (11.9, 7.2); 3.10, dd (11.9, 7.2) |
| 4 | 145.2, C | 141.7, C | 137.9, C | 139.3, C | ||||
| 5 | 129.2, CH | 7.29, overlap | 127.6, CH | 7.24, overlap | 128.2, CH | 7.25, overlap | 130.8, CH | 7.26, overlap |
| 6 | 129.9, CH | 7.27, overlap | 128.9, CH | 7.31, t (7.2) | 129.3, CH | 7.25, overlap | 129.7, CH | 7.26, overlap |
| 7 | 128.0, CH | 7.22, m | 127.2, CH | 7.23, d (7.2) | 126.4, CH | 7.18, m | 127.9, CH | 7.19, overlap |
| 8 | 129.9, CH | 7.27, overlap | 128.9, CH | 7.31, t (7.2) | 129.3, CH | 7.25, overlap | 129.7, CH | 7.26, overlap |
| 9 | 129.2, CH | 7.29, overlap | 127.6, CH | 7.24, overlap | 128.2, CH | 7.25, overlap | 130.8, CH | 7.26, overlap |
| 10 | 20.7, CH3 | 1.29, d (7.2) | 18.2, CH3 | 1.30, d (7.2) | ||||
| 1′ | 166.4, C | 165.8, C | 164.9, C | 166.5, C | ||||
| 1′-NH | 7.09, d (8.4) | 5.77, d (7.2) | 8.21, d (8.4) | 7.34, d (8.0) | ||||
| 2′ | 126.5, CH | 5.76, dt (15.6, 1.2) | 123.7, CH | 5.62, d (15.2); | 125.4, CH | 5.93, d (15.0) | 126.7, CH | 5.98, d (14.4) |
| 3′ | 143.9, CH | 6.62, td (15.0, 7.8) | 145.1, CH | 6.70, td (15.2, 7.2) | 141.6, CH | 6.53, td (15.0, 7.2) | 144.0, CH | 6.71, dt (14.4, 6.4) |
| 4′ | 42.4, CH2 | 1.97, m | 41.3, CH2 | 2.00, m | 40.6, CH2 | 2.00, m | 42.5, CH2 | 2.05, overlap |
| 5′ | 29.3, CH | 1.66, m | 27.8, CH | 1.88, m | 27.6, CH | 1.67, m | 29.3, CH | 1.72, m |
| 6′ | 23.3, CH3 | 0.86, d (6.6) | 22.3, CH3 | 0.87, d (6.4) | 21.6, CH3 | 0.87, d (6.6) | 23.3, CH3 | 0.90, d (6.4) |
| 7′ | 23.3, CH3 | 0.86, d (6.6) | 22.4, CH3 | 0.87, d (6.4) | 22.3, CH3 | 0.87, d (6.6) | 23.3, CH3 | 0.90, d (6.4) |
| 1″ | 22.5, CH3 | 1.22, d (6.6) | 21.4, CH3 | 0.94, d (6.4) | 22.0, CH3 | 1.89, d (6.0) | 24.7, CH3 | 1.06, d (6.4) |
| 2″ | 78.4, CH | 4.28, m | 23.9, CH3 | 0.93, d (6.4) | 76.9, CH | 4.27, m | 67.3, CH | 3.64, overlap |
| 3″ | 31.7, CH2 | 1.90, m; 1.38, m | 24.8, CH | 1.64, m | 29.8, CH2 | 1.76, m; 1.26, m | 38.5, CH2 | 1.37, m; 1.28, m |
| 4″ | 20.3, CH2 | 1.98, m; 1.89, m | 19.5, CH2 | 1.99, m; 1.87, m | 18.5, CH2 | 1.83, m; 1.68, m | 24.8, CH2 | 1.86, m; 1.72, m |
| 5″ | 45.3, CH | 2.77, ddd (9.0, 7.2, 2.4) | 43.8, CH | 2.58, m | 43.5, CH | 2.65, m | 49.4, CH | 2.51, m |
| 6″ | 75.3, CH | 4.16, ddd (8.4, 6.0, 1.8) | 73.8, CH | 4.11, overlap | 73.0, CH | 3.94, ddd (9.0, 6.6, 1.8) | 76.2, CH | 3.69, m |
| 7″ | 50.3, CH | 4.03, m | 49.3, CH | 4.10, overlap | 48.2, CH | 3.64, m | 51.3, CH | 3.90, m |
| 8″ | 43.3, CH2 | 1.79, m; 1.38, m | 40.8, CH2 | 1.55, m; 1.26, m | 41.0, CH2 | 1.46, m; 1.13, m | 39.8, CH2 | 1.38, m |
| 9″ | 26.2, CH | 1.79, m | 30.3, CH2 | 1.89, m; 1.42, m | 24.1, CH | 1.14, m | 25.5, CH | 1.28, m |
| 10″ | 23.1, CH3 | 0.95, d (6.6) | 78.0, CH | 4.39, m | 22.4, CH3 | 0.74, d (6.6) | 22.3, CH3 | 0.78, d (6.4) |
| 11″ | 25.2, CH3 | 0.93, d (6.6) | 22.0, CH3 | 1.31, d (7.2) | 24.2, CH3 | 0.74, d (6.6) | 25.1, CH3 | 0.80, d (6.4) |
| 12″ | 173.8, C | 173.8, C | 172.9, C | 176.2, C | ||||
| 13″-OCH3 | 50.9, CH3 | 3.49, s | 52.2, CH3 | 3.63, s | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hu, X.; Wang, Y.; Zhao, C.; Li, S.; Hu, X.; You, X.; Shen, J.; Wang, Z.; Hong, B.; Jiang, B.; et al. Mintaimycins, a Group of Novel Peptide Metabolites from Micromonospora sp. C-3509. Molecules 2022, 27, 1150. https://doi.org/10.3390/molecules27041150
Hu X, Wang Y, Zhao C, Li S, Hu X, You X, Shen J, Wang Z, Hong B, Jiang B, et al. Mintaimycins, a Group of Novel Peptide Metabolites from Micromonospora sp. C-3509. Molecules. 2022; 27(4):1150. https://doi.org/10.3390/molecules27041150
Chicago/Turabian StyleHu, Xiaomin, Ying Wang, Chunyan Zhao, Shufen Li, Xinxin Hu, Xuefu You, Jiajia Shen, Zhen Wang, Bin Hong, Bingya Jiang, and et al. 2022. "Mintaimycins, a Group of Novel Peptide Metabolites from Micromonospora sp. C-3509" Molecules 27, no. 4: 1150. https://doi.org/10.3390/molecules27041150






