Rapid Degradation of SARS-CoV-2 Spike S Protein by A Specific Serine Protease
Abstract
:1. Introduction
2. Results
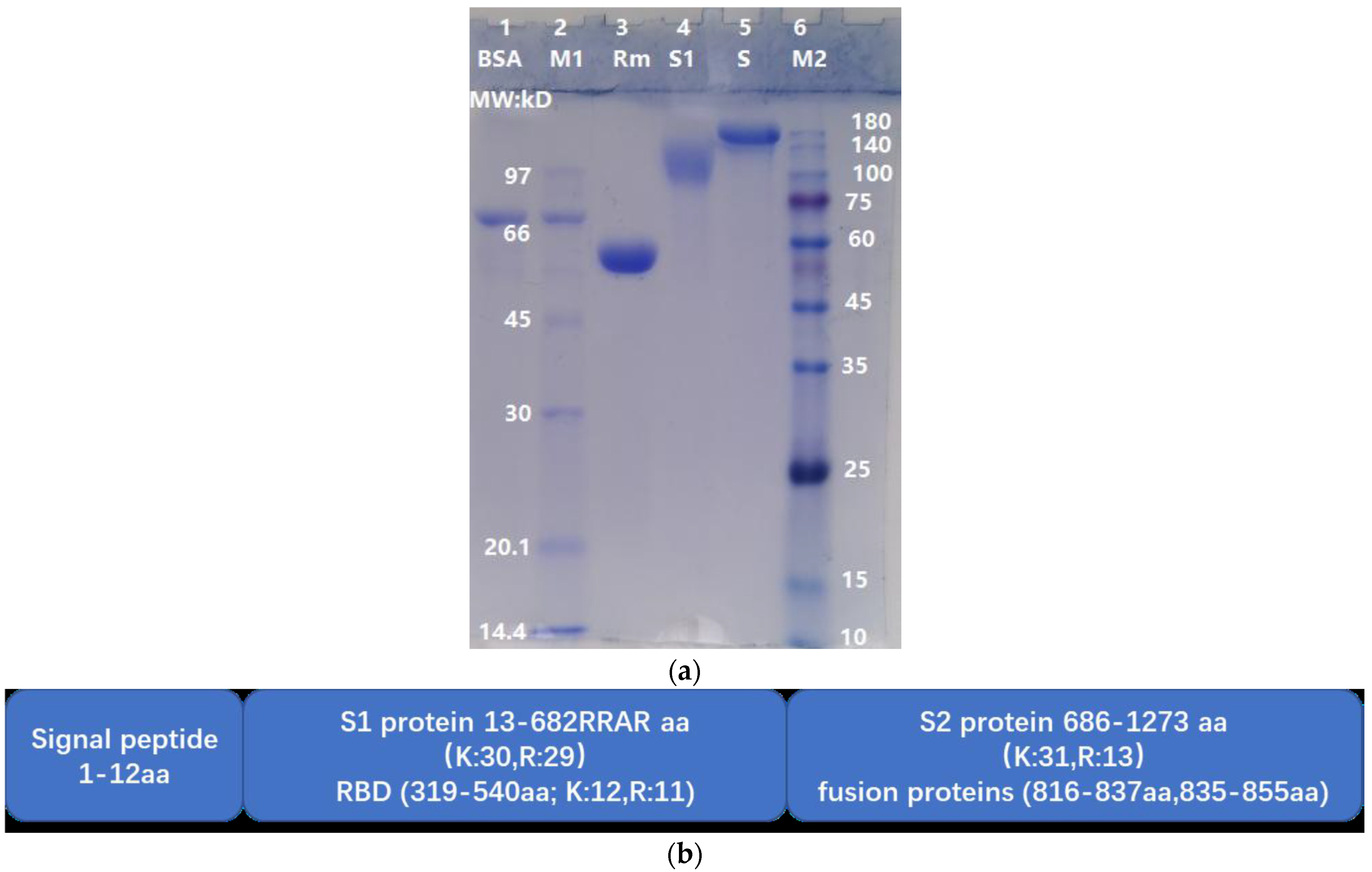
2.1. Comparison of the Molecular Weight of SARS-CoV-2 Spike Protein (S Full Length, S1 Subunit and RBD-mouseFc)
2.2. Dose-Dependent Degradation of S, S1, and RBD-mouseFc Proteins by ASPNJ
2.3. Time-Dependent Degradation of S, S1, and RBD-mouseFc Proteins by ASPNJ
2.4. Much Lower Degradation Effect of ASPNJ on Bovine Serum Albumin, BSA
2.5. Weak Degradation Effect of Trypsin on S1 Protein
2.6. The Structural Characteristics of SARS-CoV-2 Spike Protein and the Inference of ASPNJ Activity
3. Discussions
4. Materials and Methods
4.1. Viral Proteins of SARS-CoV-2
4.2. Alkaline Serine Protease with an Acidic pI (ASPNJ) and Concentration Determination
4.3. Other Reagents
4.4. SARS-CoV-2 Spike Protein (S Full Length, S1 Subunit and RBD-mouseFc) Degradation and SDS-PAGE Observation
4.5. Comparison of the Structural Characteristics and Properties of SARS-CoV-2 Wild-Type and Variant Strains
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Sample Availability Statement
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.-M.; Wang, W.; Song, Z.-G.; Hu, Y.; Tao, Z.-W.; Tian, J.-H.; Pei, Y.-Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Imran, M. The SARS-CoV-2 Spike Glycoprotein as a Drug and Vaccine Target: Structural Insights into Its Complexes with ACE2 and Antibodies. Cells 2020, 22, 2343. [Google Scholar] [CrossRef]
- Samrat, S.K.; Tharappel, A.M.; Li, Z.; Li, H. Prospect of SARS-CoV-2 spike protein: Potential role in vaccine and therapeutic development. Virus Res. 2020, 288, 198141. [Google Scholar] [CrossRef] [PubMed]
- Minol. Enhancing Readiness for Omicron (B.1.1.529): Technical Brief and Priority Actions for Member States. Zenodo 2021, 1–8. [Google Scholar] [CrossRef]
- Wang, R.; Zhang, Q.; Ge, J.; Ren, W.; Zhang, R.; Lan, J.; Ju, B.; Su, B.; Yu, F.; Chen, P.; et al. Analysis of SARS-CoV-2 variant mutations reveals neutralization escape mechanisms and the ability to use ACE2 receptors from additional species. Immunity 2021, 54, 1611–1621.e5. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.W.X.; Chiew, C.J.; Ang, L.W.; Mak, T.-M.; Cui, L.; Toh, M.P.H.S.; Lim, Y.D.; Lee, P.H.; Lee, T.H.; Chia, P.Y.; et al. Clinical and Virological Features of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Variants of Concern: A Retrospective Cohort Study Comparing B.1.1.7 (Alpha), B.1.351 (Beta), and B.1.617.2 (Delta). Clin. Infect. Dis. 2021, 23, ciab721. [Google Scholar] [CrossRef]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus. Cell 2020, 182, 812–827.e19. [Google Scholar] [CrossRef]
- Mallavarpu Ambrose, J.; Priya Veeraraghavan, V.; Kullappan, M.; Chellapandiyan, P.; Krishna Mohan, S.; Manivel, V.A. Comparison of Immunological Profiles of SARS-CoV-2 Variants in the COVID-19 Pandemic Trends: An Immunoinformatics Approach. Antibiotics 2021, 10, 535. [Google Scholar] [CrossRef]
- Hoffmann, M.; Arora, P.; Groß, R.; Seidel, A.; Hörnich, B.F.; Hahn, A.S.; Krüger, N.; Graichen, L.; Hofmann-Winkler, H.; Kempf, A.; et al. SARS-CoV-2 variants B.1.351 and P.1 escape from neutralizing antibodies. Cell 2021, 184, 2384–2393.e12. [Google Scholar] [CrossRef]
- Tse, L.V.; Meganck, R.M.; Graham, R.L.; Baric, R.S. The Current and Future State of Vaccines, Antivirals and Gene Therapies Against Emerging Coronaviruses. Front. Microbiol. 2020, 11, 658. [Google Scholar] [CrossRef] [PubMed]
- Barnes, C.O.; Jette, C.A.; Abernathy, M.E.; Dam, K.-M.A.; Esswein, S.R.; Gristick, H.B.; Malyutin, A.G.; Sharaf, N.G.; Huey-Tubman, K.E.; Lee, Y.E.; et al. SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature 2020, 588, 682–687. [Google Scholar] [CrossRef]
- Elshabrawy, H.A. SARS-CoV-2: An Update on Potential Antivirals in Light of SARS-CoV Antiviral Drug Discoveries. Vaccines 2020, 8, 335. [Google Scholar] [CrossRef] [PubMed]
- Giannis, D.; Ziogas, I.A.; Gianni, P. Coagulation disorders in coronavirus infected patients: COVID-19, SARS-CoV-1, MERS-CoV and lessons from the past. J. Clin. Virol. 2020, 127, 104362. [Google Scholar] [CrossRef]
- Lindahl, U.; Li, J. Heparin—An old drug with multiple potential targets in Covid-19 therapy. J. Thromb. Haemost. 2020, 18, 2422–2424. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Du, Y.; Kaltashov, I.A. The Utility of Native MS for Understanding the Mechanism of Action of Repurposed Therapeutics in COVID-19: Heparin as a Disruptor of the SARS-CoV-2 Interaction with Its Host Cell Receptor. Anal. Chem. 2020, 92, 10930–10934. [Google Scholar] [CrossRef] [PubMed]
- Deng, Z.; Wang, S.; Li, Q.; Ji, X.; Zhang, L.; Hong, M. Purification and characterization of a novel fibrinolytic enzyme from the polychaete, Neanthes japonica (Iznka). Bioresour. Technol. 2010, 101, 1954–1960. [Google Scholar] [CrossRef]
- Wang, S.-H.; Li, Q.; Deng, Z.-H.; Ji, X.; Jiang, X.; Ge, X.; Bo, Q.-Q.; Cui, J.-Y.; Zhang, L.-Z.; Liu, J.-K.; et al. Neanthes japonica (Iznka) fibrinolytic enzyme reduced cerebral infarction, cerebral edema and increased antioxidation in rat models of focal cerebral ischemia. Neurosci. Lett. 2011, 489, 16–19. [Google Scholar] [CrossRef]
- Ge, X.; Bo, Q.; Hong, X.; Cui, J.; Jiang, X.; Hong, M.; Liu, J. A novel acidic serine protease, ASPNJ inhibits proliferation, induces apoptosis and enhances chemo-susceptibility of acute promyelocytic leukemia cell. Leuk. Res. 2013, 37, 1697–1703. [Google Scholar] [CrossRef]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef] [Green Version]
- Daly, J.L.; Simonetti, B.; Klein, K.; Chen, K.-E.; Kavanagh Williamson, M.; Antón-Plágaro, C.; Shoemark, D.K.; Simón-Gracia, L.; Bauer, M.; Hollandi, R.; et al. Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science 2020, 370, 861–865. [Google Scholar] [CrossRef]
- Pyeon, D.; Timani, K.A.; Gulraiz, F.; He, J.J.; Park, I.-W. Function of ubiquitin (Ub) specific protease 15 (USP15) in HIV-1 replication and viral protein degradation. Virus Res. 2016, 223, 161–169. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pyeon, D.; Rojas, V.K.; Price, L.; Kim, S.; Singh, M.; Park, I.-W. HIV-1 Impairment via UBE3A and HIV-1 Nef Interactions Utilizing the Ubiquitin Proteasome System. Viruses 2019, 11, 1098. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Q.; Xiang, R.; Huo, S.; Zhou, Y.; Jiang, S.; Wang, Q.; Yu, F. Molecular mechanism of interaction between SARS-CoV-2 and host cells and interventional therapy. Signal Transduct. Target. Ther. 2021, 6, 233. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Xiao, T.; Cai, Y.; Chen, B. Structure of SARS-CoV-2 spike protein. Curr. Opin. Virol. 2021, 50, 173–182. [Google Scholar] [CrossRef]
- Zhou, T.; Tsybovsky, Y.; Gorman, J.; Rapp, M.; Cerutti, G.; Chuang, G.-Y.; Katsamba, P.S.; Sampson, J.M.; Schön, A.; Bimela, J.; et al. Cryo-EM Structures of SARS-CoV-2 Spike without and with ACE2 Reveal a pH-Dependent Switch to Mediate Endosomal Positioning of Receptor-Binding Domains. Cell Host Microbe 2020, 28, 867–879.e5. [Google Scholar] [CrossRef]
- Qiao, B.; de la Cruz, M.O. Enhanced Binding of SARS-CoV-2 Spike Protein to Receptor by Distal Polybasic Cleavage Sites. ACS Nano 2020, 14, 10616–10623. [Google Scholar] [CrossRef]
- Benton, D.J.; Wrobel, A.G.; Xu, P.; Roustan, C.; Martin, S.R.; Rosenthal, P.B.; Skehel, J.J.; Gamblin, S.J. Receptor binding and priming of the spike protein of SARS-CoV-2 for membrane fusion. Nature 2020, 588, 327–330. [Google Scholar] [CrossRef]
- Ghosh, N.; Nandi, S.; Saha, I. A review on evolution of emerging SARS-CoV-2 variants based on spike glycoprotein. Int. Immunopharmacol. 2022, 105, 108565. [Google Scholar] [CrossRef]
- Kumar, S.; Thambiraja, T.S.; Karuppanan, K.; Subramaniam, G. Omicron and Delta variant of SARS-CoV-2: A comparative computational study of spike protein. J. Med. Virol. 2021, 94, 1641–1649. [Google Scholar] [CrossRef]
- Stoscheck, C.M. Quantitation of protein. Methods Enzymol. 1990, 182, 50–68. [Google Scholar] [CrossRef]
- Brunelle, J.L.; Green, R. One-dimensional SDS-Polyacrylamide Gel Electrophoresis (1D SDS-PAGE). Methods Enzymol. 2014, 541, 151–159. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, J.; Kan, M.; Zhang, L.; Yue, Y.; Wang, S.; Hong, M.; Hong, X. Rapid Degradation of SARS-CoV-2 Spike S Protein by A Specific Serine Protease. Molecules 2022, 27, 1882. https://doi.org/10.3390/molecules27061882
Liu J, Kan M, Zhang L, Yue Y, Wang S, Hong M, Hong X. Rapid Degradation of SARS-CoV-2 Spike S Protein by A Specific Serine Protease. Molecules. 2022; 27(6):1882. https://doi.org/10.3390/molecules27061882
Chicago/Turabian StyleLiu, Jiankai, Mujie Kan, Lianzhi Zhang, Yuan Yue, Shaohua Wang, Min Hong, and Xinyu Hong. 2022. "Rapid Degradation of SARS-CoV-2 Spike S Protein by A Specific Serine Protease" Molecules 27, no. 6: 1882. https://doi.org/10.3390/molecules27061882




