Computational Screening of Newly Designed Compounds against Coxsackievirus A16 and Enterovirus A71
Abstract
:1. Introduction
2. Results and Discussion
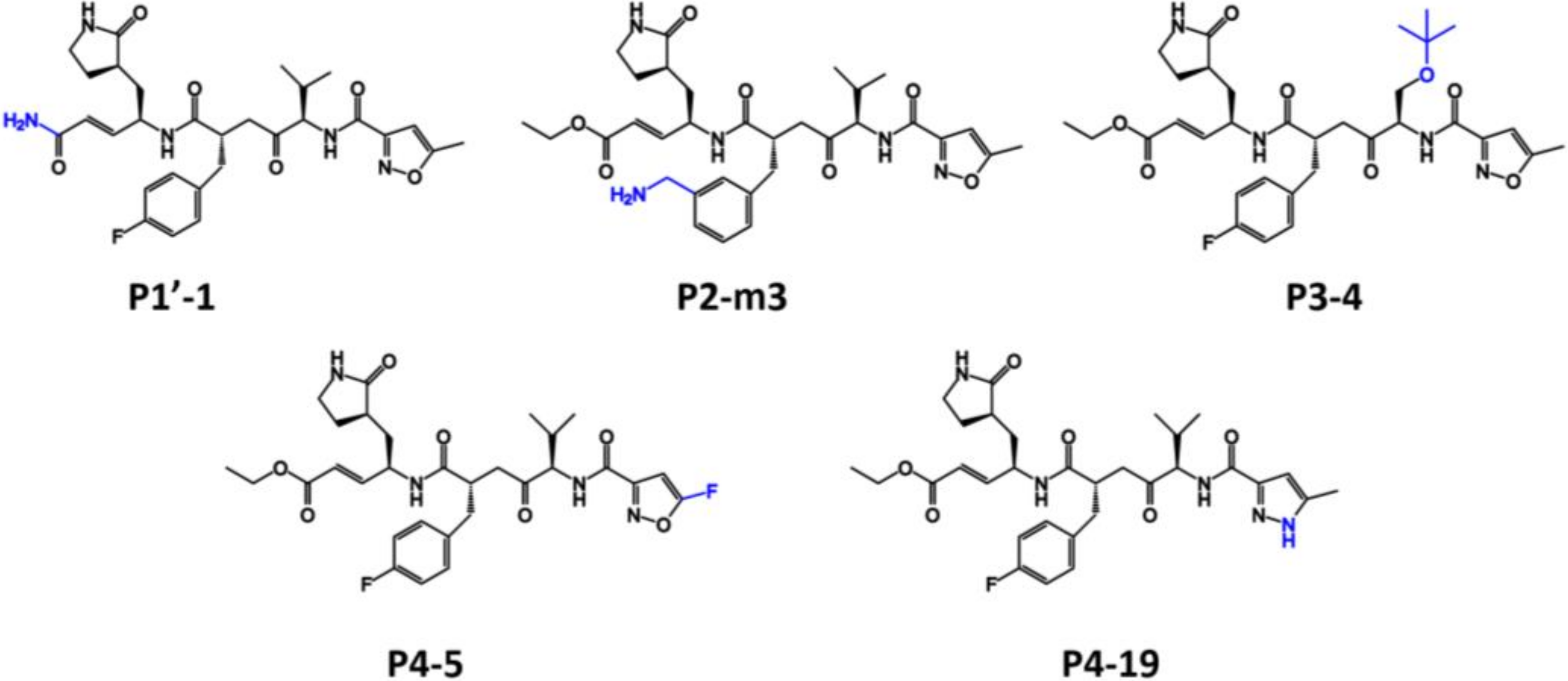
2.1. Rational Design and Screening
2.2. Stability of the Simulated Complexes
2.3. Number of Contact Atoms and H-Bonds
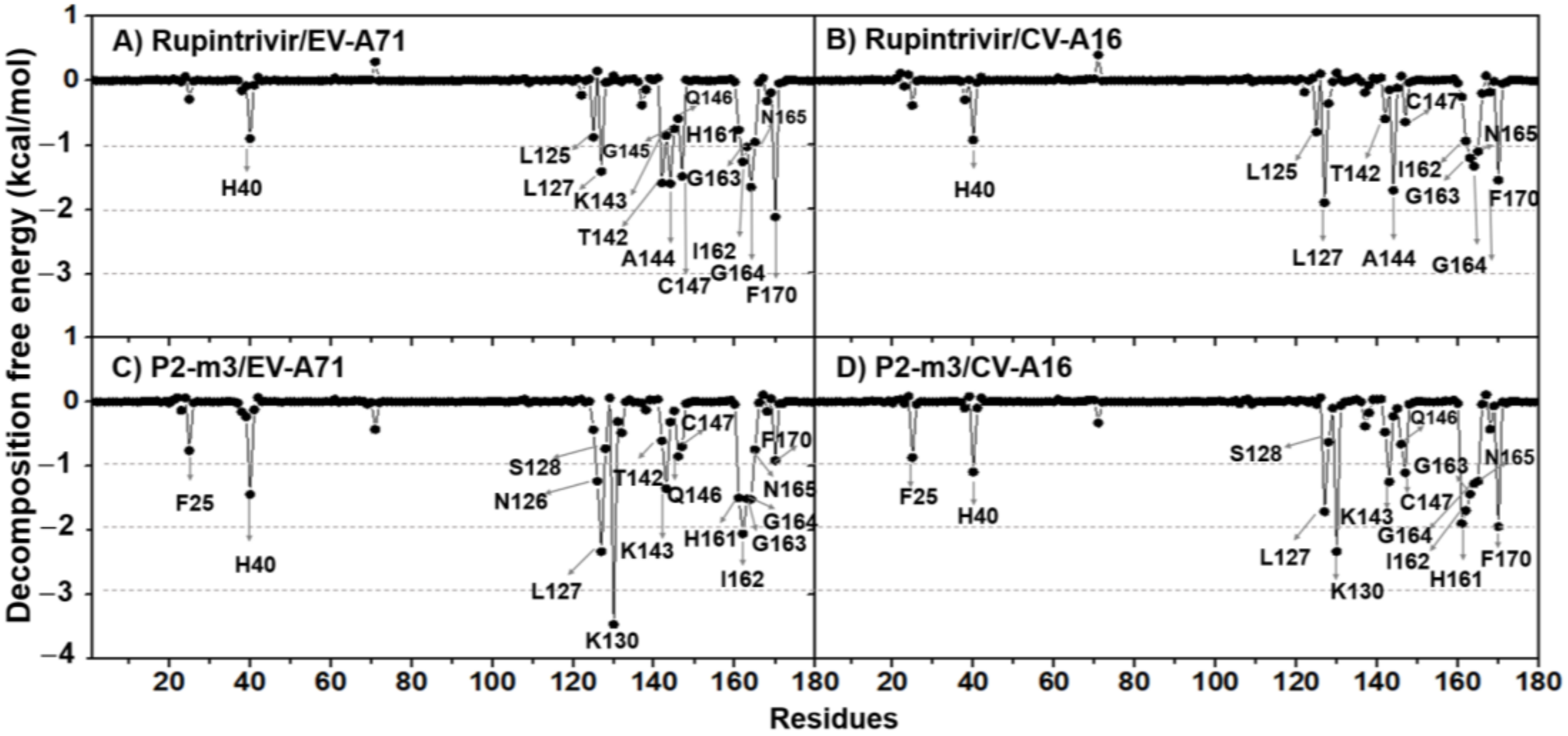
2.4. Key Binding Residues
2.5. Predicted Binding Affinity of the Potent Rupintrivir Analog
3. Materials and Methods
3.1. System Preparation and Compound Screening
3.2. Molecular Dynamics Simulations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Zhu, R.N.; Qian, Y.; Deng, J.; Xing, J.F.; Zhao, L.Q.; Wang, F.; Liao, B.; Ren, X.X.; Li, Y.; Zhang, Q. Study on the association of hand, foot and mouth disease and enterovirus 71/CA16 among children in Beijing, 2007. Zhonghua Liu Xing Bing Xue Za Zhi (Zhonghua Liuxingbingxue Zazhi) 2007, 28, 1004–1008. [Google Scholar] [PubMed]
- Sano, T.; Makino, N.; Aoyama, Y.; Ae, R.; Kojo, T.; Kotani, K.; Yanagawa, H. Temporal and geographical clustering of Kawasaki disease in Japan: 2007–2012. Pediatr. Int. 2016, 58, 1140–1145. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.Y.; Zhang, X.; Chung, P.H.; Tsao, K.C.; Lin, T.Y.; Su, L.H.; Chiu, C.H. Enterovirus 71 in Taiwan, 2004–2006: Epidemiological and virological features. Scand. J. Infect. Dis. 2008, 40, 571–574. [Google Scholar] [CrossRef]
- Chan, K.P.; Goh, K.T.; Chong, C.Y.; Teo, E.S.; Lau, G.; Ling, A.E. Epidemic hand, foot and mouth disease caused by human enterovirus 71, Singapore. Emerg. Infect. Dis. 2003, 9, 78. [Google Scholar] [CrossRef] [PubMed]
- Puenpa, J.; Chieochansin, T.; Linsuwanon, P.; Korkong, S.; Thongkomplew, S.; Vichaiwattana, P.; Theamboonlers, A.; Poovorawan, Y. Hand, foot, and mouth disease caused by coxsackievirus A6, Thailand, 2012. Emerg. Infect. Dis. 2013, 19, 641. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sarma, N. Hand, foot, and mouth disease: Current scenario and Indian perspective. Indian J. Dermatol. Venereol. Leprol. 2013, 79, 165–175. [Google Scholar] [CrossRef]
- Tsai, M.T.; Cheng, Y.H.; Liu, Y.N.; Liao, N.C.; Lu, W.W.; Kung, S.H. Real-time monitoring of human enterovirus (HEV)-infected cells and anti-HEV 3C protease potency by fluorescence resonance energy transfer. Antimicrob. Agents Chemother. 2019, 53, 748–755. [Google Scholar] [CrossRef] [Green Version]
- Omaña-Cepeda, C.; Martínez-Valverde, A.; Del Mar Sabater-Recolons, M.; Jané-Salas, E.; Marí-Roig, A.; López-López, J. A literature review and case report of hand, foot and mouth disease in an immunocompetent adult. BMC Res. Notes 2016, 9, 165. [Google Scholar] [CrossRef] [Green Version]
- Guerra, A.M.; Orille, E.; Waseem, M. Hand Foot and Mouth Disease, StatPearls. 2021. Available online: https://www.ncbi.nlm.nih.gov/books/NBK431082/ (accessed on 24 October 2021).
- Ji, T.; Han, T.; Tan, X.; Zhu, S.; Yan, D.; Yang, Q.; Song, Y.; Cui, A.; Zhang, Y.; Mao, N.; et al. Surveillance, epidemiology, and pathogen spectrum of hand, foot, and mouth disease in mainland of China from 2008 to 2017. Biosafety Health 2019, 1, 32–40. [Google Scholar] [CrossRef]
- Long, L.; Xu, L.; Xiao, Z.; Hu, S.; Luo, R.; Wang, H.; Lu, X.; Xu, Z.; Yao, X.; Zhou, L.; et al. Neurological complications and risk factors of cardiopulmonary failure of EV-A71-related hand, foot and mouth disease. Sci. Rep. 2016, 6, 23444. [Google Scholar]
- Da Silva, E.E.; Winkler, M.T.; Pallansch, M.A. Role of enterovirus 71 in acute flaccid paralysis after the eradication of poliovirus in Brazil. Emerg. Infect. Dis. 1996, 2, 231–233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, G.; Qi, J.; Chen, Z.; Xu, X.; Gao, F.; Lin, D.; Qian, W.; Liu, H.; Jiang, H.; Yan, J.; et al. Enterovirus 71 and coxsackievirus A16 3C proteases: Binding to rupintrivir and their substrates and anti-hand, foot, and mouth disease virus drug design. J. Virol. 2011, 85, 10319–10331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Fan, T.; Yao, X.; Wu, Z.; Guo, L.; Lei, X.; Wang, M.; Jin, Q.; Cui, S. Crystal structures of enterovirus 71 3C protease complexed with rupintrivir reveal the roles of catalytically important residues. J. Virol. 2011, 85, 10021–10030. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuo, C.J.; Shie, J.J.; Fang, J.M.; Yen, G.R.; Hsu, J.T.A.; Liu, H.G.; Tseng, S.N.; Chang, S.C.; Lee, C.Y.; Shih, S.R.; et al. Design, synthesis, and evaluation of 3C protease inhibitors as anti-enterovirus 71 agents. Bioorg. Med. Chem. 2008, 16, 7388–7398. [Google Scholar] [CrossRef]
- Patick, A.K.; Binford, S.L.; Brothers, M.A.; Jackson, R.L.; Ford, C.E.; Diem, M.D.; Maldonado, F.; Dragovich, P.S.; Zhou, R.; Prins, T.J.; et al. In vitro antiviral activity of AG7088, a potent inhibitor of human rhinovirus 3C protease. Antimicrob. Agents Chemother. 1999, 43, 2444–2450. [Google Scholar] [CrossRef] [Green Version]
- Ooi, M.H.; Wong, S.C.; Lewthwaite, P.; Cardosa, M.J.; Solomon, T. Clinical features, diagnosis, and management of enterovirus 71. Lancet Neurol. 2010, 9, 1097–1105. [Google Scholar] [CrossRef]
- Lee, J.C.; Shih, S.R.; Chang, T.Y.; Tseng, H.Y.; Shih, Y.F.; Yen, K.J.; Chen, W.C.; Shie, J.J.; Fang, J.M.; Liang, P.H.; et al. A mammalian cell-based reverse two-hybrid system for functional analysis of 3C viral protease of human enterovirus 71. Anal. Biochem. 2008, 375, 115–123. [Google Scholar] [CrossRef]
- Zhang, X.; Song, Z.; Qin, B.; Zhang, X.; Chen, L.; Hu, Y.; Yuan, Z. Rupintrivir is a promising candidate for treating severe cases of enterovirus-71 infection: Evaluation of antiviral efficacy in a murine infection model. Antivir. Res. 2013, 97, 264–269. [Google Scholar] [CrossRef]
- Binford, S.L.; Maldonado, F.; Brothers, M.A.; Weady, P.T.; Zalman, L.S.; Meador, J.W.; Matthews, D.A.; Patick, A.K. Conservation of amino acids in human rhinovirus 3C protease correlates with broad-spectrum antiviral activity of rupintrivir, a novel human rhinovirus 3C protease inhibitor. Antimicrob. Agents Chemother. 2005, 49, 619–626. [Google Scholar] [CrossRef] [Green Version]
- Matthews, D.A.; Dragovich, P.S.; Webber, S.E.; Fuhrman, S.A.; Patick, A.K.; Zalman, L.S.; Hendrickson, T.F.; Love, R.A.; Prins, T.J.; Marakovits, J.T.; et al. Structure-assisted design of mechanism-based irreversible inhibitors of human rhinovirus 3C protease with potent antiviral activity against multiple rhinovirus serotypes. Proc. Natl. Acad. Sci. USA 1999, 96, 11000–11007. [Google Scholar] [CrossRef] [Green Version]
- Hayden, F.G.; Turner, R.B.; Gwaltney, J.M.; Chi-Burris, K.; Gersten, M.; Hsyu, P.; Patick, A.K.; Smith, G.J., III; Zalman, L.S. Phase II, randomized, double-blind, placebo-controlled studies of ruprintrivir nasal spray 2-percent suspension for prevention and treatment of experimentally induced rhinovirus colds in healthy volunteers. Antimicrob. Agents Chemother. 2003, 47, 3907–3916. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schmidt, N.J.; Lennette, E.H.; Ho, H.H. An apparently new enterovirus isolated from patients with disease of the central nervous system. J. Infect. Dis. 1974, 129, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Meijer, A.; Froeyen, M.; Zhang, L.; Thibaut, H.J.; Baggen, J.; George, S.; Vernachio, J.; Van Kuppeveld, F.J.M.; Leyssen, P.; et al. Antiviral activity of broad-spectrum and enterovirus-specific inhibitors against clinical isolates of enterovirus D68. Antimicrob. Agents Chemother. 2015, 59, 7782–7785. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dragovich, P.S.; Prins, T.J.; Zhou, R.; Webber, S.E.; Marakovits, J.T.; Fuhrman, S.A.; Patick, A.K.; Matthews, D.A.; Lee, C.A.; Ford, C.E.; et al. Structure-based design, synthesis, and biological evaluation of irreversible human rhinovirus 3C protease inhibitors. 4. Incorporation of P1 lactam moieties as L-glutamine replacements. J. Med. Chem. 1999, 42, 1213–1224. [Google Scholar] [CrossRef] [PubMed]
- Lacroix, C.; George, S.; Leyssen, P.; Hilgenfeld, R.; Neyts, J. The enterovirus 3C protease inhibitor SG85 efficiently blocks rhinovirus replication and is not cross-resistant with rupintrivir. Antimicrob. Agents Chemother. 2015, 59, 5814–5818. [Google Scholar] [CrossRef] [Green Version]
- Jetsadawisut, W.; Nutho, B.; Meeprasert, A.; Rungrotmongkol, T.; Kungwan, N.; Wolschann, P.; Hannongbua, S. Susceptibility of inhibitors against 3C protease of coxsackievirus A16 and enterovirus A71 causing hand, foot and mouth disease: A molecular dynamics study. Biophys. Chem. 2016, 219, 9–16. [Google Scholar] [CrossRef]
- Wang, Y.; Cao, L.; Zhai, Y.; Yin, Z.; Sun, Y.; Shang, L. Structure of the Enterovirus 71 3C Protease in Complex with NK-1.8 k and Indications for the Development of Antienterovirus Protease Inhibitor. Antimicrob. Agents Chemother. 2017, 61, e00298-17. [Google Scholar] [CrossRef] [Green Version]
- Hildebrand, P.W.; Rose, A.S.; Tiemann, J.K. Bringing molecular dynamics simulation data into view. Trends Biochem. Sci. 2019, 44, 902–913. [Google Scholar] [CrossRef] [Green Version]
- Meng, X.D.; Tong, Y.; Wei, Z.N.; Wang, L.; Mai, J.Y.; Wu, Y.; Luo, Z.-Y.; Li, S.; Li, M.; Wang, S.; et al. Epidemical and etiological study on hand, foot and mouth disease following EV-A71 vaccination in Xiangyang, China. Sci. Rep. 2020, 10, 20909. [Google Scholar] [CrossRef]
- Olsson, M.H.; Søndergaard, C.R.; Rostkowski, M.; Jensen, J.H. PROPKA3: Consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 2011, 7, 525–537. [Google Scholar] [CrossRef]
- Bleiziffer, P.; Schaller, K.; Riniker, S. Machine learning of partial charges derived from high-quality quantum-mechanical calculations. J. Chem. Inf. Modeling 2018, 58, 579–590. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- MacKerell, A.D., Jr. Empirical force fields for biological macromolecules: Overview and issues. J. Comput. Chem. 2004, 25, 1584–1604. [Google Scholar] [CrossRef] [PubMed]
- Case, D.A.; Cheatham, T.E., III; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.M., Jr.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber biomolecular simulation programs. J. Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frisch, M.J.E.A. Gaussian09. 2009. Available online: http://www.gaussian.com/ (accessed on 16 July 2021).
- Case, D.A.; Babin, V.; Berryman, J.T.; Betz, R.M.; Cai, Q.; Cerutti, D.S.; Cheatham, T.E.; Darden, I.T.A.; Duke, R.E.; Gohlke, H.; et al. AMBER 16; University of California: San Francisco, CA, USA, 2016. [Google Scholar]
- Da Silva, A.W.S.; Vranken, W.F. ACPYPE-Antechamber python parser interface. BMC Res. Notes 2012, 5, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Wittayanarakul, K.; Aruksakunwong, O.; Saen-oon, S.; Chantratita, W.; Parasuk, V.; Sompornpisut, P.; Hannongbua, S. Insights into saquinavir resistance in the G48V HIV-1 protease: Quantum calculations and molecular dynamic simulations. Biophys. J. 2005, 88, 867–879. [Google Scholar] [CrossRef] [Green Version]
- Tan, J.; George, S.; Kusov, Y.; Perbandt, M.; Anemüller, S.; Mesters, J.R.; Hilgenfeld, R. 3C protease of enterovirus 68: Structure-based design of Michael acceptor inhibitors and their broad-spectrum antiviral effects against picornaviruses. J. Virol. 2013, 87, 4339–4351. [Google Scholar] [CrossRef] [Green Version]
- Knapp, B.; Frantal, S.; Cibena, M.; Schreiner, W.; Bauer, P. Is an intuitive convergence definition of molecular dynamics simulations solely based on the root mean square deviation possible? J. Comput. Biol. 2011, 18, 997–1005. [Google Scholar] [CrossRef] [Green Version]
- Roe, D.R.; Cheatham, T.E., III. PTRAJ and CPPTRAJ: Software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 2013, 9, 3084–3095. [Google Scholar] [CrossRef] [PubMed]






| Compound | Compound | ||||
|---|---|---|---|---|---|
| MM/PBSA | MM/GBSA | MM/PBSA | MM/GBSA | ||
| P1′-1 | −9.14 | −6.5 | P2-m9 | 0.4 | 0.94 |
| P1′-2 | 0.17 | 4.98 | P2-m10 | 3.25 | 8.96 |
| P1-1 | 7.82 | 5.02 | P3-1 | 0.43 | 3.24 |
| P1-2 | 20.41 | 17.58 | P3-2 | 5.67 | 7.82 |
| P1-3 | 1.37 | 1.91 | P3-3 | 2.54 | 4.15 |
| P1-4 | 8.28 | 4.21 | P3-4 | −4.9 | −2.54 |
| P1-5 | 17.76 | 14.31 | P4-1 | 1.98 | 6.27 |
| P2-p1 | 6.38 | 5.49 | P4-2 | 1.73 | 4.25 |
| P2-p2 | 14.17 | 14.15 | P4-3 | 2.28 | 7.36 |
| P2-p3 | 1.48 | 2.14 | P4-4 | 12.74 | 12.79 |
| P2-p4 | 7.55 | 9.22 | P4-5 | −5.34 | −5.68 |
| P2-p5 | 7.85 | 11.03 | P4-6 | 4.46 | 2.51 |
| P2-p6 | 0.73 | 3.55 | P4-7 | 4.65 | 4.29 |
| P2-p7 | 1.37 | 5.18 | P4-8 | 3.32 | 2.94 |
| P2-p8 | 6.65 | 7.74 | P4-9 | 6.58 | 4.92 |
| P2-p9 | 0.65 | 1.38 | P4-10 | 0.75 | 1.27 |
| P2-p10 | 1.56 | 3.93 | P4-11 | 4.89 | 3.34 |
| P2-m1 | 20.47 | 18.36 | P4-12 | 13.28 | 11.89 |
| P2-m2 | 18.59 | 17.29 | P4-13 | 1.57 | 0.47 |
| P2-m3 | −12.12 | −4.2 | P4-14 | 1.64 | 0.96 |
| P2-m4 | 2.41 | 6.29 | P4-15 | 12.68 | 11.51 |
| P2-m5 | 8.74 | 12.38 | P4-16 | 1.55 | 1.12 |
| P2-m6 | 1.75 | 1.35 | P4-17 | 3.03 | 2.52 |
| P2-m7 | 0.49 | 2.03 | P4-18 | 4.12 | 4.44 |
| P2-m8 | 1.98 | 4.71 | P4-19 | −5.55 | −6.68 |
| CV-A16 | EV-A71 | |
|---|---|---|
| P4-5 | 22.5 ± 5.0 | 16.1 ± 4.7 |
| P4-19 | 22.1 ± 5.3 | 17.4 ± 4.8 |
| P1′-1 | 19.3 ± 4.3 | 17.8 ± 4.5 |
| P2-m3 | 24.3 ± 4.3 | 22.1 ± 4.5 |
| P3-4 | 17.9 ± 4.8 | 21.6 ± 4.8 |
| Rupintrivir | 23.7 ± 4.7 | 20.0 ± 5.0 |
| CV-A16 | EV-A71 | |
|---|---|---|
| P4-5 | 4.8 ± 1.1 | 3.6 ± 1.3 |
| P4-19 | 4.0 ± 1.2 | 4.4 ± 1.2 |
| P1′-1 | 4.0 ± 1.0 | 5.1 ± 1.1 |
| P2-m3 | 6.5 ± 1.2 | 5.6 ± 1.5 |
| P3-4 | 3.3 ± 1.3 | 4.8 ± 1.1 |
| Rupintrivir | 4.5 ± 0.9 | 4.7 ± 1.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sripattaraphan, A.; Sanachai, K.; Chavasiri, W.; Boonyasuppayakorn, S.; Maitarad, P.; Rungrotmongkol, T. Computational Screening of Newly Designed Compounds against Coxsackievirus A16 and Enterovirus A71. Molecules 2022, 27, 1908. https://doi.org/10.3390/molecules27061908
Sripattaraphan A, Sanachai K, Chavasiri W, Boonyasuppayakorn S, Maitarad P, Rungrotmongkol T. Computational Screening of Newly Designed Compounds against Coxsackievirus A16 and Enterovirus A71. Molecules. 2022; 27(6):1908. https://doi.org/10.3390/molecules27061908
Chicago/Turabian StyleSripattaraphan, Amita, Kamonpan Sanachai, Warinthorn Chavasiri, Siwaporn Boonyasuppayakorn, Phornphimon Maitarad, and Thanyada Rungrotmongkol. 2022. "Computational Screening of Newly Designed Compounds against Coxsackievirus A16 and Enterovirus A71" Molecules 27, no. 6: 1908. https://doi.org/10.3390/molecules27061908







