Comparative Molecular Field Analysis (CoMFA) and Comparative Molecular Similarity Indices Analysis (CoMSIA) Studies on α1A-Adrenergic Receptor Antagonists Based on Pharmacophore Molecular Alignment
Abstract
:1. Introduction
2. Materials and Method
2.1. Data Collection
2.2. Structural Sketch and Alignment
2.3. 3D-QSAR Studies
2.4. Predictive Power of the Models
3. Results and Discussion
3.1. Alignment
3.2. CoMFA and CoMSIA Models
3.3. Predictive Power of the Models
3.4. Graphical Interpretation of the Fields
4. Conclusions
Acknowledgments
References
- Docherty, JR. Subtypes of functional α1- and α2-adrenoceptors. Eur. J. Pharmacol 1998, 361, 1–15. [Google Scholar]
- Oesterling, JE. Benign prostatic hyperplasia-medical and minimally invasive treatment options. N. Engl. J. Med 1995, 332, 99–110. [Google Scholar]
- Debnath, B; Samanta, S; Naskar, SK; Roy, K; Jha, T. QSAR study on the affinity of some arylpiperazines towards the 5-HT1A/α1-adrenergic receptor using the e-state index. Bioorg. Med. Chem. Lett 2003, 13, 2837–2842. [Google Scholar]
- Pallavicini, M; Fumagalli, L; Gobbi, M; Bolchi, C; Colleoni, S; Moroni, B; Pedretti, A; Rusconi, C; Vistoli, G; Valoti, E. QSAR study for novel series of ortho disubstituted phenoxy analogues of α1-adrenoceptor antagonist WB4101. Eur. J. Med. Chem 2006, 41, 1025–1040. [Google Scholar]
- Nowaczyk, A; Przybylski, R; Kulig, K; Malawska, B. Structure-activity relationship studies of a number of α1-adrenoceptor antagonists and antiarrhythmic agents. Mol. Inf 2010, 29, 343–351. [Google Scholar]
- Cinone, N; Carrieri, A; Strappaghetti, G; Corsano, S; Barbaro, R; Carotti, A. Comparative molecular field analysis of some pyridazinone-containing α1-Antagonists. Bioorg. Med. Chem 1999, 7, 2615–2620. [Google Scholar]
- Balle, T; Andersen, K; Søby, KK; Liljefors, T. α1-Adrenoceptor subtype selectivity: 3D-QSAR models for a new class of α1-adrenoceptor antagonists derived from the novel antipsychotic sertindole. J. Mol. Graph. Model 2003, 21, 523–534. [Google Scholar]
- Montorsi, M; Menziani, MC; Cocchi, M; Fanelli, F; De Benedetti, PG. Computer modeling of size and shape descriptors of α1-adrenergic receptor antagonists and quantitative structureaffinity/selectivity relationships. Methods 1998, 14, 239–254. [Google Scholar]
- Maciejewska, D; Zolek, T; Herold, F. CoMFA methodology in structure-activity analysis of hexahydro- and octahydropyrido[1,2-c]pyrimidine derivatives based on affinity towards 5-HT1A, 5-HT2A and α1-adrenergic receptors. J. Mol. Graph. Model 2003, 25, 353–362. [Google Scholar]
- Li, MY; Du, LP; Wu, B; Xia, L. Self-organizing molecular field analysis on α1a- adrenoceptor dihydropyridine antagonists. Bioorg. Med. Chem 2003, 11, 3945–3951. [Google Scholar]
- Shakya, N; Fatima, Z; Saxena, AK. 3D-QSAR Studies on substituted Dihydropyridin 3D-QSAR studies on substituted Dihydropyridines for their α1a-adrenergic receptor binding affinity. Med. Chem. Res 2004, 13, 812–823. [Google Scholar]
- Gupta, AK; Saxena, AK. 3D-QSAR CoMFA and CoMSIA studies on a set of diverse α1a-adrenergic receptor antagonists. Med Chem Res 2010. [Google Scholar] [CrossRef]
- Zhu, LL; Hou, TJ; Chen, LR; Xu, XJ. 3D-QSAR analyses of novel tyrosine kinase inhibitors based on pharmacophore alignment. J.Chem. Inf. Comput. Sci 2001, 41, 1032–1040. [Google Scholar]
- Chaudhaery, SS; Roy, KK; Saxena, AK. Consensus superiority of the pharmacophore- based alignment, over maximum common substructure (MCS): 3D-QSAR studies on carbamates as acetylcholinesterase inhibitors. J. Chem. Inf. Model 2009, 49, 1590–1601. [Google Scholar]
- Long, W; Liu, P; Li, Q; Xu, Y; Gao, J. 3D-QSAR studies on a class of ikk-2 inhibitors with galahad used to develop molecular alignment models. QSAR Comb. Sci 2008, 27, 1113–1119. [Google Scholar]
- Richmond, NJ; Abrams, CA; Wolohan, PR; Abrahamian, E; Willett, P; Clark, RD. GALAHAD: 1. pharmacophore identification by hypermolecular alignment of ligands in 3D. J. Comput.-Aided Mol. Des 2006, 20, 567–587. [Google Scholar]
- Brasili, L; Sorbi, C; Franchini, S; Manicardi, M; Angeli, P; Marucci, G; Leonardi, A; Poggesi, E. 1,3-Dioxolane-based ligands as a novel class of α1-adrenoceptor antagonists. J. Med. Chem 2003, 46, 1504–1511. [Google Scholar]
- Sorbi, C; Franchini, S; Tait, A; Prandi, A; Gallesi, R; Angeli, P; Marucci, G; Pirona, L; Poggesi, E; Brasili, L. 1,3-Dioxolane-based ligands as rigid analogues of naftopidil: Structureaffinity/activity relationships at α1 and 5-HT1A receptors. ChemMedChem 2009, 4, 393–399. [Google Scholar]
- Franchini, S; Tait, A; Prandi, A; Sorbi, C; Gallesi, R; Buccioni, M; Marucci, G; Stefani, CD; Cilia, A; Brasili, L. (2,2-Diphenyl-[1,3]oxathiolan-5-ylmethyl)- (3-phenyl-propyl)- amine: A potent and selective 5- HT1A receptor agonist. ChemMedChem 2009, 4, 196–203. [Google Scholar]
- Quaglia, W; Pigini, M; Piergentili, A; Giannella, M; Marucci, G; Poggesi, E; Leonardi, A; Melchiorre, C. Structure-activity relationships in 1,4-benzodioxan-related compounds. 6.1role of the dioxane unit on selectivity for α1-adrenoreceptor subtypes. J. Med. Chem 1999, 42, 2961–2968. [Google Scholar]
- Quaglia, W; Pigini, M; Piergentili, A; Giannella, M; Gentili, F; Marucci, G; Carrieri, A; Carotti, A; Poggesi, E; Leonardi, A; Melchiorre, C. Structure-activity relationships in 1,4- benzodioxan-related compounds. 7.1selectivity of 4-phenylchroman analogues for α1-adrenoreceptor subtypes. J. Med. Chem 2002, 45, 1633–1643. [Google Scholar]
- Quaglia, W; Santoni, G; Pigini, M; Piergentili, A; Gentili, F; Buccioni, M; Mosca, M; Lucciarini, R; Amantini, C; Nabissi, MI; et al. Structure-activity relationships in 1,4- benzodioxan-related compounds. 8.1 {2-[2-(4- chlorobenzyloxy)phenoxy]ethyl}-[2-(2,6- dimethoxyphenoxy)ethyl]amine (clopenphendioxan) as a tool to highlight the involvement of α1d- and α1b-adrenoreceptor subtypes in the regulation of human pc-3 prostate cancer cell apoptosis and proliferation. J. Med. Chem 2005, 48, 7750–7763. [Google Scholar]
- Quaglia, W; Piergentili, A; Del, BF; Farande, Y; Giannella, M; Pigini, M; Rafaiani, G; Carrieri, A; Amantini, C; Lucciarini, R; et al. Structure-activity relationships in 1,4-benzodioxan-related compounds. 9.1from 1,4-benzo- dioxane to 1,4-dioxane ring as a promising template of novel α1d-adrenoreceptor antagonists, 5-HT1A full agonists, and cytotoxic agents. J. Med. Chem 2008, 51, 6359–6370. [Google Scholar]
- Carrieri, A; Piergentili, A; Del, BF; Giannella, M; Pigini, M; Leonardi, A; Fanelli, F; Quaglia, W. Structure-activity relationships in 1,4-benzodioxan-related compounds. 10. Novel α1-adrenoreceptor antagonists related to openphendioxan: Synthesis, biological evaluation, and α1d computational study. Bioorg. Med. Chem 2010, 18, 7065–7077. [Google Scholar]
- Leonardi, A; Barlocco, D; Montesano, F; Cignarella, G; Motta, G; Testa, R; Poggesi, E; Seeber, M; De Benedetti, PG; Fanelli, F. Synthesis, screening, and molecular modeling of new potent and selective antagonists at the α1d adrenergic receptor. J. Med. Chem 2004, 47, 1900–1918. [Google Scholar]
- Saxena, AK; Prathipati, P. Comparison of MLR, PLS and GA-MLR in QSAR analysis. SAR QSAR Environ. Res 2003, 14, 433–445. [Google Scholar]
- Yung-Chi, C; Prusoff, WH. Relationship between the inhibition constant (KI) and the concentration of inhibitor which causes 50 per cent inhibition (I50) of an enzymatic reaction. Biochem. Pharmacol 1973, 22, 3099–3108. [Google Scholar]
- Clark, M; Cramer, RD; Opdenbosch, NV. Validation of the general purpose tripos 5.2 force field. J. Comput. Chem 1989, 10, 982–1012. [Google Scholar]
- Purcell, WP; Singer, JA. A brief review and table of semiempirical parameters used in the Hüeckel molecular orbital method. J. Chem. Eng. Data 1967, 12, 235–246. [Google Scholar]
- Zhao, X; Yuan, M; Huang, BY; Ji, H; Zhu, L. Ligand-based pharmacophore model of N-Aryl and N-Heteroaryl piperazine α1A-adrenoceptors antagonists using GALAHAD. J. Mol. Graph. Model 2010, 29, 126–136. [Google Scholar]
- Cramer, RD; Patterson, DE; Bunce, JD. Comparative molecular field analysis (CoMFA). 1. Effect of shape on binding of steroids to carrier proteins. J. Am. Chem. Soc 1988, 110, 5959–5967. [Google Scholar]
- Klebe, G; Abraham, U; Mietzner, T. Molecular similarity indices in a comparative analysis (CoMSIA) of drug molecules to correlate and predict their biological activity. J. Med. Chem 1994, 37, 4130–4146. [Google Scholar]
- Klebe, G; Abraham, U. Comparative molecular similarity index analysis (CoMSIA) to study hydrogen-bonding properties and to score combinatorial libraries. J. Comput.-Aided Mol. Des 1999, 13, 1–10. [Google Scholar]
- Wold, S. Validation of QSAR's. QSAR 1991, 10, 191–193. [Google Scholar]
- Shao, J. Linear model selection by cross-validation. J. Am. Stat. Assoc 1993, 88, 486–494. [Google Scholar]
- Cruciani, G; Baroni, M; Clementi, S; Costantino, G; Riganelli, D; Skagerberg, B. Predictive ability of regression models. Part I: Standard deviation of prediction errors (SDEP). J. Chemom 1992, 6, 335–346. [Google Scholar]
- Baroni, M; Clementi, S; Cruciani, G; Costantino, G; Riganelli, D; Oberrauch, E. Predictive ability of regression models. Part II: Selection of the best predictive PLS model. J. Chemom 1992, 6, 347–356. [Google Scholar]
- Tripos. GALAHAD® Manual. Available online: http://www.tripos.com accessed on 19 October 2011.
- Böhm, M; Strzebeche, J; Klebe, G. Three-dimensional quantitative structure-activity relationship analyses using comparative molecular field analysis and comparative molecular similarity indices analysis to elucidate selectivity differences of inhibitors binding to trypsin, thrombin, and factor Xa. J. Med. Chem 1999, 42, 458–477. [Google Scholar]
- Hattotuwagama, CK; Doytchinova, IA; Flower, DR. In silico prediction of peptide binding affinity to class i mouse major histocompatibility complexes: A comparative molecular similarity index analysis (CoMSIA) Study. J. Chem. Inf. Model 2005, 45, 1415–1423. [Google Scholar]
- Li, MY; Fang, H; Du, LP; Xia, L; Wang, BH. Computational studies of the binding site of α1A-adrenoceptor antagonists. J. Mol. Model 2008, 14, 957–966. [Google Scholar]
- Kinsella, GK; Rozas, I; Watson, GW. Computational study of antagonist/α1A adrenoceptor complexes--observations of conformational variations on the formation of ligand/receptor complexes. J. Med. Chem 2006, 49, 501–510. [Google Scholar]






| Series | Compd. No. | R1 | R2 | R3 | X | Ki(nM) | pKi |
|---|---|---|---|---|---|---|---|
| I | 1 | 4-Cl | −OCH3 | −OCH3 | O | 0.251 | 9.60 |
| 2 | 2-CH3 | −OCH3 | −OCH3 | O | 1.58 | 8.80 | |
| 3 | 3-CH3 | −OCH3 | −OCH3 | O | 0.398 | 9.40 | |
| 4 | 4-CH3 | −OCH3 | −OCH3 | O | 1.99 | 8.70 | |
| 5 | 3-OCH3 | −OCH3 | −OCH3 | O | 1.99 | 8.70 | |
| 6 | 4-OCH3 | −OCH3 | −OCH3 | O | 1.99 | 8.70 | |
| 7 | H | −OCH3 | −OCH3 | S | 3.16 | 8.50 | |
| 8 | H | −OC2H5 | −OC2H5 | O | 3.16 | 8.50 | |
| 9 | H | −OC2H5 | −H | O | 2.00 | 8.70 | |
| II | 10 | −Cl | − | - | - | 441.29 | 6.36 |
| 11 | −CH | − | - | - | 467.33 | 6.33 | |
| 12 | −CN | − | - | - | 170.88 | 6.77 | |
| 13 | −Br | − | - | - | 301.13 | 6.52 | |
| 14 | −F | −F | - | - | 108.10 | 6.97 | |
| 15 | −Cl | −CH3 | - | - | 100.44 | 7.00 | |
| 16 | −CH3 | −CH3 | - | - | 95.42 | 7.02 | |
| 17 | −CH3 | −Cl | - | - | 152.20 | 6.82 | |
| 18 | −CN | −Cl | - | - | 402.56 | 6.39 | |
| 19 | −Cl | −F | - | - | 86.16 | 7.06 | |
| III | 20 |  | −OCH3 | −OCH3 | - | 0.08 | 10.05 |
| 21 | 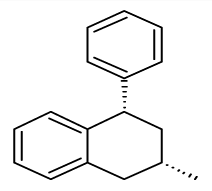 | −OCH3 | −OCH3 | - | 0.51 | 9.29 | |
| 22 |  | −OCH3 | −OCH3 | - | 3.23 | 8.49 | |
| IV | 23 |  | 2-Cl | 5-Cl | - | 95.68 | 7.02 |
| 24 |  | 2-Cl | 5-Cl | - | 264.40 | 6.58 | |
| 25 |  | 2-Cl | 5-Cl | - | 73.57 | 7.13 | |
| 26 |  | 2-OCH3 | - | - | 23.4 | 7.63 | |
| 27 |  | 2-OCH3 | - | - | 43.54 | 7.36 | |
| 28 |  | 2-Cl | 5-Cl | - | 21.77 | 7.66 | |
| 29 |  | 2-OCH3 | - | - | 5.88 | 8.23 | |
| 30 |  | 2-OCH3 | - | - | 7.94 | 8.10 | |
| 31 |  | 2-OCH3 | - | - | 28.84 | 7.54 | |
| V | 32 | - | - | - | - | 629.04 | 6.20 |
| Series | Compd. No. | R1 | R2 | R3 | X | Ki(nM) | pKi |
|---|---|---|---|---|---|---|---|
| I | 33 | 2-Cl | −OCH3 | −OCH3 | O | 1.41 | 8.85 |
| II | 34 | −Br | −Br | - | - | 91.25 | 7.04 |
| 35 | −Cl | −I | - | - | 383.60 | 6.42 | |
| III | 36 |  | −OCH3 | - | - | 27.54 | 7.57 |
| 37 |  | −OCH3 | −OCH3 | - | 2.34 | 8.63 | |
| 38 |  | −OCH3 | −OCH3 | - | 0.40 | 9.40 | |
| 39 |  | −OCH3 | −OCH3 | - | 72.44 | 7.14 | |
| IV | 40 |  | −Cl | −Cl | - | 2.28 | 8.64 |
| 41 |  | −OCH3 | - | - | 235.52 | 6.63 | |
| 42 |  | −Cl | -Cl | - | 47.86 | 7.32 | |
| 43 | 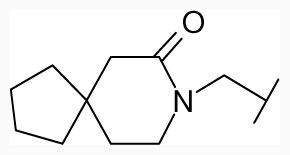 | −Cl | -Cl | - | 33.93 | 7.47 | |
| V | 44 |  | 31.62 | 7.50 | |||
| Parameters a | CoMFA | CoMSIA b | ||||
|---|---|---|---|---|---|---|
| A | B | C | D | E | ||
| Optimal PLS component | 4 | 3 | 3 | 3 | 2 | 3 |
| q2 | 0.840 | 0.874 | 0.866 | 0.842 | 0.856 | 0.840 |
| Scv | 0.476 | 0.407 | 0.419 | 0.456 | 0.427 | 0.459 |
| r2 | 0.988 | 0.980 | 0.982 | 0.977 | 0.961 | 0.975 |
| SEE | 0.128 | 0.160 | 0.154 | 0.174 | 0.222 | 0.180 |
| F | 555.64 | 469.24 | 510.31 | 394.83 | 357.74 | 370.67 |
| r2cv | 0.837 | 0.864 | ||||
| Fractions | ||||||
| Steric | 0.460 | 0.104 | 0.203 | 0.153 | 0.133 | |
| Electrostatic | 0.540 | 0.217 | 0.386 | 0.297 | 0.264 | 0.347 |
| Hydrophobic | 0.212 | 0.410 | 0.336 | 0.399 | ||
| Donnor | 0.183 | 0.360 | ||||
| Acceptor | 0.285 | 0.214 | 0.243 | 0.254 | ||
| r2pred | 0.694 | 0.646 | 0.581 | 0.576 | 0.663 | 0.671 |
| Compd. No. | pKi (exp.) | pKi (pred.) | ΔpKi a | ||
|---|---|---|---|---|---|
| CoMFA | CoMSIA | CoMFA | CoMSIA | ||
| Training set | |||||
| 1 | 9.60 | 9.620 | 9.561 | −0.0197 | 0.0388 |
| 2 | 8.80 | 8.743 | 8.896 | 0.0572 | −0.0957 |
| 3 | 9.40 | 9.410 | 9.273 | −0.0103 | 0.1265 |
| 4 | 8.70 | 8.593 | 8.680 | 0.1072 | 0.0201 |
| 5 | 8.70 | 8.659 | 8.687 | 0.0412 | 0.0127 |
| 6 | 8.70 | 8.649 | 8.777 | 0.0505 | −0.0771 |
| 7 | 8.50 | 8.510 | 8.611 | −0.0103 | −0.1113 |
| 8 | 8.50 | 8.440 | 8.676 | 0.0602 | −0.1755 |
| 9 | 8.70 | 8.689 | 8.843 | 0.0108 | −0.1432 |
| 10 | 6.36 | 6.505 | 6.694 | −0.1453 | −0.3342 |
| 11 | 6.33 | 6.385 | 6.474 | −0.0548 | −0.1435 |
| 12 | 6.77 | 6.652 | 6.716 | 0.1176 | 0.0540 |
| 13 | 6.52 | 6.559 | 6.605 | −0.0385 | −0.0847 |
| 14 | 6.97 | 6.923 | 6.901 | 0.0472 | 0.0688 |
| 15 | 7.00 | 7.113 | 7.035 | −0.1129 | −0.0352 |
| 16 | 7.02 | 6.880 | 6.690 | 0.1404 | 0.3303 |
| 17 | 6.82 | 6.793 | 6.656 | 0.0275 | 0.1644 |
| 18 | 6.39 | 6.475 | 6.641 | −0.0852 | −0.2509 |
| 19 | 7.06 | 7.011 | 7.024 | 0.0494 | 0.0363 |
| 20 | 10.05 | 10.126 | 9.768 | −0.0759 | 0.2821 |
| 21 | 9.29 | 9.294 | 9.181 | −0.0042 | 0.1095 |
| 22 | 8.49 | 8.601 | 8.751 | −0.1107 | −0.2609 |
| 23 | 7.02 | 6.871 | 6.661 | 0.1492 | 0.3588 |
| 24 | 6.58 | 6.590 | 6.691 | −0.0096 | −0.1111 |
| 25 | 7.13 | 7.177 | 7.202 | −0.0472 | −0.072 |
| 26 | 7.63 | 7.653 | 7.404 | −0.0229 | 0.2259 |
| 27 | 7.36 | 7.365 | 7.290 | −0.0055 | 0.0698 |
| 28 | 7.66 | 7.697 | 7.423 | −0.0371 | 0.2370 |
| 29 | 8.23 | 8.275 | 8.179 | −0.0451 | 0.0512 |
| 30 | 8.10 | 8.070 | 8.265 | 0.0305 | −0.1652 |
| 31 | 7.54 | 7.576 | 7.631 | −0.0364 | −0.0906 |
| 32 | 6.20 | 6.217 | 6.235 | −0.0173 | −0.0351 |
| Test set | |||||
| 33 | 8.85 | 8.587 | 9.010 | 0.2629 | −0.1603 |
| 34 | 7.04 | 6.766 | 6.613 | 0.2737 | 0.4265 |
| 35 | 6.42 | 6.669 | 6.653 | −0.2490 | −0.2334 |
| 36 | 7.57 | 8.514 | 8.370 | −0.9437 | −0.800 |
| 37 | 8.63 | 8.174 | 8.723 | 0.4563 | −0.0931 |
| 38 | 9.40 | 9.804 | 9.128 | −0.4037 | 0.2719 |
| 39 | 7.14 | 7.916 | 7.652 | −0.7758 | −0.5117 |
| 40 | 8.64 | 7.996 | 7.506 | 0.6440 | 1.1341 |
| 41 | 6.63 | 6.584 | 6.848 | 0.0461 | −0.2182 |
| 42 | 7.32 | 7.794 | 6.801 | −0.4744 | 0.5191 |
| 43 | 7.47 | 6.974 | 7.079 | 0.4955 | 0.3908 |
| 44 | 7.50 | 7.341 | 7.852 | 0.1589 | −0.3518 |
| r2predb | 0.694 | 0.671 | |||
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhao, X.; Chen, M.; Huang, B.; Ji, H.; Yuan, M. Comparative Molecular Field Analysis (CoMFA) and Comparative Molecular Similarity Indices Analysis (CoMSIA) Studies on α1A-Adrenergic Receptor Antagonists Based on Pharmacophore Molecular Alignment. Int. J. Mol. Sci. 2011, 12, 7022-7037. https://doi.org/10.3390/ijms12107022
Zhao X, Chen M, Huang B, Ji H, Yuan M. Comparative Molecular Field Analysis (CoMFA) and Comparative Molecular Similarity Indices Analysis (CoMSIA) Studies on α1A-Adrenergic Receptor Antagonists Based on Pharmacophore Molecular Alignment. International Journal of Molecular Sciences. 2011; 12(10):7022-7037. https://doi.org/10.3390/ijms12107022
Chicago/Turabian StyleZhao, Xin, Minsheng Chen, Biyun Huang, Hong Ji, and Mu Yuan. 2011. "Comparative Molecular Field Analysis (CoMFA) and Comparative Molecular Similarity Indices Analysis (CoMSIA) Studies on α1A-Adrenergic Receptor Antagonists Based on Pharmacophore Molecular Alignment" International Journal of Molecular Sciences 12, no. 10: 7022-7037. https://doi.org/10.3390/ijms12107022
APA StyleZhao, X., Chen, M., Huang, B., Ji, H., & Yuan, M. (2011). Comparative Molecular Field Analysis (CoMFA) and Comparative Molecular Similarity Indices Analysis (CoMSIA) Studies on α1A-Adrenergic Receptor Antagonists Based on Pharmacophore Molecular Alignment. International Journal of Molecular Sciences, 12(10), 7022-7037. https://doi.org/10.3390/ijms12107022




