Interaction of Classical Platinum Agents with the Monomeric and Dimeric Atox1 Proteins: A Molecular Dynamics Simulation Study
Abstract
:1. Introduction
2. Results
2.1. The Characteristics of Platinum Agents Combining with a Monomeric Atox1 Protein
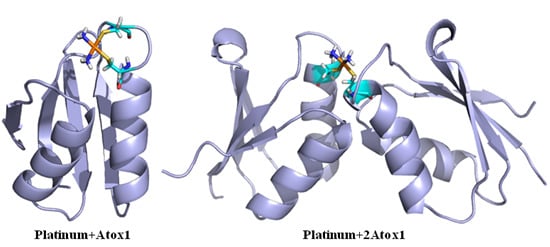
2.1.1. The Structure Characteristics of Platinum + Atox1 Complexes
2.1.2. Free Energy and Interaction Analysis of Platinum + Atox1 Complexes
2.1.3. Dynamical Fluctuation Analysis for Platinum + Atox1 Complexes
2.2. The Characteristics of Platinum Agents Combining with a Dimeric Atox1 Protein
3. Discussion
3.1. The Structure Analysis of Platinum Agents Combining with the Monomeric and Dimeric Atox1 Protein
3.2. Comparison on the Structural Characteristics of the Oxaliplatin + Atox1 and Oxaliplatin + 2Atox1 Models
4. Models and Methods
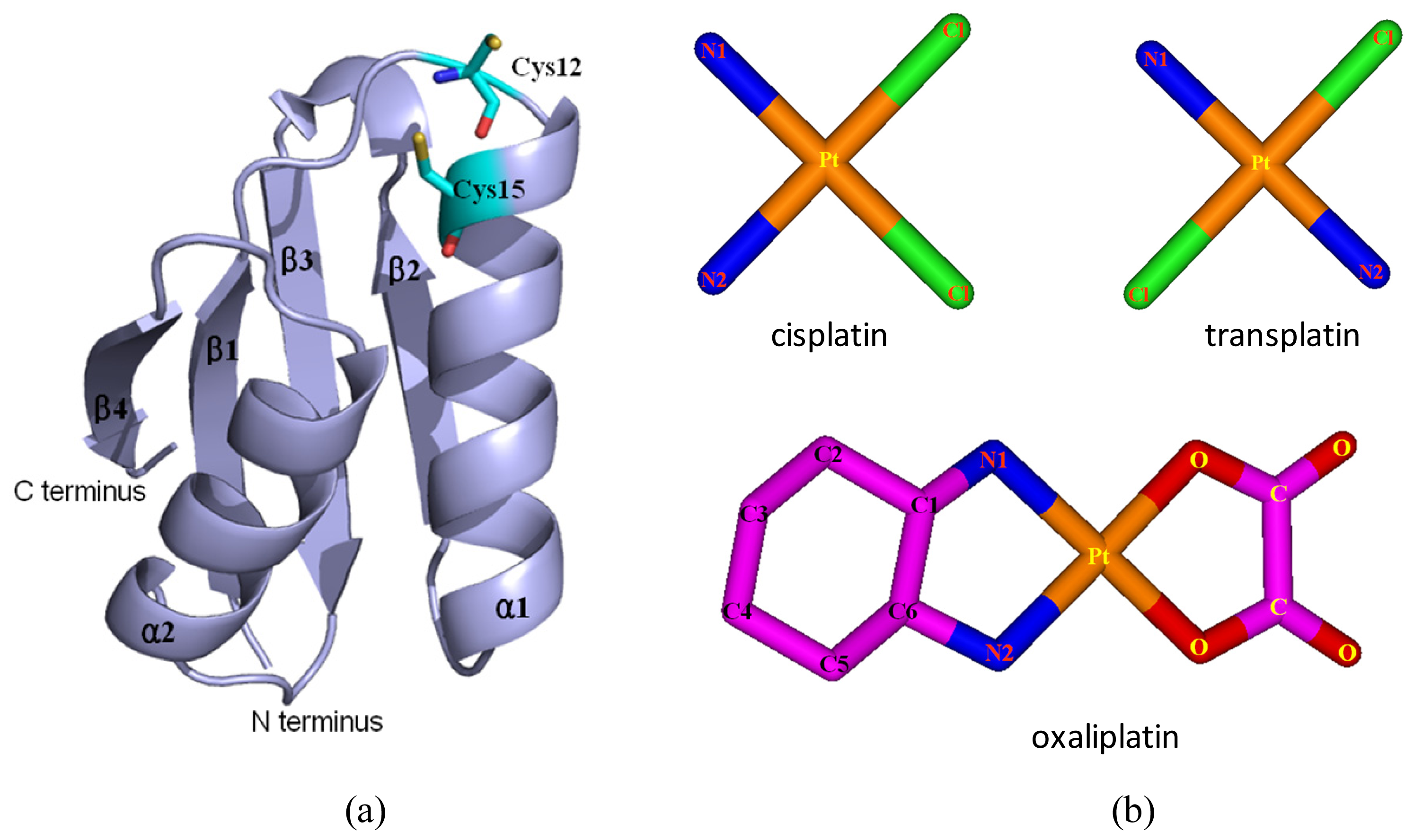
4.1. Initial Structures
4.2. Force Field Parameter Preparation
4.3. Molecular Dynamics Simulation Protocols
4.4. Free-Energy Analyses
4.5. Fluctuation Analyses
4.6. Calculation of Angle between Two Helices
4.7. DynDom and Surface Analyses
5. Conclusions
Supplementary Information
ijms-15-00075-s001.pdfAcknowledgments
Conflicts of Interest
References
- Jamieson, E.R.; Lippard, S.J. Structure, recognition, and processing of cisplatin-DNA adducts. Chem. Rev 1999, 99, 2467–2498. [Google Scholar]
- Brabec, V. DNA Modifications by Antitumor Platinum and Ruthenium Compounds: Their Recognition and Repair. In Progress in Nucleic Acid Research and Molecular Biology; Academic Press: Brno, Czech Republic, 2002; pp. 1–68. [Google Scholar]
- Paquet, F.; Boudvillain, M.; Lancelot, G.; Leng, M. NMR solution structure of a DNA dodecamer containing a transplatin interstrand GN7-CN3 cross-link. Nucleic Acids Res 1999, 27, 4261–4268. [Google Scholar]
- Heringova, P.; Woods, J.; Mackay, F.S.; Kasparkova, J.; Sadler, P.J.; Brabec, V. Transplatin is cytotoxic when photoactivated: Enhanced formation of DNA cross-links. J. Med. Chem 2006, 49, 7792–7798. [Google Scholar]
- Almeida, G.M.; Duarte, T.L.; Steward, W.P.; Jones, G.D.D. Detection of oxaliplatin-induced DNA crosslinks in vitro and in cancer patients using the alkaline comet assay. DNA Repair 2006, 5, 219–225. [Google Scholar]
- Woynarowski, J.M.; Faivre, S.; Herzig, M.C.S.; Arnett, B.; Chapman, W.G.; Trevino, A.V.; Raymond, E.; Chaney, S.G.; Vaisman, A.; Varchenko, M.; et al. Oxaliplatin-induced damage of cellular DNA. Mol. Pharmacol 2000, 58, 920–927. [Google Scholar]
- Hannon, M.J. Metal-based anticancer drugs: From a past anchored in platinum chemistry to a post-genomic future of diverse chemistry and biology. Pure Appl. Chem 2007, 79, 2243–2261. [Google Scholar]
- Kostova, I. Platinum complexes as anticancer agents. Recent Pat. Anticancer Drug Discov 2006, 1, 1–22. [Google Scholar]
- Reedijk, J. Platinum anticancer coordination compounds: Study of DNA binding inspires new drug design. Eur. J. Inorg. Chem 2009, 2009, 1303–1312. [Google Scholar]
- Sherman, S.E.; Gibson, D.; Wang, A.H.; Lippard, S.J. X-ray structure of the major adduct of the anticancer drug cisplatin with DNA: Cis-[Pt(NH3)2(d(pGpG))]. Science 1985, 230, 412–417. [Google Scholar]
- Zhao, T.; King, F.L. Direct determination of the primary binding site of cisplatin on cytochrome C by mass spectrometry. J. Am. Soc. Mass Spectr 2009, 20, 1141–1147. [Google Scholar]
- Sze, C.; Khairallah, G.; Xiao, Z.; Donnelly, P.; O’Hair, R.J.; Wedd, A. Interaction of cisplatin and analogues with a met-rich protein site. J. Biol. Inorg. Chem 2009, 14, 163–165. [Google Scholar]
- Howell, S.B.; Safaei, R.; Larson, C.A.; Sailor, M.J. Copper transporters and the cellular pharmacology of the platinum-containing cancer drugs. Mol. Pharmacol 2010, 77, 887–894. [Google Scholar]
- Kuo, M.; Chen, H.W.; Song, I.-S.; Savaraj, N.; Ishikawa, T. The roles of copper transporters in cisplatin resistance. Cancer Metastasis Rev 2007, 26, 71–83. [Google Scholar]
- Kalayda, G.; Wagner, C.; BuSz, I.; Reedijk, J.; Jaehde, U. Altered localisation of the copper efflux transporters ATP7A and ATP7B associated with cisplatin resistance in human ovarian carcinoma cells. BMC Cancer 2008, 8, 175. [Google Scholar]
- Lin, X.; Okuda, T.; Holzer, A.; Howell, S.B. The copper transporter CTR1 regulates cisplatin uptake in saccharomyces cerevisiae. Mol. Pharmacol 2002, 62, 1154–1159. [Google Scholar]
- Ishida, S.; Lee, J.; Thiele, D.J.; Herskowitz, I. Uptake of the anticancer drug cisplatin mediated by the copper transporter Ctr1 in yeast and mammals. Proc. Natl. Acad. Sci. USA 2002, 99, 14298–14302. [Google Scholar]
- Holzer, A.K.; Samimi, G.; Katano, K.; Naerdemann, W.; Lin, X.; Safaei, R.; Howell, S.B. The copper influx transporter human copper transport protein 1 regulates the uptake of cisplatin in human ovarian carcinoma cells. Mol. Pharmacol 2004, 66, 817–823. [Google Scholar]
- Komatsu, M.; Sumizawa, T.; Mutoh, M.; Chen, Z.-S.; Terada, K.; Furukawa, T.; Yang, X.-L.; Gao, H.; Miura, N.; Sugiyama, T.; et al. Copper-transporting P-type adenosine triphosphatase (ATP7B) is associated with cisplatin resistance. Cancer Res 2000, 60, 1312–1316. [Google Scholar]
- Nakayama, K.; Kanzaki, A.; Ogawa, K.; Miyazaki, K.; Neamati, N.; Takebayashi, Y. Copper-transporting P-type adenosine triphosphatase (ATP7B) as a cisplatin based chemoresistance marker in ovarian carcinoma: Comparative analysis with expression of MDR1, MRP1, MRP2, LRP and BCRP. Int. J. Cancer 2002, 101, 488–495. [Google Scholar]
- Anastassopoulou, I.; Banci, L.; Bertini, I.; Cantini, F.; Katsari, E.; Rosato, A. Solution structure of the apo and copper(I)-loaded human metallochaperone HAH1. Biochemistry 2004, 43, 13046–13053. [Google Scholar]
- Safaei, R.; Maktabi, M.H.; Blair, B.G.; Larson, C.A.; Howell, S.B. Effects of the loss of Atox1 on the cellular pharmacology of cisplatin. J. Inorg. Biochem 2009, 103, 333–341. [Google Scholar]
- Boal, A.K.; Rosenzweig, A.C. Crystal structures of cisplatin bound to a human copper chaperone. J. Am. Chem. Soc 2009, 131, 14196–14197. [Google Scholar]
- Arnesano, F.; Banci, L.; Bertini, I.; Felli, I.C.; Losacco, M.; Natile, G. Probing the interaction of cisplatin with the human copper chaperone Atox1 by solution and in-cell NMR spectroscopy. J. Am. Chem. Soc 2011, 133, 18361–18369. [Google Scholar]
- Palm, M.E.; Weise, C.F.; Lundin, C.; Wingsle, G.; Nygren, Y.; Björn, E.; Naredi, P.; Wolf-Watz, M.; Wittung-Stafshede, P. Cisplatin binds human copper chaperone Atox1 and promotes unfolding in vitro. Proc. Natl. Acad. Sci. USA 2011, 108, 6951–6956. [Google Scholar]
- Hua, H.; Günther, V.; Georgiev, O.; Schaffner, W. Distorted copper homeostasis with decreased sensitivity to cisplatin upon chaperone Atox1 deletion in Drosophila. BioMetals 2011, 24, 445–453. [Google Scholar]
- Hemmingsson, O.; Zhang, Y.; Still, M.; Naredi, P. ASNA1, an ATPase targeting tail-anchored proteins, regulates melanoma cell growth and sensitivity to cisplatin and arsenite. Cancer Chemother. Pharmcol 2009, 63, 491–499. [Google Scholar]
- Hemmingsson, O.; Nöjd, M.; Kao, G.; Naredi, P. Increased sensitivity to platinating agents and arsenite in human ovarian cancer by downregulation of ASNA1. Oncol. Rep 2009, 22, 869–875. [Google Scholar]
- Palm-Espling, M.E.; Wittung-Stafshede, P. Reaction of platinum anticancer drugs and drug derivatives with a copper transporting protein, Atox1. Biochem. Pharmacol 2012, 83, 874–881. [Google Scholar]
- Mizumura, Y.; Matsumura, Y.; Hamaguchi, T.; Nishiyama, N.; Kataoka, K.; Kawaguchi, T.; Hrushesky, W.J.M.; Moriyasu, F.; Kakizoe, T. Cisplatin-incorporated polymeric micelles eliminate nephrotoxicity, while maintaining antitumor activity. Cancer Sci 2001, 92, 328–336. [Google Scholar]
- Casini, A.; Mastrobuoni, G.; Temperini, C.; Gabbiani, C.; Francese, S.; Moneti, G.; Supuran, C.T.; Scozzafava, A.; Messori, L. ESI mass spectrometry and X-ray diffraction studies of adducts between anticancer platinum drugs and hen egg white lysozyme. Chem. Commun. (Camb.) 2007, 156–158. [Google Scholar]
- Jerremalm, E.; Wallin, I.; Ehrsson, H. New insights into the biotransformation and pharmacokinetics of oxaliplatin. J. Pharm. Sci 2009, 98, 3879–3885. [Google Scholar]
- Brocklehurst, S.M.; Perham, R.N. Prediction of the three-dimensional structures of the biotinylated domain from yeast pyruvate carboxylase and of the lipoylated H-protein from the pea leaf glycine cleavage system: A new automated method for the prediction of protein tertiary structure. Protein Sci 1993, 2, 626–639. [Google Scholar]
- Chuprina, V.P.; Rullmann, J.A.C.; Lamerichs, R.M.J.N.; van Boom, J.H.; Boelens, R.; Kaptein, R. Structure of the complex of lac repressor headpiece and an 11 base-pair half-operator determined by nuclear magnetic resonance spectroscopy and restrained molecular dynamics. J. Mol. Biol 1993, 234, 446–462. [Google Scholar]
- Gutmanas, A.; Billeter, M. Specific DNA recognition by the Antp homeodomain: MD simulations of specific and nonspecific complexes. Proteins 2004, 57, 772–782. [Google Scholar]
- Boal, A.K.; Rosenzweig, A.C. Structural biology of copper trafficking. Chem. Rev 2009, 109, 4760–4779. [Google Scholar]
- Ralle, M.; Lutsenko, S.; Blackburn, N.J. X-ray absorption spectroscopy of the copper chaperone HAH1 reveals a linear two-coordinate Cu(I) center capable of adduct formation with exogenous thiols and phosphines. J. Biol. Chem 2003, 278, 23163–23170. [Google Scholar]
- Laskowski, R.A.; Luscombe, N.M.; Swindells, M.B.; Thornton, J.M. Protein clefts in molecular recognition and function. Protein Sci 1996, 5, 2438–2452. [Google Scholar]
- Binkowski, T.A.; Naghibzadeh, S.; Liang, J. CASTp: Computed atlas of surface topography of proteins. Nucleic Acids Res 2003, 31, 3352–3355. [Google Scholar]
- Liang, J.; Woodward, C.; Edelsbrunner, H. Anatomy of protein pockets and cavities: Measurement of binding site geometry and implications for ligand design. Protein Sci 1998, 7, 1884–1897. [Google Scholar]
- Wu, Y.; Pradhan, P.; Havener, J.; Boysen, G.; Swenberg, J.A.; Campbell, S.L.; Chaney, S.G. NMR solution structure of an oxaliplatin 1,2-d(GG) intrastrand cross-link in a DNA dodecamer duplex. J. Mol. Biol 2004, 341, 1251–1269. [Google Scholar]
- Carloni, P.; Andreoni, W. Platinum-modified nucleobase pairs in the solid state: A theoretical study. J. Phys. Chem 1996, 100, 17797–17800. [Google Scholar]
- Arnesano, F.; Losacco, M.; Natile, G. An updated view of cisplatin transport. Eur. J. Inorg. Chem 2013, 2013, 2701–2711. [Google Scholar]
- Guex, N.; Peitsch, M.C. SWISS-MODEL and the Swiss-PdbView: An enviroment for comparative protein modeling. Electrophoresis 1997, 18, 2714–2723. [Google Scholar]
- Wang, G.; Li, C.; Wang, Y.; Chen, G. Cooperative assembly of co-Smad4 MH1 with R-Smsd1/3 MH1 on DNA: A molecular dynamics simulation study. PLoS One 2013, 8, e53841. [Google Scholar]
- Morris, G.M.; Goodsell, D.S.; Halliday, R.S.; Huey, R.; Hart, W.E.; Belew, R.K.; Olson, A.J. Automated docking using a lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem 1998, 19, 1639–1662. [Google Scholar]
- Formaneck, M.S.; Ma, L.; Cui, Q. Effects of temperature and salt concentration on the structural stability of human lymphotactin: Insights from molecular simulations. J. Am. Chem. Soc 2006, 128, 9506–9517. [Google Scholar]
- Case, D.A.; Darden, T.A.; Cheatham, T.E., Iii; Simmerling, C.L.; Wang, J.; Duke, R.E.; Luo, R.; Merz, K.M.; Pearlman, D.A.; Crowley, M. Amber 9; University of California: San Francisco, CA, USA, 2006. [Google Scholar]
- Ruiz, J.; Rodríguez, V.; Cutillas, N.; Espinosa, A.; Hannon, M.J. Novel C,N-chelate platinum(II) antitumor complexes bearing a lipophilic ethisterone pendant. J. Inorg. Biochem 2011, 105, 525–531. [Google Scholar]
- Mantri, Y.; Lippard, S.J.; Baik, M.-H. Bifunctional binding of cisplatin to DNA: Why does cisplatin form 1,2-intrastrand cross-links with AG but not with GA? J. Am. Chem. Soc 2007, 129, 5023–5030. [Google Scholar]
- Kobayashi, A.; Ohbayashi, K.; Aoki, R.; Chang, H.-C.; Kato, M. Synthesis, structure and photophysical properties of a flavin-based platinum(II) complex. Dalton Trans 2011, 40, 3484–3489. [Google Scholar]
- Lavery, R.; Sklenar, H. The definition of generalized helicoidal parameters and of axis curvature for irregular nucleic acids. J. Biomol. Struct. Dyn 1988, 6, 63–91. [Google Scholar]
- Frisch, M.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G.A. Gaussian 09, Revision A. 02; Gaussian Inc: Wallingford, CT, USA, 2009; Volume 270, p. 271. [Google Scholar]
- Williams, K.M.; Rowan, C.; Mitchell, J. Effect of amine ligand bulk on the interaction of methionine with platinum(II) diamine complexes. Inorg. Chem 2004, 43, 1190–1196. [Google Scholar]
- Lee, M.C.; Duan, Y. Distinguish protein decoys by using a scoring function based on a new amber force field, short molecular dynamics simulations, and the generalized born solvent model. Proteins 2004, 55, 620–634. [Google Scholar]
- Duan, Y.; Wu, C.; Chowdhury, S.; Lee, M.C.; Xiong, G.; Zhang, W.; Yang, R.; Cieplak, P.; Luo, R.; Lee, T.; et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem 2003, 24, 1999–2012. [Google Scholar]
- Yao, S.; Plastaras, J.P.; Marzilli, L.G. A molecular mechanics amber-type force field for modeling platinum complexes of guanine derivatives. Inorg. Chem 1994, 33, 6061–6077. [Google Scholar]
- Cheatham, T.E.; Srinivasan, J.; Case, D.A.; Kollman, P.A. Molecular dynamics and continuum solvent studies of the stability of polyG-polyC and polyA-polyT DNA duplexes in solution. J. Biomol. Struct. Dyn 1998, 16, 265–280. [Google Scholar]
- Kollman, P.A.; Massova, I.; Reyes, C.; Kuhn, B.; Huo, S.; Chong, L.; Lee, M.; Lee, T.; Duan, Y.; Wang, W.; et al. Calculating structures and free energies of complex molecules: Combining molecular mechanics and continuum models. Acc. Chem. Res 2000, 33, 889–897. [Google Scholar]
- Jayaram, B.; Sprous, D.; Young, M.A.; Beveridge, D.L. Free energy analysis of the conformational preferences of A and B forms of DNA in solution. J. Am. Chem. Soc 1998, 120, 10629–10633. [Google Scholar]
- Srinivasan, J.; Cheatham, T.E.; Cieplak, P.; Kollman, P.A.; Case, D.A. Continuum solvent studies of the stability of DNA, RNA, and phosphoramidate–DNA helices. J. Am. Chem. Soc 1998, 120, 9401–9409. [Google Scholar]
- McQuarrie, D.A. Statistical Mechanics; Harper and Row: New York, NY, USA, 1976. [Google Scholar]
- Kottalam, J.; Case, D.A. Langevin modes of macromolecules: Applications to crambin and DNA hexamers. Biopolymers 1990, 29, 1409–1421. [Google Scholar]
- Frank, J. Introduction to Computational Chemistry; Wiley: New York, NY, USA, 1999. [Google Scholar]
- Sadiq, S.K.; de Fabritiis, G. Explicit solvent dynamics and energetics of HIV-1 protease flap opening and closing. Proteins 2010, 78, 2873–2885. [Google Scholar]
- Yap, K.L.; Ames, J.B.; Swindells, M.B.; Ikura, M. Diversity of conformational states and changes within the EF-hand protein superfamily. Proteins 1999, 37, 499–507. [Google Scholar]
- Yap, K.; Ames, J.; Swindells, M.; Ikura, M. Vector Geometry Mapping. In Calcium-Binding Protein Protocols. Volume 2: Methods and Techniques; Vogel, H., Ed.; Springer: New York, NY, USA, 2002; Volume 173, pp. 317–324. [Google Scholar]
- Hayward, S.; Berendsen, H.J.C. Systematic analysis of domain motions in proteins from conformational change: New results on citrate synthase and T4 lysozyme. Proteins 1998, 30, 144–154. [Google Scholar]
- Dundas, J.; Ouyang, Z.; Tseng, J.; Binkowski, A.; Turpaz, Y.; Liang, J. CASTp: Computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res 2006, 34, 116–118. [Google Scholar]
- Dolinsky, T.J.; Czodrowski, P.; Li, H.; Nielsen, J.E.; Jensen, J.H.; Klebe, G.; Baker, N.A. PDB2PQR: Expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res 2007, 35, 522–525. [Google Scholar]
- Dolinsky, T.J.; Nielsen, J.E.; McCammon, J.A.; Baker, N.A. PDB2PQR: An automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res 2004, 32, 665–667. [Google Scholar]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem 2004, 25, 1157–1174. [Google Scholar]
- Miyamoto, S.; Kollman, P.A. Settle: An analytical version of the shake and rattle algorithm for rigid water models. J. Comput. Chem 1992, 13, 952–962. [Google Scholar]
- Connolly, M.L. Analytical molecular surface calculation. J. Appl. Crystallogr 1983, 16, 548–558. [Google Scholar]









| Component | CisPt + Atox1 | TransPt + Atox1 | OxaliPt + Atox1 |
|---|---|---|---|
| Receptor | Atox1 | Atox1 | Atox1 |
| Ligand | Cisplatin ligand | Transplatin ligand | Oxaliplatin ligand |
| ΔEele | −46.47 | 11.24 | −35.54 |
| ΔEvdw | −0.64 | −2.47 | −9.09 |
| ΔEint | 0.00 | 0.00 | 0.00 |
| ΔGnp/solv | −8.86 | −11.10 | −14.93 |
| ΔGpb/solv | 45.06 | −0.39 | 41.56 |
| ΔGnp | −9.50 | −13.57 | −24.02 |
| ΔGpb | −1.41 | 10.85 | 6.02 |
| ΔHbinding | −10.91 | −2.73 | −18.00 |
| TΔS | −16.19 | −15.14 | −13.87 |
| ΔGbinding | 5.28 | 12.41 | −4.13 |
| Hydrogen bond | OxaliPt + Atox1 |
|---|---|
| (Cys15)CB H–N1 | 99.0 |
| (Cys15)CA H–N2 | 77.6 |
| N1 H–CA(Cys15) | 30.6 |
| Hydrophobic contact | |
| (Val8)CG1 C4(oxaliplatin) | 65.4 |
| (Val8)CG1 C5(oxaliplatin) | 88.7 |
| (Val8)CG1 C3(oxaliplatin) | 47.2 |
| (Asp9)CB C5(oxaliplatin) | 90.3 |
| (Cys15)CA/CB C1(oxaliplatin) | 197.2 |
| (Ala18)CB C2(oxaliplatin) | 76.3 |
| (Thr58)CB C4(oxaliplatin) | 50.9 |
| (Thr58)CB/CG2 C3(oxaliplatin) | 126.5 |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, X.; Li, C.; Wang, Y.; Chen, G. Interaction of Classical Platinum Agents with the Monomeric and Dimeric Atox1 Proteins: A Molecular Dynamics Simulation Study. Int. J. Mol. Sci. 2014, 15, 75-99. https://doi.org/10.3390/ijms15010075
Wang X, Li C, Wang Y, Chen G. Interaction of Classical Platinum Agents with the Monomeric and Dimeric Atox1 Proteins: A Molecular Dynamics Simulation Study. International Journal of Molecular Sciences. 2014; 15(1):75-99. https://doi.org/10.3390/ijms15010075
Chicago/Turabian StyleWang, Xiaolei, Chaoqun Li, Yan Wang, and Guangju Chen. 2014. "Interaction of Classical Platinum Agents with the Monomeric and Dimeric Atox1 Proteins: A Molecular Dynamics Simulation Study" International Journal of Molecular Sciences 15, no. 1: 75-99. https://doi.org/10.3390/ijms15010075