1. Introduction
Bone marrow derived mesenchymal stem cells (BMSCs) are mesodermal multipotent stem cells with self-renewal capacities and possess the ability to differentiate beyond their lineage into endoderm and even ectoderm cells
in vitro [
1,
2]. Our previous study showed that BMSCs could be differentiated into neural progenitor-like cells (NPCs) under a specifically induced microenvironment [
3]. We found a combination of epidermal growth factor (EGF), basic fibroblast growth factor (bFGF), and insulin-like growth factor 1 (IGF-1) could significantly improve the quality of the derived NPCs, as the addition of IGF-1 enhances cell proliferation and survivability compared to the published protocol used only EGF and bFGF. An identical combination of growth factors was also reported to provide an optimal niche for embryonic striatal stem cell maintenance [
4]. However, the molecular mechanism of IGF-1 addition on BMSC-derived NPCs maintenance is still unclear. We believed that alteration to gene expression by microRNAs play important role in the enhancement of cellular activities.
MicroRNAs are short noncoding RNA with 18 to 22 nucleotides that regulates gene expression at posttranscriptional levels by base pairing with targeted messenger RNA (mRNA) [
5]. MicroRNAs bind on the 3'-untranslated region of mRNA by perfect base pairing, leading to mRNA cleavage. In contrast, binding with imperfect base pairing may cause translational repression or deadenylation [
5]. A single microRNA may regulate hundreds of target mRNAs and single target mRNA may be regulated by several microRNAs. Therefore, microRNA-mRNA interaction forms a complex gene regulatory network. MicroRNAs also regulate genes at the transcriptional level by modulating DNA methylation and histone modification. For instance, miR-10 is required for hypermethylation in gastric cancer, and the mechanism was predicted by targeting the
HOXA1 gene [
6]. Similarly, miR-874, a putative tumor suppressor in human cancers, can target histone deacetylase 1 in head and neck squamous cell carcinoma and contributes to cell proliferation [
7]. Taken together, microRNAs can be considered important players in the control of epigenetics modification.
MicroRNAs have also emerged as powerful regulators of diverse biological processes, including cell differentiation [
8,
9], proliferation [
10] and apoptosis [
11,
12]. MicroRNAs are also involved as key modulators of neuronal development, neuroplasticity, and disease pathogenesis, such as neurodegenerative disease and traumatic brain injuries [
11,
13]. The substantial value of microRNAs has been reported for medical diagnostic and prognostic determination, which eventually will lead to novel therapeutic intervention [
14]. In addition, microRNAs have critical roles in stem cell differentiation and the derivation of induced pluripotent stem cells [
15].
To elucidate the functions of microRNAs in stem cell differentiation, global profiling, such as microRNA microarray, microRNA sequencing, real-time PCR, and next generation sequencing of microRNA [
16], may be carried out to find differentially and uniquely expressed microRNAs involved in the differentiation of BMSCs into neural lineages. Several microRNA expression studies have been carried out, such as the elucidation of genes involved between the mouse frontal cortex and hippocampus [
17] and microRNA expression pattern changes in spinal-cord injury [
18]. However, no study to date has characterized microRNA expression patterns in BMSC-derived NPCs under the influence of IGF-1. This information is critical since changes in cellular physiology, such as apoptosis and growth rate, are closely related to their microRNA-mRNA interactome within cells.
Microarray of microRNA can reveal differential expression of several microRNAs and microRNA family. We hypothesized that microRNA family may act as a whole in regulating specific cellular functions and pathways during the differentiation of BMSCs into NPCs. Therefore, the aim of this study was to identify the biological functions and pathways that might be involved due to the expression of key microRNAs during the differentiation of BMSCs into NPCs under the effect of IGF-1.
3. Discussion
The present profiling work was the first to decipher the microRNA expression patterns in adult rat BMSC-derived NPCs. Microarray-based microRNA profiling is a common method to discover the differentially and uniquely expressed microRNAs. This procedure is the first step in the screening process of overexpressed microRNAs that in turn regulate gene expression.
Both bFGF and EGF have been shown to be crucial factors in differentiating BMSCs into neural lineage cells [
21]. IGF-1 is another polypeptide important for nervous system development [
22]. By stimulating the IGF-1 receptor, IGF-1 acts as an important mitogenic factor to promote neural cell proliferation and survival
in vitro and
in vivo [
23].
In our previous study, we confirmed that a combination of bFGF, EGF and IGF-1 could induce the neuronal differentiation (NPCs) of BMSCs and significantly (
p < 0.05) enhance BMSCs-derived NPCs viability in undifferentiated state [
3]. However, the molecular mechanisms such as microRNAs profiling, associated with this enhanced differentiation remain to be elucidated. Therefore, in this study, we described the microRNA signatures of NPCs (P1) specifically at Day 1, 3 and 5 after induction. These signatures showed the pattern of microRNA changes during differentiation. However, one of the main limitations in deciphering the mechanism of epigenetic modification and maintenance of NPCs is the complexity of biological pathways associated with regulating microRNAs. Therefore, it is difficult to reveal the specific biological pathways based on the analysis of a handful of predicted genes.
Among the 389 rat-specific microRNA probes on the GeneChip® microRNA 2.0 array, we discovered 46 microRNAs up-regulated or down-regulated due to the effects of growth factors. However, microRNAs with inconsistent expression throughout the treatment days from Groups A and B were excluded from analysis. Only microRNAs differentially expressed at least on two consecutive treatment days were taken into account.
There are 30 microRNAs (
Table 1 and
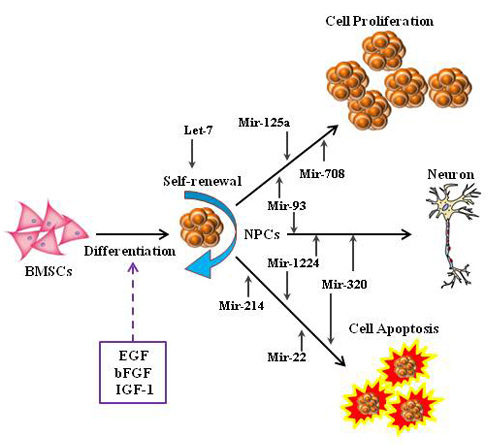
Table 2) consistently expressed in two patterns (up-pattern or down-pattern) from Day 1 to Day 5. Both up- and down-patterned microRNAs were observed in Group A; however, in the presence of IGF-1, the majority of microRNAs were down-regulated and only miR-496 was up-regulated. Since the down-regulation of microRNAs may cause the up-regulation of targeted genes, we hypothesized that the presence of IGF-1 triggers the expression of certain genes by down-regulating key microRNAs (miR-1224, miR-125a-3p, miR-214, miR-22, miR-320, miR-708, and miR-93), which in turn enhance NPCs proliferation and survivability.
In Groups A and B, let-7 family microRNAs were observed to be consistently down-regulated. These data suggest contribution of different members of one microRNA cluster or family act as a whole in regulating specific signaling pathways. MicroRNAs from miR-8 family and miR-34 family were reported to be involved in the regulation of ceramide signaling pathway in the frontal cortex and dopamine signaling pathway in the hippocampus respectively [
17]. In addition, Itesako
et al. concluded that down-regulation of miR-195/497 cluster contributed to bladder cancer progression by targeting
BIRC5 and
WNT7A genes [
24]. Therefore, our finding is supported by previous experiments regarding signaling pathway targeted by either microRNA families or clusters. Besides, Cimadamore
et al. reported that LIN28 binds to precursor let-7 microRNA and blocks the production of mature let-7i microRNAs, inhibiting neuronal differentiation by targeting
MASH1 and
NGN1 genes [
25]. This process suggests that the down-regulation of the let-7 microRNA family promotes cell proliferation during early neurogenesis.
miR-93, a microRNA frequently associated with TGF-β signaling in controlling cell cycle arrest [
26], cell proliferation, and differentiation [
27], was also down-regulated in both Groups A and B. Moreover, we discovered that miR-1224 and miR-125a-3p, which were initially up-regulated in Group A, became down-regulated at day 3 and 5 in Group B post-induced with IGF-1. Both miR-1224 and miR-125a-3p play important roles in maintaining cell proliferation and survivability. The down-regulation of miR-1224 positively regulates TNF-α gene expression [
28], which strongly influences NPC survival, proliferation, and neuronal differentiation [
29,
30].
The down-regulation of miR-125a-3p has been associated with a reduction in apoptosis by targeting p53 mRNA and an increase in cell proliferation and migration by up-regulating Fyn expression [
31,
32]. Stronger p53 protein expression has been associated with the induction of apoptosis and inhibition of cell proliferation [
33]. Similarly, the reduction of Fyn and its downstream proteins causes cell cycle arrest at G2/M stage, consequently, reducing cell viability and migration [
32]. These data support our findings that the down-regulation of mir-125-3p reduced apoptosis and increased cell proliferation possibly by a p53- and Fyn-regulated manner.
Furthermore, the introduction of IGF-1 triggers the down-regulation of several microRNAs (miR-214, miR-22, miR-320 and miR-708) with their inconsistent differential expression in Group A. The inhibition of miR-214 has been reported to decrease the level of apoptosis in HeLa cells [
34]. Yu
et al. reported that miR-22 is a pivotal candidate for gene therapy in cerebral ischemic injury due to its neuroprotective effect [
35]. The down-regulation of miR-708 was reported to be involved in the enhancement of cell proliferation [
36] and the expression of miR-708 has been reported to be associated with PI3K/Akt signaling pathways [
37]. In summary, a group of microRNAs with individual roles forms a complex functional network, and together they contribute to the enhancement of BMSC-derived NPCs.
Differences in functional involvement of down-regulated microRNAs in Groups A and B were measured using GO and pathway analyses. Biological relevant of each GO terms were determined by enrichment score. Top-ranked annotation groups indicated by higher enrichment score are indicated as more significant biologically [
38,
39]. Both gene groups in A and B were generally associated with cell growth and survival signaling functions (
Table 4 and
Figure 4). The GO term “regulation of programmed cell death” (GO: 0043067) was overrepresented in both groups, resulting from the activation of endogenous cellular processes that play important roles in cell epigenetic modification [
40]. The regulation of non-apoptotic programmed cell death is most likely due to the effects of combinatorial EGF and bFGF in which the synergistic treatment of both growth factors modify the BMSCs fate into a neural lineage.
Zhao
et al. reported that EGF and bFGF trigger the expression of miR-9, which targets the nuclear receptor TLX controlling NSC proliferation and fate determination [
41]. Chuang and Jones reported that small interfering RNAs closely related to microRNAs are involved in the histone modification and DNA methylation that regulate the mechanism of epigenetic modification [
42]. However, the gene list of Group B (EGF + bFGF + IGF-1) showed a higher important level with an enrichment score of 15.76 (34 genes involved) than did Group A (EGF + bFGF) with an enrichment score of 13.31 (29 genes involved) in GO term “negative regulation of apoptotic process” (GO: 0043066). These data indicate that the down-regulation of key microRNAs in Group B due to IGF-1 triggers some yet to be determined biological processes that reduce apoptotic process. Our GO analysis data is consistent with our apoptosis assay findings described earlier.
Furthermore, functional networks analysis of up-regulated genes associated with GO term “negative regulation of apoptosis process” by the algorithm GeneMANIA. It revealed several genes such as
Akt1 (v-Akt murine thymoma viral oncogene homolog 1),
Tp53 (tumor protein p53),
Pten (phosphatase and tensin homolog) and
Bcl2 (B-cell lymphoma 2). Protein kinase B or Akt, a key protein involved in the activation of PI3K-Akt pathway and is crucial in promoting cell survivability [
43], is inhibited by the key microRNAs (miR-22, miR-214, miR-125a-3p, miR-320 and let-7 family) that are down-regulated with the addition of IGF-1. Chen
et al. reported that down-regulation of miR-133b significantly overexpressed
Akt1 mRNA, which increased T24 bladder cancer cell proliferation and reduced cell apoptosis [
44]. Overexpression of
Akt1 gene also triggers the expression of
Pten, an antagonist of the PI3K pathway [
45]. It has been reported that up-regulation of miR-93 stimulated cell proliferation and inhibited apoptosis through Akt pathway by targeting
Pten and
Cdkn1a [
46]. Wang
et al. showed experimental evidence of miR-21 inhibition triggered the overexpression of
Pten and
Ptpn14, which was a result of suppressed intrahepatic cholangiocarcinoma cell proliferation and growth [
47].
Moreover, down-regulation of key microRNAs also triggers the expression of anti-apoptotic gene
Bcl-2 and tumor suppressor
p53. Down-regulation of miR-451 and miR-885-5p in neuron-like cells has been shown to increase
Bcl-2 expression and lead to reduction of apoptosis activity [
48]. Similarly, Lin
et al. showed that inhibition of miR-34a restored the down-regulated expression of anti-apoptotic gene
Bcl-2 and thus decreased apoptotic rate in pancreatic β-cell [
49]. In contrast,
Tp53 has been described as essential mediator of cell cycle arrest or pro-apoptotic gene, which has strong genetic and biochemical ties with
Bcl-2 [
50]. Yoo
et al. reported that inhibition of cell growth may occur through up-regulation of p53 which leading to G1/S cell cycle arrest [
51]. Zhang
et al. also reported that SOX4 transcription factor inhibits Glioblastoma cell growth partly via the activation of p53-p21 signaling which induces G0/G1 cell cycle arrest [
52]. Therefore, taking all together we could postulate that down-regulation of key microRNAs due to IGF-1 induction simultaneously activate the agonist and antagonist of cell growth, which on the one hand, enhanced cell proliferation, but on the other hand, might control the rate of cell proliferation.
Cell growth and survival-related pathways, including the pathways in cancer (KEGG 5200), MAPK signaling (KEGG 4010), apoptosis (KEGG 4210), cytokine-cytokine receptor interaction (KEGG 4060), and TGF-β (KEGG 4350) cell cycle, were significantly (
p < 0.05, FDR < 0.01) enriched in the predicted target genes of down-regulated microRNAs in Groups A and B. This suggests that the down-regulation of microRNAs due to growth factors is biologically functional (
Table 5). Up-regulated genes due to the down-regulation of microRNAs by EGF, bFGF, and IGF-1 treatment were predicted to target EGFR, FGFR, and IGFR, respectively, via cytokine-cytokine receptor interaction in the cancer pathway. This leads to the activation of Ras protein, which is involved in MAPK and PI3K-Akt signaling pathways. The MAPK [
53,
54] and PI3K-Akt pathways [
43] have been widely reported to be involved in cell proliferation and evading apoptosis. Moreover, the effects of growth factors on the Ras/MAPK/ERK signaling pathway inhibit the TGF-β pathway, which blocks cells from apoptosis and neurogenesis. At the same time, the activated MAPK pathway also triggers the expression of c-Myc regulator genes, which inhibit cyclin-dependent kinase inhibitor 2B on the TGF-β pathway, and leads to cell cycle arrest at G1 phase. In addition, the involvement of the apoptosis pathway showed that down-regulated microRNAs regulate the effect of IAP and Bcl-2, which suppress apoptotic activity [
55,
56]. Therefore, we postulate that the crosstalk between the MAPK and TGF-β pathways is the underlying mechanism of maintaining NPCs in undifferentiated state and producing less apoptotic cells [
57].
The other functionally related pathways identified were the ErbB (KEGG 4010) and Toll-like signaling pathways (KEGG 4620). EGF targets the ErbB-1 (EGFR) receptor in the ErbB signaling pathway, which similarly leads to the activation of the MAPK and PI3K-Akt signaling pathways and, thus, regulates cell proliferation. The ErbB signaling pathway has been shown to promote the proliferation of neoplastic Schwann cells [
58]. The Toll-like signaling pathway was reported to modulate the NPC cell-fate decision and provide neuroprotective effects through the activation of Toll-like receptors (TLRs), such as TLR2 and TLR4 [
59,
60]. In summary, each pathway is interconnected with each other and, as a result, promotes NPCs proliferation and reduces cells apoptosis.