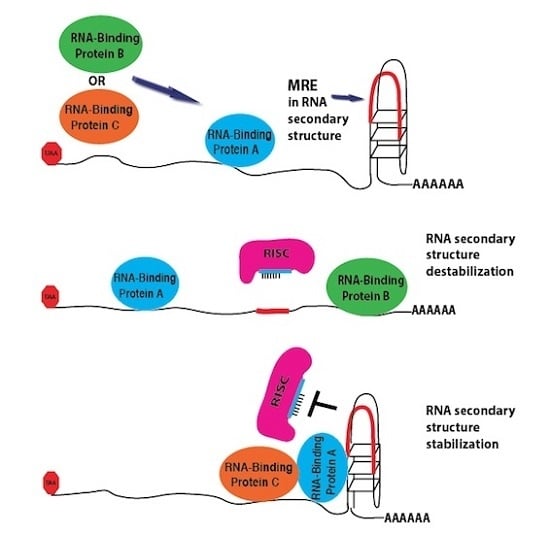
RNA Secondary Structure Modulates FMRP’s Bi-Functional Role in the MicroRNA Pathway
Abstract
:1. Introduction
2. Fragile X Mental Retardation Protein (FMRP), Structure and Function
3. FMRP as a Translation Suppressor
4. The Role of FMRP in MicroRNA (miRNA)-Mediated Translational Regulation
5. RNA Secondary Structure and Its Effect on miRNA Translational Suppression
6. Resolving RNA Secondary Structure: The Role of Moloney Leukemia Virus 10 (MOV10) and RNA Helicases in miRNA Function
7. FMRP Modulates an Agonistic or Antagonistic miRNA Function through Its Interaction with MOV10
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lozano, R.; Rosero, C.A.; Hagerman, R.J. Fragile X spectrum disorders. Intractab. Rare Dis. Res. 2014, 3, 134–146. [Google Scholar] [CrossRef] [PubMed]
- Kelleher, R.J., III; Bear, M.F. The autistic neuron: Troubled translation? Cell 2008, 135, 401–406. [Google Scholar] [CrossRef] [PubMed]
- Hagerman, R.J.; Berry-Kravis, E.; Kaufmann, W.E.; Ono, M.Y.; Tartaglia, N.; Lachiewicz, A.; Kronk, R.; Delahunty, C.; Hessl, D.; Visootsak, J.; et al. Advances in the Treatment of Fragile X Syndrome. Pediatrics 2009, 123, 378–390. [Google Scholar] [CrossRef] [PubMed]
- Kau, A.S.; Meyer, W.A.; Kaufmann, W.E. Early development in males with Fragile X Syndrome: A review of the literature. Microsc. Res. Tech. 2002, 57, 174–178. [Google Scholar] [CrossRef] [PubMed]
- Ashley, C.T.J.; Wilkinson, K.D.; Reines, D.; Warren, S.T. FMR1 protein: Conserved RNP family domains and selective RNA binding. Science 1993, 262, 563–566. [Google Scholar] [CrossRef] [PubMed]
- Brown, V.; Jin, P.; Ceman, S.; Darnell, J.C.; O’Donnell, W.T.; Tenenbaum, S.A.; Jin, X.; Feng, Y.; Wilkinson, K.D.; Keene, J.D.; et al. Microarray identification of FMRP-associated brain mRNAs and altered mRNA translational profiles in Fragile X Syndrome. Cell 2001, 107, 477–487. [Google Scholar] [CrossRef]
- Pieretti, M.; Zhang, F.; Fu, Y.H.; Warren, S.T.; Oostra, B.A.; Caskey, C.T.; Nelson, D.L. Absence of expression of the FMR-1 gene in Fragile X Syndrome. Cell 1991, 66, 817–822. [Google Scholar] [CrossRef]
- Collins, S.C.; Bray, S.M.; Suhl, J.A.; Cutler, D.J.; Coffee, B.; Zwick, M.E.; Warren, S.T. Identification of novel FMR1 variants by massively parallel sequencing in developmentally delayed males. Am. J. Med. Genet. Part A 2010, 152A, 2512–2520. [Google Scholar] [CrossRef] [PubMed]
- Myrick, L.K.; Hashimoto, H.; Cheng, X.; Warren, S.T. Human FMRP contains an integral tandem Agenet (Tudor) and KH motif in the amino terminal domain. Hum. Mol. Genet. 2015, 24, 1733–1740. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Chen, Z.; Fu, Y.; He, Q.; Jiang, L.; Zheng, J.; Gao, Y.; Mei, P.; Chen, Z.; Ren, X. The amino-terminal structure of human fragile X mental retardation protein obtained using precipitant-immobilized imprinted polymers. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Absher, D.; Eberhart, D.E.; Brown, V.; Malter, H.E.; Warren, S.T. FMRP associates with polyribosomes as an mRNP, and the I304N mutation of severe Fragile X Syndrome abolishes this association. Mol. Cell 1997, 1, 109–118. [Google Scholar] [CrossRef]
- Devys, D.; Lutz, Y.; Rouyer, N.; Bellocq, J.-P.; Mandel, J.-L. The FMR-1 protein is cytoplasmic, most abundant in neurons, and appears normal in carriers of the fragile X premutation. Nat. Genet. 1993, 4, 335–340. [Google Scholar] [CrossRef] [PubMed]
- Hinds, H.; Ashely, C.; Sutcliffe, J.; Nelson, D.; Warren, S.; Housman, D.; Schalling, M. Tissue specific expression of FMR-1 provides evidence for a functional role in Fragile X Syndrome. Nat. Genet. 1993, 3, 36–43. [Google Scholar] [CrossRef] [PubMed]
- Sittler, A.; Devys, D.; Weber, C.; Mandel, J.L. Alternative splicing of exon 14 determines nuclear or cytoplasmic localisation of FMR1 protein isoforms. Hum. Mol. Genet. 1996, 5, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Eichler, E.E.; Richards, S.; Gibbs, R.A.; Nelson, D.L. Fine structure of the human FMR1 gene. Hum. Mol. Genet. 1993, 2, 1147–1153. [Google Scholar] [CrossRef] [PubMed]
- Siomi, H.; Siomi, M.C.; Nussbaum, R.L.; Dreyfuss, G. The protein product of the fragile X gene, FMR1, has characteristics of an RNA-binding protein. Cell 1993, 74, 291–298. [Google Scholar] [CrossRef]
- Darnell, J.C.; Fraser, C.E.; Mostovetsky, O.; Stefani, G.; Jones, T.A.; Eddy, S.R.; Darnell, R.B. Kissing complex RNAs mediate interaction between the Fragile-X mental retardation protein KH2 domain and brain polyribosomes. Genes Dev. 2005, 19, 903–918. [Google Scholar] [CrossRef] [PubMed]
- Ascano, M.; Mukherjee, N.; Bandaru, P.; Miller, J.B.; Nusbaum, J.D.; Corcoran, D.L.; Langlois, C.; Munschauer, M.; Dewell, S.; Hafner, M.; et al. FMRP targets distinct mRNA sequence elements to regulate protein expression. Nature 2012, 492, 382–386. [Google Scholar] [CrossRef] [PubMed]
- Suhl, J.A.; Chopra, P.; Anderson, B.R.; Bassell, G.J.; Warren, S.T. Analysis of FMRP mRNA target datasets reveals highly associated mRNAs mediated by G-quadruplex structures formed via clustered WGGA sequences. Hum. Mol. Genet. 2014, 23, 5479–5491. [Google Scholar] [CrossRef] [PubMed]
- Phan, A.T.; Kuryavyi, V.; Darnell, J.C.; Serganov, A.; Majumdar, A.; Ilin, S.; Raslin, T.; Polonskaia, A.; Chen, C.; Clain, D.; et al. Structure-function studies of FMRP RGG peptide recognition of an RNA duplex-quadruplex junction. Nat. Struct. Mol. Biol. 2011, 18, 796–804. [Google Scholar] [CrossRef] [PubMed]
- Darnell, J.C.; Jensen, K.B.; Jin, P.; Brown, V.; Warren, S.T.; Darnell, R.B. Fragile X mental retardation protein targets G quartet mRNAs important for neuronal function. Cell 2001, 107, 489–499. [Google Scholar] [CrossRef]
- Schaeffer, C.; Bardoni, B.; Mandel, J.L.; Ehresmann, B.; Ehresmann, C.; Moine, H. The fragile X mental retardation protein binds specifically to its mRNA via a purine quartet motif. EMBO J. 2001, 20, 4803–4813. [Google Scholar] [CrossRef] [PubMed]
- Ashley, C.T.; Sutcliffe, J.S.; Kunst, C.B.; Leiner, H.A.; Eichler, E.E.; Nelson, D.L.; Warren, S.T. Human and murine FMR-1: Alternative splicing and translational initiation downstream of the CGG-repeat. Nat. Genet. 1993, 4, 244–251. [Google Scholar] [CrossRef] [PubMed]
- Sung, Y.J.; Dolzhanskaya, N.; Nolin, S.L.; Brown, T.; Currie, J.R.; Denman, R.B. The fragile X mental retardation protein FMRP binds elongation factor 1A mRNA and negatively regulates its translation in vivo. J. Biol. Chem. 2003, 278, 15669–15678. [Google Scholar] [CrossRef] [PubMed]
- Brown, V.; Small, K.; Lakkis, L.; Feng, Y.; Gunter, C.; Wilkinson, K.D.; Warren, S.T. Purified recombinant FMRP exhibits selective RNA binding as an intrinsic property of the fragile X mental retardation protein. J. Biol. Chem. 1998, 273, 15521–15527. [Google Scholar] [CrossRef] [PubMed]
- Darnell, J.C.; van Driesche, S.J.; Zhang, C.; Hung, K.Y.; Mele, A.; Fraser, C.E.; Stone, E.F.; Chen, C.; Fak, J.J.; et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 2011, 146, 247–261. [Google Scholar] [CrossRef] [PubMed]
- Vasilyev, N.; Polonskaia, A.; Darnell, J.C.; Darnell, R.B.; Patel, D.J.; Serganov, A. Crystal structure reveals specific recognition of a G-quadruplex RNA by a β-turn in the RGG motif of FMRP. Proc. Natl. Acad. Sci. USA 2015, 112, E5391–E5400. [Google Scholar] [CrossRef] [PubMed]
- Myrick, L.K.; Deng, P.-Y.; Hashimoto, H.; Oh, Y.M.; Cho, Y.; Poidevin, M.J.; Suhl, J.A.; Visootsak, J.; Cavalli, V.; Jin, P.; et al. Independent role for presynaptic FMRP revealed by an FMR1 missense mutation associated with intellectual disability and seizures. Proc. Natl. Acad. Sci. USA 2015, 112, 949–956. [Google Scholar] [CrossRef] [PubMed]
- Deng, P.Y.; Rotman, Z.; Blundon, J.A.; Cho, Y.; Cui, J.; Cavalli, V.; Zakharenko, S.S.; Klyachko, V.A. FMRP regulates neurotransmitter release and synaptic information transmission by modulating action potential duration via BK channels. Neuron 2013, 77, 696–711. [Google Scholar] [CrossRef] [PubMed]
- Adams-Cioaba, M.A.; Guo, Y.; Bian, C.; Amaya, M.F.; Lam, R.; Wasney, G.A.; Vedadi, M.; Xu, C.; Min, J. Structural studies of the tandem Tudor domains of Fragile X mental retardation related proteins FXR1 and FXR2. PLoS ONE 2010, 5, e13559. [Google Scholar] [CrossRef] [PubMed]
- Alpatov, R.; Lesch, B.J.; Nakamoto-Kinoshita, M.; Blanco, A.; Chen, S.; Stutzer, A.; Armache, K.J.; Simon, M.D.; Xu, C.; Ali, M.; et al. A chromatin-dependent role of the fragile X mental retardation protein FMRP in the DNA damage response. Cell 2014, 157, 869–881. [Google Scholar] [CrossRef] [PubMed]
- Maurer-Stroh, S.; Dickens, N.J.; Hughes-Davies, L.; Kouzarides, T.; Eisenhaber, F.; Ponting, C.P. The Tudor domain ‘Royal Family’: Tudor, plant Agenet, Chromo, PWWP and MBT domains. Trends Biochem. Sci. 2003, 28, 69–74. [Google Scholar] [CrossRef]
- Ramos, A.; Hollingworth, D.; Adinolfi, S.; Castets, M.; Kelly, G.; Frenkiel, T.A.; Bardoni, B.; Pastore, A. The structure of the N-terminal domain of the fragile X mental retardation protein: A platform for protein–protein interaction. Structure 2006, 14, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Zalfa, F.; Adinolfi, S.; Napoli, I.; Kuhn-Holsken, E.; Urlaub, H.; Achsel, T.; Pastore, A.; Bagni, C. Fragile X Mental Retardation Protein (FMRP) binds specifically to the brain cytoplasmic RNAs BC1/BC200 via a novel RNA-binding motif. J. Biol. Chem. 2005, 280, 33403–33410. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Bellini, M.; Ceman, S. Fragile X mental retardation protein FMRP binds mRNAs in the nucleus. Mol. Cell. Biol. 2009, 29, 214–228. [Google Scholar] [CrossRef] [PubMed]
- Kato, M.; Han, T.W.; Xie, S.; Shi, K.; Du, X.; Wu, L.C.; Mirzaei, H.; Goldsmith, E.J.; Longgood, J.; Pei, J.; et al. Cell-free formation of RNA granules: Low complexity sequence domains form dynamic fibers within hydrogels. Cell 2012, 149, 753–767. [Google Scholar] [CrossRef] [PubMed]
- Coletta, A.; Pinney, J.W.; Solis, D.Y.; Marsh, J.; Pettifer, S.R.; Attwood, T.K. Low-complexity regions within protein sequences have position-dependent roles. BMC Syst. Biol. 2010, 4, 43. [Google Scholar] [CrossRef] [PubMed]
- Siomi, M.C.; Zhang, Y.; Siomi, H.; Dreyfuss, G. Specific sequences in the Fragile X Syndrome protein FMR1 and the FXR proteins mediate their binding to 60S ribosomal subunits and the interactions among them. Mol. Cell. Biol. 1996, 16, 3825–3832. [Google Scholar] [CrossRef] [PubMed]
- Laggerbauer, B.; Ostareck, D.; Keidel, E.M.; Ostareck-Lederer, A.; Fischer, U. Evidence that fragile X mental retardation protein is a negative regulator of translation. Hum. Mol. Genet. 2001, 10, 329–338. [Google Scholar] [CrossRef] [PubMed]
- Ceman, S.; O’Donnell, W.T.; Reed, M.; Patton, S.; Pohl, J.; Warren, S.T. Phosphorylation influences the translation state of FMRP-associated polyribosomes. Hum. Mol. Genet. 2003, 12, 3295–3305. [Google Scholar] [CrossRef] [PubMed]
- Chen, E.; Sharma, M.R.; Shi, X.; Agrawal, R.K.; Joseph, S. Fragile X mental retardation protein regulates translation by binding directly to the ribosome. Mol. Cell 2014, 54, 407–417. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; O’Connor, J.P.; Siomi, M.C.; Srinivasan, S.; Dutra, A.; Nussbaum, R.L.; Dreyfuss, G. The fragile X mental retardation syndrome protein interacts with novel homologs FXR1 and FXR2. EMBO J. 1995, 14, 5358–5366. [Google Scholar] [PubMed]
- Dockendorff, T.C.; Su, H.S.; McBride, S.M.; Yang, Z.; Choi, C.H.; Siwicki, K.K.; Sehgal, A.; Jongens, T.A. Drosophila lacking dfmr1 activity show defects in circadian output and fail to maintain courtship interest. Neuron 2002, 34, 973–984. [Google Scholar] [CrossRef]
- Napoli, I.; Mercaldo, V.; Boyl, P.P.; Eleuteri, B.; Zalfa, F.; De Rubeis, S.; Di Marino, D.; Mohr, E.; Massimi, M.; Falconi, M.; et al. The Fragile X Syndrome protein represses activity-dependent translation through CYFIP1, a new 4E-BP. Cell 2008, 134, 1042. [Google Scholar] [CrossRef] [PubMed]
- Genheden, M.; Kenney, J.W.; Johnston, H.E. BDNF stimulation of protein synthesis in cortical neurons requires the MAP kinase-interacting kinase MNK1. J. Neurosci. 2015, 35, 972–984. [Google Scholar] [CrossRef] [PubMed]
- Panja, D.; Kenney, J.W.; D’Andrea, L.; Zalfa, F.; Vedeler, A.; Wibrand, K.; Fukunaga, R.; Bagni, C.; Proud, C.G.; Bramham, C.R. Two-stage translational control of dentate gyrus LTP consolidation is mediated by sustained BDNF-TrkB signaling to MNK. Cell Rep. 2014, 9, 1430–1445. [Google Scholar] [CrossRef] [PubMed]
- Huber, K.M.; Gallagher, S.M.; Warren, S.T.; Bear, M.F. Altered synaptic plasticity in a mouse model of fragile X mental retardation. Proc. Natl. Acad. Sci. USA 2002, 99, 7746–7750. [Google Scholar] [CrossRef] [PubMed]
- Sidorov, M.S.; Auerbach, B.D.; Bear, M.F. Fragile X mental retardation protein and synaptic plasticity. Mol. Brain 2013, 6, 15. [Google Scholar] [CrossRef] [PubMed]
- Muddashetty, R.S.; Nalavadi, V.C.; Gross, C.; Yao, X.; Xing, L.; Laur, O.; Warren, S.T.; Bassell, G.J. Reversible inhibition of PSD-95 mRNA translation by miR-125a, FMRP phosphorylation, and mGluR signaling. Mol. Cell 2011, 42, 673–688. [Google Scholar] [CrossRef] [PubMed]
- Kenny, P.J.; Zhou, H.; Kim, M.; Skariah, G.; Khetani, R.S.; Drnevich, J.; Arcila, M.L.; Kosik, K.S.; Ceman, S. MOV10 and FMRP regulate AGO2 association with microRNA recognition elements. Cell Rep. 2014, 9, 1729–1741. [Google Scholar] [CrossRef] [PubMed]
- Caudy, A.A.; Myers, M.; Hannon, G.J.; Hammond, S.M. Fragile X-related protein and VIG associate with the RNA interference machinery. Genes Dev. 2002, 16, 2491–2496. [Google Scholar] [CrossRef] [PubMed]
- Ishizuka, A.; Siomi, M.C.; Siomi, H. A Drosophila fragile X protein interacts with components of RNAi and ribosomal proteins. Genes Dev. 2002, 16, 2497–2508. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, E.; Li, K.W.; Rajan, N.; De Rubeis, S.; Fiers, M. FXR2P Exerts a Positive Translational Control and Is Required for the Activity-Dependent Increase of PSD95 Expression. J. Neurosci. 2015, 35, 9402–9408. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.-L.; Zong, R.; Li, Z.; Biswas, M.H.U.; Fang, Z.; Nelson, D.L.; Gao, F.-B. FXR1P But not FMRP Regulates the Levels of Mammalian Brain-Specific microRNA-9 and microRNA-124. J. Neurosci. 2011, 31, 13705–13709. [Google Scholar] [CrossRef] [PubMed]
- Jin, P.; Zarnescu, D.C.; Ceman, S.; Nakamoto, M.; Mowrey, J.; Jongens, T.A.; Nelson, D.L.; Moses, K.; Warren, S.T. Biochemical and genetic interaction between the fragile X mental retardation protein and the microRNA pathway. Nat. Neurosci. 2004, 7, 113–117. [Google Scholar] [CrossRef] [PubMed]
- Cheever, A.; Ceman, S. Phosphorylation of FMRP inhibits association with Dicer. RNA 2009, 15, 362–369. [Google Scholar] [CrossRef] [PubMed]
- Kedersha, N.; Anderson, P. Mammalian stress granules and processing bodies. Methods Enzymol. 2007, 431, 61–81. [Google Scholar] [PubMed]
- Teixeira, D.; Sheth, U.; Valencia-Sanchez, M.A.; Brengues, M.; Parker, R. Processing bodies require RNA for assembly and contain nontranslating mRNAs. RNA 2005, 11, 371–382. [Google Scholar] [CrossRef] [PubMed]
- Parker, R.; Sheth, U. P bodies and the control of mRNA translation and degradation. Mol. Cell 2007, 25, 635–646. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.K.; Kim, H.H.; Kuwano, Y.; Abdelmohsen, K.; Srikantan, S.; Subaran, S.S.; Gleichmann, M.; Mughal, M.R.; Martindale, J.L.; Yang, X. hnRNP C promotes APP translation by competing with FMRP for APP mRNA recruitment to P bodies. Nat. Struct. Mol. Biol. 2010, 17, 732–739. [Google Scholar] [CrossRef] [PubMed]
- Edbauer, D.; Neilson, J.R.; Foster, K.A.; Wang, C.-F.; Seeburg, D.P.; Batterton, M.N.; Tada, T.; Dolan, B.M.; Sharp, P.A.; Sheng, M. Regulation of synaptic structure and function by FMRP-associated microRNAs miR-125b and miR-132. Neuron 2010, 65, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Sievers, C.; Schlumpf, T.; Sawarkar, R.; Comoglio, F.; Paro, R. Mixture models and wavelet transforms reveal high confidence RNA-protein interaction sites in MOV10 PAR-CLIP data. Nucleic Acids Res. 2012, 40, e160. [Google Scholar] [CrossRef] [PubMed]
- Meister, G.; Landthaler, M.; Peters, L.; Chen, P.Y.; Urlaub, H.; Luhrmann, R.; Tuschl, T. Identification of novel argonaute-associated proteins. Curr. Biol. 2005, 15, 2149–2155. [Google Scholar] [CrossRef] [PubMed]
- Van Kouwenhove, M.; Kedde, M.; Agami, R. MicroRNA regulation by RNA-binding proteins and its implications for cancer. Nat. Rev. Cancer 2011, 11, 644–656. [Google Scholar] [CrossRef] [PubMed]
- Kedde, M.; van Kouwenhove, M.; Zwart, W.; Oude Vrielink, J.A.F.; Elkon, R.; Agami, R. A Pumilio-induced RNA structure switch in p27-3′ UTR controls miR-221 and miR-222 accessibility. Nat. Cell Biol. 2010, 12, 1014–1020. [Google Scholar] [CrossRef] [PubMed]
- Fabian, M.R.; Sonenberg, N. The mechanics of miRNA-mediated gene silencing: A look under the hood of miRISC. Nat. Struct. Mol. Biol. 2012, 19, 586–593. [Google Scholar] [CrossRef] [PubMed]
- Kedde, M.; Strasser, M.J.; Boldajipour, B.; Oude Vrielink, J.A.F.; Slanchev, K.; Le Sage, C.; Nagel, R.; Voorhoeve, P.M.; van Duijse, J.; ÿrom, U.A.; et al. RNA-Binding Protein Dnd1 Inhibits microRNA Access to Target mRNA. Cell 2007, 131, 1273–1286. [Google Scholar] [CrossRef] [PubMed]
- Goswami, S.; Tarapore, R.S.; Strong, A.M.P.; TeSlaa, J.J.; Grinblat, Y.; Setaluri, V.; Spiegelman, V.S. MicroRNA-340-mediated degradation of microphthalmia-associated transcription factor (MITF) mRNA is inhibited by coding region determinant-binding protein (CRD-BP). J. Biol. Chem. 2015, 290, 384–395. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.A.; Damianov, A.; Lin, C.H.; Fontes, M.; Parikshak, N.N.; Anderson, E.S.; Geschwind, D.H.; Black, D.L.; Martin, K.C. Cytoplasmic Rbfox1 regulates the expression of synaptic and autism-related genes. Neuron 2016, 89, 113–128. [Google Scholar] [CrossRef] [PubMed]
- Xue, Y.; Ouyang, K.; Huang, J.; Zhou, Y.; Ouyang, H.; Li, H.; Wang, G.; Wu, Q.; Wei, C.; Bi, Y.; et al. Direct conversion of fibroblasts to neurons by reprogramming PTB-regulated microRNA circuits. Cell 2013, 152, 82–96. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.H.; Kuwano, Y.; Srikantan, S.; Lee, E.K.; Martindale, J.L.; Gorospe, M. HuR recruits let-7/RISC to repress c-Myc expression. Genes Dev. 2009, 23, 1743–1748. [Google Scholar] [CrossRef] [PubMed]
- Srikantan, S.; Abdelmohsen, K.; Lee, E.K.; Tominaga, K.; Subaran, S.S.; Kuwano, Y.; Kulshrestha, R.; Panchakshari, R.; Kim, H.H.; Yang, X.; et al. Translational control of TOP2A influences doxorubicin efficacy. Mol. Cell. Biol. 2011, 31, 3790–3801. [Google Scholar] [CrossRef] [PubMed]
- Kundu, P.; Fabian, M.R.; Sonenberg, N.; Bhattacharyya, S.N.; Filipowicz, W. HuR protein attenuates miRNA-mediated repression by promoting miRISC dissociation from the target RNA. Nucleic Acids Res. 2012, 40, 5088–5100. [Google Scholar] [CrossRef] [PubMed]
- Long, D.; Lee, R.; Williams, P.; Chan, C.Y.; Ambros, V.; Ding, Y. Potent effect of target structure on microRNA function. Nat. Struct. Mol. Biol. 2007, 14, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Arthanari, H.; Bolton, P.H. Functional and dysfunctional roles of quadruplex DNA in cells. Chem. Biol. 2001, 8, 221–230. [Google Scholar] [CrossRef]
- Kostadinov, R.; Malhotra, N.; Viotti, M.; Shine, R.; D’Antonio, L.; Bagga, P. GRSDB: A database of quadruplex forming G-rich sequences in alternatively processed mammalian pre-mRNA sequences. Nucleic Acids Res. 2006, 34 (Suppl. S1), D119–D124. [Google Scholar] [CrossRef] [PubMed]
- Williamson, J.R.; Raghuraman, M.K.; Cech, T.R. Monovalent cation-induced structure of telomeric DNA: The G-quartet model. Cell 1989, 59, 871–880. [Google Scholar] [CrossRef]
- Creacy, S.D.; Routh, E.D.; Iwamoto, F.; Nagamine, Y.; Akman, S.A.; Vaughn, J.P. G4 resolvase 1 binds both DNA and RNA tetramolecular quadruplex with high affinity and is the major source of tetramolecular quadruplex G4-DNA and G4-RNA resolving activity in HeLa cell lysates. J. Biol. Chem. 2008, 283, 34626–34634. [Google Scholar] [CrossRef] [PubMed]
- Beaudoin, J.-D.; Perreault, J.-P. 5′-UTR G-quadruplex structures acting as translational repressors. Nucleic Acids Res. 2010, 38, 7022–7036. [Google Scholar] [CrossRef] [PubMed]
- Morris, M.J.; Basu, S. An unusually stable G-quadruplex within the 5′ UTR of the MT3 matrix metalloproteinase mRNA represses translation in eukaryotic cells. Biochemistry 2009, 48, 5313–5319. [Google Scholar] [CrossRef] [PubMed]
- Millevoi, S.; Moine, H.; Vagner, S. G-quadruplexes in RNA biology. Wiley Interdiscip. Rev. RNA 2012, 3, 495–507. [Google Scholar] [CrossRef] [PubMed]
- Ameres, S.L.; Martinez, J.; Schroeder, R. Molecular basis for target RNA recognition and cleavage by human RISC. Cell 2007, 130, 101–112. [Google Scholar] [CrossRef] [PubMed]
- Jo, M.H.; Shin, S.; Jung, S.R.; Kim, E.; Song, J.J.; Hohng, S. Human argonaute 2 has diverse reaction pathways on target RNAs. Mol. Cell 2015, 59, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Stefanovic, S.; Bassell, G.J.; Mihailescu, M.R. G quadruplex RNA structures in PSD-95 mRNA: Potential regulators of miR-125a seed binding site accessibility. RNA 2015, 21, 48–60. [Google Scholar] [CrossRef] [PubMed]
- Lattmann, S.; Giri, B.; Vaughn, J.P.; Akman, S.A.; Nagamine, Y. Role of the amino terminal RHAU-specific motif in the recognition and resolution of guanine quadruplex-RNA by the DEAH-box RNA helicase RHAU. Nucleic Acids Res. 2010, 38, 6219–6233. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Ma, W.; Nie, J.; Zhang, C.; Zhang, J.; Yao, G.; Han, J.; Xu, J.; Hu, B.; Du, Y.; et al. A G-quadruplex DNA structure resolvase, RHAU, is essential for spermatogonia differentiation. Cell Death Dis. 2015, 6, e1610. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, P.; Grosse, F. Human DHX9 helicase preferentially unwinds RNA-containing displacement loops (R-loops) and G-quadruplexes. DNA Repair 2011, 10, 654–665. [Google Scholar] [CrossRef] [PubMed]
- Gregersen, L.H.; Schueler, M.; Munschauer, M.; Mastrobuoni, G.; Chen, W.; Kempa, S.; Dieterich, C.; Landthaler, M. MOV10 is a 5′ to 3′ RNA helicase contributing to UPF1 mRNA target degradation by translocation along 3′ UTRs. Mol. Cell 2014, 54, 573–585. [Google Scholar] [CrossRef] [PubMed]
- Jankowsky, E. RNA helicases at work: Binding and rearranging. Trends Biochem. Sci. 2011, 36, 19–29. [Google Scholar] [CrossRef] [PubMed]
- Goodier, J.L.; Cheung, L.E.; Kazazian, H.H., Jr. MOV10 RNA helicase is a potent inhibitor of retrotransposition in cells. PLoS Genet. 2012, 8, e1002941. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, J.; Jia, R.; Cheng, V.; Xu, X.; Qiao, W.; Guo, F.; Liang, C.; Cen, S. The MOV10 helicase inhibits LINE-1 mobility. J. Biol. Chem. 2013, 288, 21148–21160. [Google Scholar] [CrossRef] [PubMed]




© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kenny, P.; Ceman, S. RNA Secondary Structure Modulates FMRP’s Bi-Functional Role in the MicroRNA Pathway. Int. J. Mol. Sci. 2016, 17, 985. https://doi.org/10.3390/ijms17060985
Kenny P, Ceman S. RNA Secondary Structure Modulates FMRP’s Bi-Functional Role in the MicroRNA Pathway. International Journal of Molecular Sciences. 2016; 17(6):985. https://doi.org/10.3390/ijms17060985
Chicago/Turabian StyleKenny, Phillip, and Stephanie Ceman. 2016. "RNA Secondary Structure Modulates FMRP’s Bi-Functional Role in the MicroRNA Pathway" International Journal of Molecular Sciences 17, no. 6: 985. https://doi.org/10.3390/ijms17060985