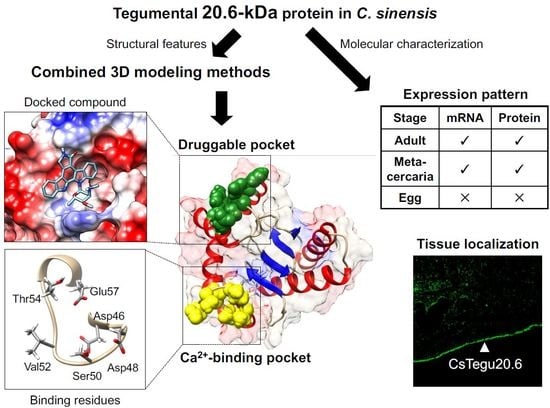
Molecular and Structural Characterization of the Tegumental 20.6-kDa Protein in Clonorchis sinensis as a Potential Druggable Target
Abstract
:1. Introduction
2. Results and Discussion
2.1. Physico-Chemical and Functional Characterization
2.2. Sequence-Based Similarities
2.3. Improved and Full-Length 3D Models Using a Combined Approach
2.4. Structure Validation
2.5. Structure-Based Similarity
2.6. Overall Structural Features and Dimerization
2.7. EF-Hand Calcium-Binding Site
2.8. Virtual Inhibitor Screening
2.9. Production of Recombinant CsTegu20.6
2.10. Stage-Specific Expression
2.11. Immunolocalization in Adult C. sinensis
3. Materials and Methods
3.1. Sequence Identification and Characterization
3.2. Combined 3D Modeling Methods and Refinement
3.3. Model Evaluation and Quality Assessment
3.4. Structural Comparison and Visualization
3.5. Protein-Compound Interaction and Drug-Likeness
3.6. Ethics Statement
3.7. Adult Worms and Sera
3.8. Plasmid Construction
3.9. Expression and Purification
3.10. Preparation of Antiserum
3.11. Analysis of Protein Expression during C. sinensis Developmental Stages
3.12. Transcript Analysis of Developmental cDNA Libraries in C. sinensis
3.13. Immunolocalization of CsTegu20.6 in C. sinensis Adult Worms
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| STU | Staurosporine |
| ANP | Phosphoaminophosphonic acid-adenylate ester |
| BK3 | 3-(naphthalen-1-ylmethyl)-1-(piperidin-4-ylmethyl)-1H-pyrazolo[3,4-d]pyrimidin-4-amine |
| CRK | 4-{(z)-[2-[3-(methylsulfanyl)propanoyl]-5-oxo-1-(2-oxoethyl)-1,5-dihydro-4h-imidazol-4-ylidene]methyl}benzenolate |
| DTQ | 4-[3-hydroxyanilino]-6,7-dimethoxyquinazoline |
| MRD | (4r)-2-methylpentane-2,4-diol |
References
- Yu, S.H.; Kawanaka, M.; Li, X.M.; Xu, L.Q.; Lan, C.G.; Rui, L. Epidemiological investigation on Clonorchis sinensis in human population in an area of South China. Jpn. J. Infect. Dis. 2003, 56, 168–171. [Google Scholar] [PubMed]
- Wang, K.X.; Zhang, R.B.; Cui, Y.B.; Tian, Y.; Cai, R.; Li, C.P. Clinical and epidemiological features of patients with clonorchiasis. World J. Gastroenterol. 2004, 10, 446–448. [Google Scholar] [PubMed]
- Rim, H.J. Clonorchiasis: An update. J. Helminthol. 2005, 79, 269–281. [Google Scholar] [CrossRef] [PubMed]
- Keiser, J.; Utzinger, J. Emerging foodborne trematodiasis. Emerg. Infect. Dis. 2005, 11, 1507–1514. [Google Scholar] [CrossRef] [PubMed]
- Bouvard, V.; Baan, R.; Straif, K.; Grosse, Y.; Secretan, B.; El Ghissassi, F.; Benbrahim-Tallaa, L.; Guha, N.; Freeman, C.; Galichet, L.; et al. A review of human carcinogens—Part B: Biological agents. Lancet Oncol. 2009, 10, 321–322. [Google Scholar] [CrossRef]
- Kim, T.S.; Cho, S.H.; Huh, S.; Kong, Y.; Sohn, W.M.; Hwang, S.S.; Chai, J.Y.; Lee, S.H.; Park, Y.K.; Oh, D.K.; et al. A nationwide survey on the prevalence of intestinal parasitic infections in the Republic of Korea, 2004. Korean J. Parasitol. 2009, 47, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Choi, M.H.; Park, S.K.; Li, Z.; Ji, Z.; Yu, G.; Feng, Z.; Xu, L.; Cho, S.Y.; Rim, H.J.; Lee, S.H.; et al. Effect of control strategies on prevalence, incidence and re-infection of clonorchiasis in endemic areas of China. PLoS Negl. Trop. Dis. 2010, 4, e601. [Google Scholar] [CrossRef] [PubMed]
- Tinga, N.; De, N.; Vien, H.V.; Chau, L.; Toan, N.D.; Kager, P.A.; Vries, P.J. Little effect of praziquantel or artemisinin on clonorchiasis in Northern Vietnam. A pilot study. Trop. Med. Int. Health 1999, 4, 814–818. [Google Scholar] [CrossRef] [PubMed]
- Loukas, A.; Tran, M.; Pearson, M.S. Schistosome membrane proteins as vaccines. Int. J. Parasitol. 2007, 37, 257–263. [Google Scholar] [CrossRef] [PubMed]
- Mulvenna, J.; Moertel, L.; Jones, M.K.; Nawaratna, S.; Lovas, E.M.; Gobert, G.N.; Colgrave, M.; Jones, A.; Loukas, A.; McManus, D.P. Exposed proteins of the Schistosoma japonicum tegument. Int. J. Parasitol. 2010, 40, 543–554. [Google Scholar] [CrossRef] [PubMed]
- Van Hellemond, J.J.; Retra, K.; Brouwers, J.F.; van Balkom, B.W.; Yazdanbakhsh, M.; Shoemaker, C.B.; Tielens, A.G. Functions of the tegument of schistosomes: Clues from the proteome and lipidome. Int. J. Parasitol. 2006, 36, 691–699. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.K.; Gobert, G.N.; Zhang, L.; Sunderland, P.; McManus, D.P. The cytoskeleton and motor proteins of human schistosomes and their roles in surface maintenance and host-parasite interactions. Bioessays 2004, 26, 752–765. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Xu, H.; Zhang, Z.; Zeng, S.; Gan, W.; Yu, X.; Hu, X. Cloning and expression of 21.1-kDa tegumental protein of Clonorchis sinensis and human antibody response to it as a trematode-nematode pan-specific serodiagnosis antigen. Parasitol. Res. 2011, 108, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Zhou, Z.; Hu, X.; Wei, Q.; Xu, J.; Wu, Z.; Yu, X. A novel tegumental protein 31.8 kDa of Clonorchis sinensis: Sequence analysis, expression, and immunolocalization. Parasitol. Res. 2007, 102, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Hu, X.; Huang, Y.; Hu, H.; Ma, C.; Chen, X.; Hu, F.; Xu, J.; Lu, F.; Wu, Z.; et al. Molecular cloning and identification of a novel Clonorchis sinensis gene encoding a tegumental protein. Parasitol. Res. 2007, 101, 737–742. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Xia, H.; Hu, X.; Huang, Y.; Li, Y.; Li, L.; Ma, C.; Chen, X.; Hu, F.; Xu, J.; et al. Oral administration of a Bacillus subtilis spore-based vaccine expressing Clonorchis sinensis tegumental protein 22.3 kDa confers protection against Clonorchis sinensis. Vaccine 2008, 26, 1817–1825. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Yoo, W.G.; Lee, M.R.; Kim, D.W.; Lee, W.J.; Kang, J.M.; Na, B.K.; Ju, J.W. Identification and characterization of a novel 21.6-kDa tegumental protein from Clonorchis sinensis. Parasitol. Res. 2012, 110, 2061–2066. [Google Scholar] [CrossRef] [PubMed]
- Yoo, W.G.; Kim, D.W.; Ju, J.W.; Cho, P.Y.; Kim, T.I.; Cho, S.H.; Choi, S.H.; Park, H.S.; Kim, T.S.; Hong, S.J. Developmental transcriptomic features of the carcinogenic liver fluke, Clonorchis sinensis. PLoS Negl. Trop. Dis. 2011, 5, e1208. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Cullis, D.N.; Feig, L.A.; Baleja, J.D. Solution structure of the Reps1 EH domain and characterization of its binding to NPF target sequences. Biochemistry 2001, 40, 6776–6785. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, K.F.; Strand, M. Molecular characterization of a 20.8-kDa Schistosoma mansoni antigen. Sequence similarity to tegumental associated antigens and dynein light chains. J. Biol. Chem. 1997, 272, 14509–14515. [Google Scholar] [CrossRef] [PubMed]
- Gouet, P.; Robert, X.; Courcelle, E. ESPript/ENDscript: Extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 2003, 31, 3320–3323. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Bordoli, L.; Kiefer, F.; Arnold, K.; Benkert, P.; Battey, J.; Schwede, T. Protein structure homology modeling using SWISS-MODEL workspace. Nat. Protoc. 2009, 4, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 2015, 43, W174–W181. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhang, Y. Protein Structure and Function Prediction Using I-TASSER. Curr. Protoc. Bioinform. 2015, 52, 5.8.1–5.8.15. [Google Scholar]
- Thomas, C.M.; Fitzsimmons, C.M.; Dunne, D.W.; Timson, D.J. Comparative biochemical analysis of three members of the Schistosoma mansoni TAL family: Differences in ion and drug binding properties. Biochimie 2015, 108, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Banford, S.; Drysdale, O.; Hoey, E.M.; Trudgett, A.; Timson, D.J. FhCaBP3: A Fasciola hepatica calcium binding protein with EF-hand and dynein light chain domains. Biochimie 2013, 95, 751–758. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Y.; Faraggi, E.; Zhao, H.; Zhou, Y. Improving protein fold recognition and template-based modeling by employing probabilistic-based matching between predicted one-dimensional structural properties of query and corresponding native properties of templates. Bioinformatics 2011, 27, 2076–2082. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Zhang, Y. How significant is a protein structure similarity with TM-score = 0.5? Bioinformatics 2010, 26, 889–895. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Zhang, Y. Improving the physical realism and structural accuracy of protein models by a two-step atomic-level energy minimization. Biophys. J. 2011, 101, 2525–2534. [Google Scholar] [CrossRef] [PubMed]
- Omotuyi, O.I.; Ueda, H. A Novel Unified Ab Initio and Template-Based Approach to GPCR Modeling: Case of EDG-LPA Receptors. Curr. Bioinform. 2013, 8, 603–610. [Google Scholar] [CrossRef]
- Park, H.; Seok, C. Refinement of unreliable local regions in template-based protein models. Proteins 2012, 80, 1974–1986. [Google Scholar] [CrossRef] [PubMed]
- Heo, L.; Park, H.; Seok, C. GalaxyRefine: Protein structure refinement driven by side-chain repacking. Nucleic Acids Res. 2013, 41, W384–W388. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; Rullmannn, J.A.; MacArthur, M.W.; Kaptein, R.; Thornton, J.M. AQUA and PROCHECK-NMR: Programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 1996, 8, 477–486. [Google Scholar] [CrossRef] [PubMed]
- Wiederstein, M.; Sippl, M.J. ProSA-web: Interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res. 2007, 35, W407–W410. [Google Scholar] [CrossRef] [PubMed]
- Benkert, P.; Biasini, M.; Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 2011, 27, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Colovos, C.; Yeates, T.O. Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Sci. 1993, 2, 1511–1519. [Google Scholar] [CrossRef] [PubMed]
- McGuffin, L.J.; Buenavista, M.T.; Roche, D.B. The ModFOLD4 server for the quality assessment of 3D protein models. Nucleic Acids Res. 2013, 41, W368–W372. [Google Scholar] [CrossRef] [PubMed]
- Rose, P.W.; Bi, C.; Bluhm, W.F.; Christie, C.H.; Dimitropoulos, D.; Dutta, S.; Green, R.K.; Goodsell, D.S.; Prlic, A.; Quesada, M.; et al. The RCSB Protein Data Bank: New resources for research and education. Nucleic Acids Res. 2013, 41, D475–D482. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.H.; Bharatham, K.; Sherman, W.A.; Mihalek, I. deconSTRUCT: General purpose protein database search on the substructure level. Nucleic Acids Res. 2010, 38, W590–W594. [Google Scholar] [CrossRef] [PubMed]
- Delarue, M.; Poterszman, A.; Nikonov, S.; Garber, M.; Moras, D.; Thierry, J.C. Crystal structure of a prokaryotic aspartyl tRNA-synthetase. EMBO J. 1994, 13, 3219–3229. [Google Scholar] [PubMed]
- Holm, L.; Rosenstrom, P. Dali server: Conservation mapping in 3D. Nucleic Acids Res. 2010, 38, W545–W549. [Google Scholar] [CrossRef] [PubMed]
- Makokha, M.; Huang, Y.J.; Montelione, G.; Edison, A.S.; Barbar, E. The solution structure of the pH-induced monomer of dynein light-chain LC8 from Drosophila. Protein Sci. 2004, 13, 727–734. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Park, H.; Ko, J.; Seok, C. GalaxyGemini: A web server for protein homo-oligomer structure prediction based on similarity. Bioinformatics 2013, 29, 1078–1080. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, B.M.; Hofmann, N.E.; Arroyo-Olarte, R.D.; Nickl, B.; Hoehne, W.; Jungblut, P.R.; Lucius, R.; Scheerer, P.; Gupta, N. Dynein light chain 8a of Toxoplasma gondii, a unique conoid-localized β-strand-swapped homodimer, is required for an efficient parasite growth. FASEB J. 2013, 27, 1034–1047. [Google Scholar] [CrossRef] [PubMed]
- Gifford, J.L.; Walsh, M.P.; Vogel, H.J. Structures and metal-ion-binding properties of the Ca2+-binding helix-loop-helix EF-hand motifs. Biochem. J. 2007, 405, 199–221. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Roy, A.; Zhang, Y. Protein-ligand binding site recognition using complementary binding-specific substructure comparison and sequence profile alignment. Bioinformatics 2013, 29, 2588–2595. [Google Scholar] [CrossRef] [PubMed]
- Heo, L.; Shin, W.H.; Lee, M.S.; Seok, C. GalaxySite: Ligand-binding-site prediction by using molecular docking. Nucleic Acids Res. 2014, 42, W210–W214. [Google Scholar] [CrossRef] [PubMed]
- Cheng, F.; Li, W.; Zhou, Y.; Shen, J.; Wu, Z.; Liu, G.; Lee, P.W.; Tang, Y. admetSAR: A comprehensive source and free tool for assessment of chemical ADMET properties. J. Chem. Inf. Model. 2012, 52, 3099–3105. [Google Scholar] [CrossRef] [PubMed]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 2001, 46, 3–26. [Google Scholar] [CrossRef]
- Becker, S.; Jaffe, C.L. Effect of protein kinase inhibitors on the growth, morphology, and infectivity of Leishmania promastigotes. Parasitol. Res. 1997, 83, 273–280. [Google Scholar] [CrossRef] [PubMed]
- Gazarini, M.L.; Garcia, C.R. Interruption of the blood-stage cycle of the malaria parasite, Plasmodium chabaudi, by protein tyrosine kinase inhibitors. Braz. J. Med. Biol. Res. 2003, 36, 1465–1469. [Google Scholar] [CrossRef] [PubMed]
- Park, B.S.; Abdel-Azeem, A.Z.; Al-Sanea, M.M.; Yoo, K.H.; Tae, J.S.; Lee, S.H. Staurosporine analogues from microbial and synthetic sources and their biological activities. Curr. Med. Chem. 2013, 20, 3872–3902. [Google Scholar] [CrossRef] [PubMed]
- Ju, J.W.; Joo, H.N.; Lee, M.R.; Cho, S.H.; Cheun, H.I.; Kim, J.Y.; Lee, Y.H.; Lee, K.J.; Sohn, W.M.; Kim, D.M.; et al. Identification of a serodiagnostic antigen, legumain, by immunoproteomic analysis of excretory-secretory products of Clonorchis sinensis adult worms. Proteomics 2009, 9, 3066–3078. [Google Scholar] [CrossRef] [PubMed]
- Mehlhorn, H. Encyclopedic Reference of Parasitology, 2nd ed.; Springer: New York, NY, USA, 2001. [Google Scholar]
- Cho, P.Y.; Kim, T.I.; Whang, S.M.; Hong, S.J. Gene expression profile of Clonorchis sinensis metacercariae. Parasitol. Res. 2008, 102, 277–282. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhao, J.; Wang, X.; Xu, X.; Pan, W. Evaluation of six novel antigens as potential biomarkers for the early immunodiagnosis of schistosomiasis. Parasites Vectors 2015, 8, 447. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.W.; Yoo, W.G.; Lee, S.; Lee, M.R.; Kim, Y.J.; Cho, S.H.; Lee, W.J.; Ju, J.W. ClonorESTdb: A comprehensive database for Clonorchis sinensis EST sequences. BMC Res. Notes 2014, 7, 388. [Google Scholar] [CrossRef] [PubMed]
- Min, X.J.; Butler, G.; Storms, R.; Tsang, A. OrfPredictor: Predicting protein-coding regions in EST-derived sequences. Nucleic Acids Res. 2005, 33, W677–W680. [Google Scholar] [CrossRef] [PubMed]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Humana Press: New York, NY, USA, 2005; pp. 571–607. [Google Scholar]
- Quevillon, E.; Silventoinen, V.; Pillai, S.; Harte, N.; Mulder, N.; Apweiler, R.; Lopez, R. InterProScan: Protein domains identifier. Nucleic Acids Res. 2005, 33, W116–W120. [Google Scholar] [CrossRef] [PubMed]
- Bendtsen, J.D.; Nielsen, H.; von Heijne, G.; Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 2004, 340, 783–795. [Google Scholar] [CrossRef] [PubMed]
- Emanuelsson, O.; Nielsen, H.; Brunak, S.; von Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J. Mol. Biol. 2000, 300, 1005–1016. [Google Scholar] [CrossRef] [PubMed]
- Krogh, A.; Larsson, B.; von Heijne, G.; Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 2001, 305, 567–580. [Google Scholar] [CrossRef] [PubMed]
- Pruitt, K.D.; Tatusova, T.; Brown, G.R.; Maglott, D.R. NCBI Reference Sequences (RefSeq): Current status, new features and genome annotation policy. Nucleic Acids Res. 2012, 40, D130–D135. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T.; Cozzetto, D. DISOPRED3: Precise disordered region predictions with annotated protein-binding activity. Bioinformatics 2015, 31, 857–863. [Google Scholar] [CrossRef] [PubMed]
- Lovell, S.C.; Davis, I.W.; Arendall, W.B., 3rd; de Bakker, P.I.; Word, J.M.; Prisant, M.G.; Richardson, J.S.; Richardson, D.C. Structure validation by Cα geometry: PHI, PSI and Cβ deviation. Proteins 2003, 50, 437–450. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; Watson, J.D.; Thornton, J.M. ProFunc: A server for predicting protein function from 3D structure. Nucleic Acids Res. 2005, 33, W89–W93. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.C.; Meng, E.C.; Morris, J.H.; Pettersen, E.F.; Ferrin, T.E. Enhancing UCSF Chimera through web services. Nucleic Acids Res. 2014, 42, W478–W484. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; Swindells, M.B. LigPlot+: Multiple ligand-protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 2011, 51, 2778–2786. [Google Scholar] [CrossRef] [PubMed]
- Labbe, C.M.; Rey, J.; Lagorce, D.; Vavrusa, M.; Becot, J.; Sperandio, O.; Villoutreix, B.O.; Tuffery, P.; Miteva, M.A. MTiOpenScreen: A web server for structure-based virtual screening. Nucleic Acids Res. 2015, 43, W448–W454. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.Y.; Kang, S.Y.; Ahn, I.Y.; Cho, S.Y.; Hong, S.J. Molecular cloning and characterization of an antigenic protein with a repeating region from Clonorchis sinensis. Korean J. Parasitol. 2001, 39, 57–66. [Google Scholar] [CrossRef] [PubMed]










| PDB ID | Aln. Score a | Aln. Length b | RMSD (Å) c | Avg. dL d | Genom. Z e | Target Molecule |
|---|---|---|---|---|---|---|
| 1YO3_A | 67.31 | 78 | 1.95 | 1.20 | −5.46 | Dynein light chain 1 |
| 1RE6_A | 66.35 | 78 | 1.98 | 0.20 | −7.05 | Dynein light chain 2 |
| 1JFJ_A | 49.94 | 65 | 3.12 | 2.00 | −3.16 | Calcium-binding protein |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, Y.-J.; Yoo, W.G.; Lee, M.-R.; Kang, J.-M.; Na, B.-K.; Cho, S.-H.; Park, M.-Y.; Ju, J.-W. Molecular and Structural Characterization of the Tegumental 20.6-kDa Protein in Clonorchis sinensis as a Potential Druggable Target. Int. J. Mol. Sci. 2017, 18, 557. https://doi.org/10.3390/ijms18030557
Kim Y-J, Yoo WG, Lee M-R, Kang J-M, Na B-K, Cho S-H, Park M-Y, Ju J-W. Molecular and Structural Characterization of the Tegumental 20.6-kDa Protein in Clonorchis sinensis as a Potential Druggable Target. International Journal of Molecular Sciences. 2017; 18(3):557. https://doi.org/10.3390/ijms18030557
Chicago/Turabian StyleKim, Yu-Jung, Won Gi Yoo, Myoung-Ro Lee, Jung-Mi Kang, Byoung-Kuk Na, Shin-Hyeong Cho, Mi-Yeoun Park, and Jung-Won Ju. 2017. "Molecular and Structural Characterization of the Tegumental 20.6-kDa Protein in Clonorchis sinensis as a Potential Druggable Target" International Journal of Molecular Sciences 18, no. 3: 557. https://doi.org/10.3390/ijms18030557