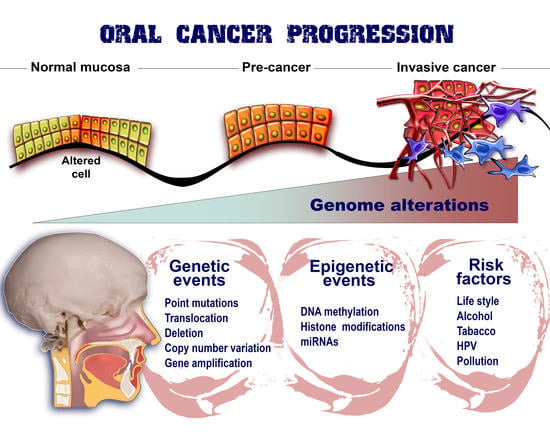
Current Insights into Oral Cancer Epigenetics
Abstract
:1. Introduction
2. Epigenetics—An Emerging Concept
3. DNA Methylation
4. Histone Modifications
5. Nucleosome Positioning and Histone Variants
6. Non-Coding RNAs—miRNAs
7. Epigenetic Landscape in Oral Cancer
8. DNA Methylation in Oral Cancer
9. Histone Modifications in Oral Cancer
10. Epigenetic Alterations of miRNAs in Oral Cancer
11. Epigenetic Therapies in Oral Cancer
12. Conclusions
Acknowledgments
Conflicts of Interest
References
- Irimie, A.I.; Braicu, C.; Cojocneanu-Petric, R.; Berindan-Neagoe, I.; Campian, R.S. Novel technologies for oral squamous carcinoma biomarkers in diagnostics and prognostics. Acta Odontol. Scand. 2015, 73, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Hema, K.N.; Smitha, T.; Sheethal, H.S.; Mirnalini, S.A. Epigenetics in oral squamous cell carcinoma. J. Oral Maxillofac. Pathol. 2017, 21, 252–259. [Google Scholar] [CrossRef] [PubMed]
- Parkin, D.M.; Bray, F.; Ferlay, J.; Pisani, P. Global cancer statistics, 2002. CA Cancer J. Clin. 2005, 55, 74–108. [Google Scholar] [CrossRef] [PubMed]
- Stewart, B.W.; Greim, H.; Shuker, D.; Kauppinen, T. Defence of iarc monographs. Lancet 2003, 361, 1300. [Google Scholar] [CrossRef]
- Irimie, A.I.; Braicu, C.; Zanoaga, O.; Pileczki, V.; Gherman, C.; Berindan-Neagoe, I.; Campian, R.S. Epigallocatechin-3-gallate suppresses cell proliferation and promotes apoptosis and autophagy in oral cancer ssc-4 cells. Onco Targets Ther. 2015, 8, 461–470. [Google Scholar] [PubMed]
- Perez-Sayans, M.; Somoza-Martin, J.M.; Barros-Angueira, F.; Reboiras-Lopez, M.D.; Gandara Rey, J.M.; Garcia-Garcia, A. Genetic and molecular alterations associated with oral squamous cell cancer (review). Oncol. Rep. 2009, 22, 1277–1282. [Google Scholar] [CrossRef] [PubMed]
- Irimie, A.I.; Braicu, C.; Pileczki, V.; Petrushev, B.; Soritau, O.; Campian, R.S.; Berindan-Neagoe, I. Knocking down of p53 triggers apoptosis and autophagy, concomitantly with inhibition of migration on ssc-4 oral squamous carcinoma cells. Mol. Cell. Biochem. 2016, 419, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Lingen, M.W.; Pinto, A.; Mendes, R.A.; Franchini, R.; Czerninski, R.; Tilakaratne, W.M.; Partridge, M.; Peterson, D.E.; Woo, S.B. Genetics/epigenetics of oral premalignancy: Current status and future research. Oral Dis. 2011, 17 (Suppl. 1), 7–22. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Kelly, T.K.; Jones, P.A. Epigenetics in cancer. Carcinogenesis 2010, 31, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Heyn, H.; Esteller, M. DNA methylation profiling in the clinic: Applications and challenges. Nat. Rev. Genet. 2012, 13, 679–692. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.A.; Baylin, S.B. The fundamental role of epigenetic events in cancer. Nat. Rev. Genet. 2002, 3, 415–428. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.A.; Baylin, S.B. The epigenomics of cancer. Cell 2007, 128, 683–692. [Google Scholar] [CrossRef] [PubMed]
- Yoo, C.B.; Jones, P.A. Epigenetic therapy of cancer: Past, present and future. Nat. Rev. Drug Discov. 2006, 5, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Kornberg, R.D. Chromatin structure: A repeating unit of histones and DNA. Science 1974, 184, 868–871. [Google Scholar] [CrossRef] [PubMed]
- Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002, 16, 6–21. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Leung, F.C. An evaluation of new criteria for cpg islands in the human genome as gene markers. Bioinformatics 2004, 20, 1170–1177. [Google Scholar] [CrossRef] [PubMed]
- Illingworth, R.; Kerr, A.; Desousa, D.; Jorgensen, H.; Ellis, P.; Stalker, J.; Jackson, D.; Clee, C.; Plumb, R.; Rogers, J.; et al. A novel cpg island set identifies tissue-specific methylation at developmental gene loci. PLoS Biol. 2008, 6, e22. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.M.; Bird, A. DNA methylation landscapes: Provocative insights from epigenomics. Nat. Rev. Genet. 2008, 9, 465–476. [Google Scholar] [CrossRef] [PubMed]
- Eckhardt, F.; Lewin, J.; Cortese, R.; Rakyan, V.K.; Attwood, J.; Burger, M.; Burton, J.; Cox, T.V.; Davies, R.; Down, T.A.; et al. DNA methylation profiling of human chromosomes 6, 20 and 22. Nat. Genet. 2006, 38, 1378–1385. [Google Scholar] [CrossRef] [PubMed]
- Baylin, S.B.; Jones, P.A. A decade of exploring the cancer epigenome-biological and translational implications. Nat. Rev. Cancer 2011, 11, 726–734. [Google Scholar] [CrossRef] [PubMed]
- Dawson, M.A.; Kouzarides, T. Cancer epigenetics: From mechanism to therapy. Cell 2012, 150, 12–27. [Google Scholar] [CrossRef] [PubMed]
- Robertson, K.D. DNA methylation and human disease. Nat. Rev. Genet. 2005, 6, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Ley, T.J.; Ding, L.; Walter, M.J.; McLellan, M.D.; Lamprecht, T.; Larson, D.E.; Kandoth, C.; Payton, J.E.; Baty, J.; Welch, J.; et al. Dnmt3a mutations in acute myeloid leukemia. N. Engl. J. Med. 2010, 363, 2424–2433. [Google Scholar] [CrossRef] [PubMed]
- Kouzarides, T. Chromatin modifications and their function. Cell 2007, 128, 693–705. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, B.E.; Meissner, A.; Lander, E.S. The mammalian epigenome. Cell 2007, 128, 669–681. [Google Scholar] [CrossRef] [PubMed]
- Haberland, M.; Montgomery, R.L.; Olson, E.N. The many roles of histone deacetylases in development and physiology: Implications for disease and therapy. Nat. Rev. Genet. 2009, 10, 32–42. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y. Histone lysine demethylases: Emerging roles in development, physiology and disease. Nat. Rev. Genet. 2007, 8, 829–833. [Google Scholar] [CrossRef] [PubMed]
- Bannister, A. The Role of Epigenetics in Cancer. Available online: http://www.abcam.com/index.html?pageconfig=resource&rid=10755 (accessed on 27 February 2018).
- Arif, M.; Vedamurthy, B.M.; Choudhari, R.; Ostwal, Y.B.; Mantelingu, K.; Kodaganur, G.S.; Kundu, T.K. Nitric oxide-mediated histone hyperacetylation in oral cancer: Target for a water-soluble hat inhibitor, ctk7a. Chem. Biol. 2010, 17, 903–913. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Pugh, B.F. Nucleosome positioning and gene regulation: Advances through genomics. Nat. Rev. Genet. 2009, 10, 161–172. [Google Scholar] [CrossRef] [PubMed]
- Yuan, G.C.; Liu, Y.J.; Dion, M.F.; Slack, M.D.; Wu, L.F.; Altschuler, S.J.; Rando, O.J. Genome-scale identification of nucleosome positions in s. Cerevisiae. Science 2005, 309, 626–630. [Google Scholar] [CrossRef] [PubMed]
- Shivaswamy, S.; Bhinge, A.; Zhao, Y.; Jones, S.; Hirst, M.; Iyer, V.R. Dynamic remodeling of individual nucleosomes across a eukaryotic genome in response to transcriptional perturbation. PLoS Biol. 2008, 6, e65. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.C.; Jeong, S.; Liang, G.; Takai, D.; Fatemi, M.; Tsai, Y.C.; Egger, G.; Gal-Yam, E.N.; Jones, P.A. Role of nucleosomal occupancy in the epigenetic silencing of the mlh1 cpg island. Cancer Cell 2007, 12, 432–444. [Google Scholar] [CrossRef] [PubMed]
- Gal-Yam, E.N.; Jeong, S.; Tanay, A.; Egger, G.; Lee, A.S.; Jones, P.A. Constitutive nucleosome depletion and ordered factor assembly at the grp78 promoter revealed by single molecule footprinting. PLoS Genet. 2006, 2, e160. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.L.; Peterson, C.L. Atp-dependent chromatin remodeling. Curr. Top. Dev. Biol. 2005, 65, 115–148. [Google Scholar] [PubMed]
- Santenard, A.; Torres-Padilla, M.E. Epigenetic reprogramming in mammalian reproduction: Contribution from histone variants. Epigenetics 2009, 4, 80–84. [Google Scholar] [CrossRef] [PubMed]
- Sarma, K.; Reinberg, D. Histone variants meet their match. Nat. Rev. Mol. Cell Biol. 2005, 6, 139–149. [Google Scholar] [CrossRef] [PubMed]
- Budisan, L.; Gulei, D.; Zanoaga, O.M.; Irimie, A.I.; Sergiu, C.; Braicu, C.; Gherman, C.D.; Berindan-Neagoe, I. Dietary intervention by phytochemicals and their role in modulating coding and non-coding genes in cancer. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Gulei, D.; Magdo, L.; Jurj, A.; Raduly, L.; Cojocneanu-Petric, R.; Moldovan, A.; Moldovan, C.; Florea, A.; Pasca, S.; Pop, L.A.; et al. The silent healer: Mir-205-5p up-regulation inhibits epithelial to mesenchymal transition in colon cancer cells by indirectly up-regulating e-cadherin expression. Cell Death Dis. 2018, 9, 66. [Google Scholar] [CrossRef] [PubMed]
- Gulei, D.; Mehterov, N.; Nabavi, S.M.; Atanasov, A.G.; Berindan-Neagoe, I. Targeting ncrnas by plant secondary metabolites: The ncrnas game in the balance towards malignancy inhibition. Biotechnol. Adv. 2017. [Google Scholar] [CrossRef] [PubMed]
- Pop-Bica, C.; Gulei, D.; Cojocneanu-Petric, R.; Braicu, C.; Petrut, B.; Berindan-Neagoe, I. Understanding the role of non-coding rnas in bladder cancer: From dark matter to valuable therapeutic targets. Int. J. Mol. Sci. 2017, 18. [Google Scholar] [CrossRef] [PubMed]
- Berindan-Neagoe, I.; Calin, G.A. Molecular pathways: Micrornas, cancer cells, and microenvironment. Clin. Cancer Res. 2014, 20, 6247–6253. [Google Scholar] [CrossRef] [PubMed]
- Braicu, C.; Calin, G.A.; Berindan-Neagoe, I. Micrornas and cancer therapy—From bystanders to major players. Curr. Med. Chem. 2013, 20, 3561–3573. [Google Scholar] [CrossRef] [PubMed]
- Berindan-Neagoe, I.; Monroig Pdel, C.; Pasculli, B.; Calin, G.A. Micrornaome genome: A treasure for cancer diagnosis and therapy. CA Cancer J. Clin. 2014, 64, 311–336. [Google Scholar] [CrossRef] [PubMed]
- Zaharie, F.; Muresan, M.S.; Petrushev, B.; Berce, C.; Gafencu, G.A.; Selicean, S.; Jurj, A.; Cojocneanu-Petric, R.; Lisencu, C.I.; Pop, L.A.; et al. Exosome-carried microrna-375 inhibits cell progression and dissemination via bcl-2 blocking in colon cancer. J. Gastrointest. Liver Dis. 2015, 24, 435–443. [Google Scholar]
- Fabbri, M.; Garzon, R.; Cimmino, A.; Liu, Z.; Zanesi, N.; Callegari, E.; Liu, S.; Alder, H.; Costinean, S.; Fernandez-Cymering, C.; et al. Microrna-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3a and 3b. Proc. Natl. Acad. Sci. USA 2007, 104, 15805–15810. [Google Scholar] [CrossRef] [PubMed]
- Friedman, J.M.; Liang, G.; Liu, C.C.; Wolff, E.M.; Tsai, Y.C.; Ye, W.; Zhou, X.; Jones, P.A. The putative tumor suppressor microrna-101 modulates the cancer epigenome by repressing the polycomb group protein ezh2. Cancer Res. 2009, 69, 2623–2629. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, M.; Calin, G.A. Epigenetics and mirnas in human cancer. Adv. Genet. 2010, 70, 87–99. [Google Scholar] [PubMed]
- Kala, R.; Peek, G.W.; Hardy, T.M.; Tollefsbol, T.O. Micrornas: An emerging science in cancer epigenetics. J. Clin. Bioinform. 2013, 3, 6. [Google Scholar] [CrossRef] [PubMed]
- Kozaki, K.; Imoto, I.; Mogi, S.; Omura, K.; Inazawa, J. Exploration of tumor-suppressive micrornas silenced by DNA hypermethylation in oral cancer. Cancer Res. 2008, 68, 2094–2105. [Google Scholar] [CrossRef] [PubMed]
- Gomes, C.C.; Gomez, R.S. Microrna and oral cancer: Future perspectives. Oral Oncol. 2008, 44, 910–914. [Google Scholar] [CrossRef] [PubMed]
- Gasche, J.A.; Goel, A. Epigenetic mechanisms in oral carcinogenesis. Future Oncol. 2012, 8, 1407–1425. [Google Scholar] [CrossRef] [PubMed]
- Mascolo, M.; Siano, M.; Ilardi, G.; Russo, D.; Merolla, F.; De Rosa, G.; Staibano, S. Epigenetic disregulation in oral cancer. Int. J. Mol. Sci. 2012, 13, 2331–2353. [Google Scholar] [CrossRef] [PubMed]
- Jithesh, P.V.; Risk, J.M.; Schache, A.G.; Dhanda, J.; Lane, B.; Liloglou, T.; Shaw, R.J. The epigenetic landscape of oral squamous cell carcinoma. Br. J. Cancer 2013, 108, 370–379. [Google Scholar] [CrossRef] [PubMed]
- Piyathilake, C.J.; Bell, W.C.; Jones, J.; Henao, O.L.; Heimburger, D.C.; Niveleau, A.; Grizzle, W.E. Pattern of nonspecific (or global) DNA methylation in oral carcinogenesis. Head Neck 2005, 27, 1061–1067. [Google Scholar] [CrossRef] [PubMed]
- Guerrero-Preston, R.; Baez, A.; Blanco, A.; Berdasco, M.; Fraga, M.; Esteller, M. Global DNA methylation: A common early event in oral cancer cases with exposure to environmental carcinogens or viral agents. P. R. Health Sci. J. 2009, 28, 24–29. [Google Scholar] [PubMed]
- Baba, S.; Yamada, Y.; Hatano, Y.; Miyazaki, Y.; Mori, H.; Shibata, T.; Hara, A. Global DNA hypomethylation suppresses squamous carcinogenesis in the tongue and esophagus. Cancer Sci. 2009, 100, 1186–1191. [Google Scholar] [CrossRef] [PubMed]
- Supic, G.; Kozomara, R.; Jovic, N.; Zeljic, K.; Magic, Z. Prognostic significance of tumor-related genes hypermethylation detected in cancer-free surgical margins of oral squamous cell carcinomas. Oral Oncol. 2011, 47, 702–708. [Google Scholar] [CrossRef] [PubMed]
- Aoki, K.; Taketo, M.M. Adenomatous polyposis coli (apc): A multi-functional tumor suppressor gene. J. Cell. Sci. 2007, 120, 3327–3335. [Google Scholar] [CrossRef] [PubMed]
- Reya, T.; Clevers, H. Wnt signalling in stem cells and cancer. Nature 2005, 434, 843–850. [Google Scholar] [CrossRef] [PubMed]
- Barker, N.; Clevers, H. Mining the wnt pathway for cancer therapeutics. Nat. Rev. Drug Discov. 2006, 5, 997–1014. [Google Scholar] [CrossRef] [PubMed]
- Iwai, S.; Katagiri, W.; Kong, C.; Amekawa, S.; Nakazawa, M.; Yura, Y. Mutations of the apc, beta-catenin, and axin 1 genes and cytoplasmic accumulation of beta-catenin in oral squamous cell carcinoma. J. Cancer Res. Clin. Oncol. 2005, 131, 773–782. [Google Scholar] [CrossRef] [PubMed]
- Rigi-Ladiz, M.A.; Kordi-Tamandani, D.M.; Torkamanzehi, A. Analysis of hypermethylation and expression profiles of apc and atm genes in patients with oral squamous cell carcinoma. Clin. Epigenet. 2011, 3, 6. [Google Scholar] [CrossRef] [PubMed]
- Uesugi, H.; Uzawa, K.; Kawasaki, K.; Shimada, K.; Moriya, T.; Tada, A.; Shiiba, M.; Tanzawa, H. Status of reduced expression and hypermethylation of the apc tumor suppressor gene in human oral squamous cell carcinoma. Int. J. Mol. Med. 2005, 15, 597–602. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Eiberg, H.; Krogdahl, A.; Liu, C.J.; Sorensen, J.A. Cytoplasmic expression of e-cadherin and beta-catenin correlated with loh and hypermethylation of the apc gene in oral squamous cell carcinomas. J. Oral Pathol. Med. 2005, 34, 116–119. [Google Scholar] [CrossRef] [PubMed]
- Ali, J.; Sabiha, B.; Jan, H.U.; Haider, S.A.; Khan, A.A.; Ali, S.S. Genetic etiology of oral cancer. Oral Oncol. 2017, 70, 23–28. [Google Scholar] [CrossRef] [PubMed]
- Lo Muzio, L.; Pannone, G.; Staibano, S.; Mignogna, M.D.; Rubini, C.; Mariggio, M.A.; Procaccini, M.; Ferrari, F.; De Rosa, G.; Altieri, D.C. Survivin expression in oral squamous cell carcinoma. Br. J. Cancer 2003, 89, 2244–2248. [Google Scholar] [CrossRef] [PubMed]
- Khan, Z.; Tiwari, R.P.; Mulherkar, R.; Sah, N.K.; Prasad, G.B.; Shrivastava, B.R.; Bisen, P.S. Detection of survivin and p53 in human oral cancer: Correlation with clinicopathologic findings. Head Neck 2009, 31, 1039–1048. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, P.K.; Goel, A.; Mittal, R.D. Survivin: A molecular biomarker in cancer. Indian J. Med. Res. 2015, 141, 389–397. [Google Scholar] [PubMed]
- Chen, Y.K.; Hsue, S.S.; Lin, L.M. Survivin expression is regulated by an epigenetic mechanism for dmba-induced hamster buccal-pouch squamous-cell carcinomas. Arch. Oral Biol. 2005, 50, 593–598. [Google Scholar] [CrossRef] [PubMed]
- Hsue, S.S.; Wang, W.C.; Chen, Y.K.; Lin, L.M. Expression of inhibitors of apoptosis family protein in 7,12-dimethylbenz[a]anthracene-induced hamster buccal-pouch squamous-cell carcinogenesis is associated with mutant p53 accumulation and epigenetic changes. Int. J. Exp. Pathol. 2008, 89, 309–320. [Google Scholar] [CrossRef] [PubMed]
- Gulei, D.; Mehterov, N.; Ling, H.; Stanta, G.; Braicu, C.; Berindan-Neagoe, I. The “good-cop bad-cop” tgf-beta role in breast cancer modulated by non-coding rnas. Biochim. Biophys. Acta 2017, 1861, 1661–1675. [Google Scholar] [CrossRef] [PubMed]
- Viswanathan, M.; Tsuchida, N.; Shanmugam, G. Promoter hypermethylation profile of tumor-associated genes p16, p15, hmlh1, mgmt and e-cadherin in oral squamous cell carcinoma. Int. J. Cancer 2003, 105, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Maruya, S.; Issa, J.P.; Weber, R.S.; Rosenthal, D.I.; Haviland, J.C.; Lotan, R.; El-Naggar, A.K. Differential methylation status of tumor-associated genes in head and neck squamous carcinoma: Incidence and potential implications. Clin. Cancer Res. 2004, 10, 3825–3830. [Google Scholar] [CrossRef] [PubMed]
- Saito, Y.; Takazawa, H.; Uzawa, K.; Tanzawa, H.; Sato, K. Reduced expression of e-cadherin in oral squamous cell carcinoma: Relationship with DNA methylation of 5′ cpg island. Int. J. Oncol. 1998, 12, 293–298. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, S.; Sasaki, A.; Mese, H.; Alcalde, R.E.; Tsuji, T.; Matsumura, T. The e-cadherin gene is silenced by cpg methylation in human oral squamous cell carcinomas. Int. J. Cancer 2001, 93, 667–673. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.W.; Chow, V.; Lam, K.Y.; Wei, W.I.; Yuen, A. Loss of e-cadherin expression resulting from promoter hypermethylation in oral tongue carcinoma and its prognostic significance. Cancer 2002, 94, 386–392. [Google Scholar] [CrossRef] [PubMed]
- Vered, M.; Allon, I.; Buchner, A.; Dayan, D. E-cadherin in oral scc: An analysis of the confusing literature and new insights related to its immunohistochemical expression. Histol. Histopathol. 2012, 27, 141–150. [Google Scholar] [PubMed]
- Yin, Y.; Shen, W.H. Pten: A new guardian of the genome. Oncogene 2008, 27, 5443–5453. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.J.; Liu, V.W.; Wang, Y.; Tsang, P.C.; Ngan, H.Y. Differential DNA methylation profiles in gynecological cancers and correlation with clinico-pathological data. BMC Cancer 2006, 6, 212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kang, Y.H.; Lee, H.S.; Kim, W.H. Promoter methylation and silencing of pten in gastric carcinoma. Lab. Investig. 2002, 82, 285–291. [Google Scholar] [CrossRef] [PubMed]
- Salvesen, H.B.; MacDonald, N.; Ryan, A.; Jacobs, I.J.; Lynch, E.D.; Akslen, L.A.; Das, S. Pten methylation is associated with advanced stage and microsatellite instability in endometrial carcinoma. Int. J. Cancer 2001, 91, 22–26. [Google Scholar] [CrossRef]
- Soria, J.C.; Lee, H.Y.; Lee, J.I.; Wang, L.; Issa, J.P.; Kemp, B.L.; Liu, D.D.; Kurie, J.M.; Mao, L.; Khuri, F.R. Lack of pten expression in non-small cell lung cancer could be related to promoter methylation. Clin. Cancer Res. 2002, 8, 1178–1184. [Google Scholar] [PubMed]
- Kurasawa, Y.; Shiiba, M.; Nakamura, M.; Fushimi, K.; Ishigami, T.; Bukawa, H.; Yokoe, H.; Uzawa, K.; Tanzawa, H. Pten expression and methylation status in oral squamous cell carcinoma. Oncol. Rep. 2008, 19, 1429–1434. [Google Scholar] [PubMed]
- Lee, J.I.; Soria, J.C.; Hassan, K.A.; El-Naggar, A.K.; Tang, X.; Liu, D.D.; Hong, W.K.; Mao, L. Loss of pten expression as a prognostic marker for tongue cancer. Arch. Otolaryngol. Head Neck Surg. 2001, 127, 1441–1445. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Samaranayake, L.P.; Zhou, H.; Xiao, L. Homozygous deletion of the pten tumor-suppressor gene is not a feature in oral squamous cell carcinoma. Oral Oncol. 2000, 36, 95–99. [Google Scholar] [CrossRef]
- Mavros, A.; Hahn, M.; Wieland, I.; Koy, S.; Koufaki, O.N.; Strelocke, K.; Koch, R.; Haroske, G.; Schackert, H.K.; Eckelt, U. Infrequent genetic alterations of the tumor suppressor gene pten/mmac1 in squamous cell carcinoma of the oral cavity. J. Oral Pathol. Med. 2002, 31, 270–276. [Google Scholar] [CrossRef] [PubMed]
- Squarize, C.H.; Castilho, R.M.; Santos Pinto, D., Jr. Immunohistochemical evidence of pten in oral squamous cell carcinoma and its correlation with the histological malignancy grading system. J. Oral Pathol. Med. 2002, 31, 379–384. [Google Scholar] [CrossRef] [PubMed]
- Shin, K.H.; Kim, J.M.; Rho, K.S.; Park, K.H.; Oh, J.E.; Min, B.M. Inactivation of the pten gene by mutation, exonic deletion, and loss of transcript in human oral squamous cell carcinomas. Int. J. Oncol. 2002, 21, 997–1001. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, V.; Saranath, D. Concurrent hypermethylation of multiple regulatory genes in chewing tobacco associated oral squamous cell carcinomas and adjacent normal tissues. Oral Oncol. 2004, 40, 145–153. [Google Scholar] [CrossRef]
- Huang, M.J.; Yeh, K.T.; Shih, H.C.; Wang, Y.F.; Lin, T.H.; Chang, J.Y.; Shih, M.C.; Chang, J.G. The correlation between cpg methylation and protein expression of p16 in oral squamous cell carcinomas. Int. J. Mol. Med. 2002, 10, 551–554. [Google Scholar] [PubMed]
- Kato, K.; Hara, A.; Kuno, T.; Mori, H.; Yamashita, T.; Toida, M.; Shibata, T. Aberrant promoter hypermethylation of p16 and mgmt genes in oral squamous cell carcinomas and the surrounding normal mucosa. J. Cancer Res. Clin. Oncol. 2006, 132, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Hasegawa, M.; Nelson, H.H.; Peters, E.; Ringstrom, E.; Posner, M.; Kelsey, K.T. Patterns of gene promoter methylation in squamous cell cancer of the head and neck. Oncogene 2002, 21, 4231–4236. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Cespedes, M.; Esteller, M.; Wu, L.; Nawroz-Danish, H.; Yoo, G.H.; Koch, W.M.; Jen, J.; Herman, J.G.; Sidransky, D. Gene promoter hypermethylation in tumors and serum of head and neck cancer patients. Cancer Res. 2000, 60, 892–895. [Google Scholar] [PubMed]
- Rosas, S.L.; Koch, W.; da Costa Carvalho, M.G.; Wu, L.; Califano, J.; Westra, W.; Jen, J.; Sidransky, D. Promoter hypermethylation patterns of p16, o6-methylguanine-DNA-methyltransferase, and death-associated protein kinase in tumors and saliva of head and neck cancer patients. Cancer Res. 2001, 61, 939–942. [Google Scholar] [PubMed]
- Viet, C.T.; Jordan, R.C.; Schmidt, B.L. DNA promoter hypermethylation in saliva for the early diagnosis of oral cancer. J. Calif. Dent. Assoc. 2007, 35, 844–849. [Google Scholar] [PubMed]
- Youssef, E.M.; Lotan, D.; Issa, J.P.; Wakasa, K.; Fan, Y.H.; Mao, L.; Hassan, K.; Feng, L.; Lee, J.J.; Lippman, S.M.; et al. Hypermethylation of the retinoic acid receptor-beta(2) gene in head and neck carcinogenesis. Clin. Cancer Res. 2004, 10, 1733–1742. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.S.; Issa, J.P.; Roberts, D.B.; Williams, M.D.; Weber, R.S.; Kies, M.S.; El-Naggar, A.K. Quantitative promoter hypermethylation analysis of cancer-related genes in salivary gland carcinomas: Comparison with methylation-specific pcr technique and clinical significance. Clin. Cancer Res. 2008, 14, 2664–2672. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.H.; Huang, S.F.; Chen, I.H.; Liao, C.T.; Wang, H.M.; Hsieh, L.L. Methylation of rassf1a, rassf2a, and hin-1 is associated with poor outcome after radiotherapy, but not surgery, in oral squamous cell carcinoma. Clin. Cancer Res. 2009, 15, 4174–4180. [Google Scholar] [CrossRef] [PubMed]
- Imai, T.; Toyota, M.; Suzuki, H.; Akino, K.; Ogi, K.; Sogabe, Y.; Kashima, L.; Maruyama, R.; Nojima, M.; Mita, H.; et al. Epigenetic inactivation of rassf2 in oral squamous cell carcinoma. Cancer Sci. 2008, 99, 958–966. [Google Scholar] [CrossRef] [PubMed]
- Lo Muzio, L.; Staibano, S.; Pannone, G.; Mignogna, M.D.; Mariggio, A.; Salvatore, G.; Chieffi, P.; Tramontano, D.; De Rosa, G.; Altieri, D.C. Expression of the apoptosis inhibitor survivin in aggressive squamous cell carcinoma. Exp. Mol. Pathol. 2001, 70, 249–254. [Google Scholar] [CrossRef] [PubMed]
- Czerninski, R.; Krichevsky, S.; Ashhab, Y.; Gazit, D.; Patel, V.; Ben-Yehuda, D. Promoter hypermethylation of mismatch repair genes, hmlh1 and hmsh2 in oral squamous cell carcinoma. Oral Dis. 2009, 15, 206–213. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Ramirez, I.; Ramirez-Amador, V.; Irigoyen-Camacho, M.E.; Sanchez-Perez, Y.; Anaya-Saavedra, G.; Granados-Garcia, M.; Garcia-Vazquez, F.; Garcia-Cuellar, C.M. Hmlh1 promoter methylation is an early event in oral cancer. Oral Oncol. 2011, 47, 22–26. [Google Scholar] [CrossRef] [PubMed]
- Takeshima, M.; Saitoh, M.; Kusano, K.; Nagayasu, H.; Kurashige, Y.; Malsantha, M.; Arakawa, T.; Takuma, T.; Chiba, I.; Kaku, T.; et al. High frequency of hypermethylation of p14, p15 and p16 in oral pre-cancerous lesions associated with betel-quid chewing in sri lanka. J. Oral Pathol. Med. 2008, 37, 475–479. [Google Scholar] [CrossRef] [PubMed]
- Ishida, E.; Nakamura, M.; Ikuta, M.; Shimada, K.; Matsuyoshi, S.; Kirita, T.; Konishi, N. Promotor hypermethylation of p14arf is a key alteration for progression of oral squamous cell carcinoma. Oral Oncol. 2005, 41, 614–622. [Google Scholar] [CrossRef] [PubMed]
- Sailasree, R.; Abhilash, A.; Sathyan, K.M.; Nalinakumari, K.R.; Thomas, S.; Kannan, S. Differential roles of p16ink4a and p14arf genes in prognosis of oral carcinoma. Cancer Epidemiol. Biomark. Prev. 2008, 17, 414–420. [Google Scholar] [CrossRef] [PubMed]
- Yeh, K.T.; Chang, J.G.; Lin, T.H.; Wang, Y.F.; Tien, N.; Chang, J.Y.; Chen, J.C.; Shih, M.C. Epigenetic changes of tumor suppressor genes, p15, p16, vhl and p53 in oral cancer. Oncol. Rep. 2003, 10, 659–663. [Google Scholar] [PubMed]
- Shaw, R.J.; Liloglou, T.; Rogers, S.N.; Brown, J.S.; Vaughan, E.D.; Lowe, D.; Field, J.K.; Risk, J.M. Promoter methylation of p16, rarbeta, e-cadherin, cyclin a1 and cytoglobin in oral cancer: Quantitative evaluation using pyrosequencing. Br. J. Cancer 2006, 94, 561–568. [Google Scholar] [CrossRef] [PubMed]
- Mancuso, M.; Matassa, D.S.; Conte, M.; Colella, G.; Rana, G.; Fucci, L.; Piscopo, M. H3k4 histone methylation in oral squamous cell carcinoma. Acta Biochim. Pol. 2009, 56, 405–410. [Google Scholar] [PubMed]
- Chen, Y.W.; Kao, S.Y.; Wang, H.J.; Yang, M.H. Histone modification patterns correlate with patient outcome in oral squamous cell carcinoma. Cancer 2013, 119, 4259–4267. [Google Scholar] [CrossRef] [PubMed]
- De Ruijter, A.J.; van Gennip, A.H.; Caron, H.N.; Kemp, S.; van Kuilenburg, A.B. Histone deacetylases (hdacs): Characterization of the classical hdac family. Biochem. J. 2003, 370, 737–749. [Google Scholar] [CrossRef] [PubMed]
- Sakuma, T.; Uzawa, K.; Onda, T.; Shiiba, M.; Yokoe, H.; Shibahara, T.; Tanzawa, H. Aberrant expression of histone deacetylase 6 in oral squamous cell carcinoma. Int. J. Oncol. 2006, 29, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Staibano, S.; Mascolo, M.; Rocco, A.; Lo Muzio, L.; Ilardi, G.; Siano, M.; Pannone, G.; Vecchione, M.L.; Nugnes, L.; Califano, L.; et al. The proliferation marker chromatin assembly factor-1 is of clinical value in predicting the biological behaviour of salivary gland tumours. Oncol. Rep. 2011, 25, 13–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Staibano, S.; Mignogna, C.; Lo Muzio, L.; Mascolo, M.; Salvatore, G.; Di Benedetto, M.; Califano, L.; Rubini, C.; De Rosa, G. Chromatin assembly factor-1 (caf-1)-mediated regulation of cell proliferation and DNA repair: A link with the biological behaviour of squamous cell carcinoma of the tongue? Histopathology 2007, 50, 911–919. [Google Scholar] [CrossRef] [PubMed]
- Sethi, N.; Wright, A.; Wood, H.; Rabbitts, P. Micrornas and head and neck cancer: Reviewing the first decade of research. Eur. J. Cancer 2014, 50, 2619–2635. [Google Scholar] [CrossRef] [PubMed]
- Wiklund, E.D.; Gao, S.; Hulf, T.; Sibbritt, T.; Nair, S.; Costea, D.E.; Villadsen, S.B.; Bakholdt, V.; Bramsen, J.B.; Sorensen, J.A.; et al. Microrna alterations and associated aberrant DNA methylation patterns across multiple sample types in oral squamous cell carcinoma. PLoS ONE 2011, 6, e27840. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hsu, C.M.; Lin, P.M.; Wang, Y.M.; Chen, Z.J.; Lin, S.F.; Yang, M.Y. Circulating mirna is a novel marker for head and neck squamous cell carcinoma. Tumour Biol. 2012, 33, 1933–1942. [Google Scholar] [CrossRef] [PubMed]
- Jung, H.M.; Phillips, B.L.; Patel, R.S.; Cohen, D.M.; Jakymiw, A.; Kong, W.W.; Cheng, J.Q.; Chan, E.K. Keratinization-associated mir-7 and mir-21 regulate tumor suppressor reversion-inducing cysteine-rich protein with kazal motifs (reck) in oral cancer. J. Biol. Chem. 2012, 287, 29261–29272. [Google Scholar] [CrossRef] [PubMed]
- Siow, M.Y.; Ng, L.P.; Vincent-Chong, V.K.; Jamaludin, M.; Abraham, M.T.; Abdul Rahman, Z.A.; Kallarakkal, T.G.; Yang, Y.H.; Cheong, S.C.; Zain, R.B. Dysregulation of mir-31 and mir-375 expression is associated with clinical outcomes in oral carcinoma. Oral Dis. 2014, 20, 345–351. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.J.; Lin, S.C.; Yang, C.C.; Cheng, H.W.; Chang, K.W. Exploiting salivary mir-31 as a clinical biomarker of oral squamous cell carcinoma. Head Neck 2012, 34, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Momen-Heravi, F.; Trachtenberg, A.J.; Kuo, W.P.; Cheng, Y.S. Genomewide study of salivary micrornas for detection of oral cancer. J. Dent. Res. 2014, 93, 86S–93S. [Google Scholar] [CrossRef] [PubMed]
- Shiiba, M.; Shinozuka, K.; Saito, K.; Fushimi, K.; Kasamatsu, A.; Ogawara, K.; Uzawa, K.; Ito, H.; Takiguchi, Y.; Tanzawa, H. Microrna-125b regulates proliferation and radioresistance of oral squamous cell carcinoma. Br. J. Cancer 2013, 108, 1817–1821. [Google Scholar] [CrossRef] [PubMed]
- Ni, Y.H.; Huang, X.F.; Wang, Z.Y.; Han, W.; Deng, R.Z.; Mou, Y.B.; Ding, L.; Hou, Y.Y.; Hu, Q.G. Upregulation of a potential prognostic biomarker, mir-155, enhances cell proliferation in patients with oral squamous cell carcinoma. Oral Surg. Oral Med. Oral Pathol. Oral Radiol. 2014, 117, 227–233. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.J.; Zhang, C.Y.; Zhou, Z.T.; Ma, J.Y.; Liu, Y.; Bao, Z.X.; Jiang, W.W. Microrna-155 in oral squamous cell carcinoma: Overexpression, localization, and prognostic potential. Head Neck 2015, 37, 970–976. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.C.; Hung, P.S.; Wang, P.W.; Liu, C.J.; Chu, T.H.; Cheng, H.W.; Lin, S.C. Mir-181 as a putative biomarker for lymph-node metastasis of oral squamous cell carcinoma. J. Oral Pathol. Med. 2011, 40, 397–404. [Google Scholar] [CrossRef] [PubMed]
- Chang, K.W.; Liu, C.J.; Chu, T.H.; Cheng, H.W.; Hung, P.S.; Hu, W.Y.; Lin, S.C. Association between high mir-211 microrna expression and the poor prognosis of oral carcinoma. J. Dent. Res. 2008, 87, 1063–1068. [Google Scholar] [CrossRef] [PubMed]
- Min, A.; Zhu, C.; Peng, S.; Rajthala, S.; Costea, D.E.; Sapkota, D. Micrornas as important players and biomarkers in oral carcinogenesis. BioMed Res. Int. 2015, 2015, 186904. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suzuki, M.; Shinohara, F.; Nishimura, K.; Echigo, S.; Rikiishi, H. Epigenetic regulation of chemosensitivity to 5-fluorouracil and cisplatin by zebularine in oral squamous cell carcinoma. Int. J. Oncol. 2007, 31, 1449–1456. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.; Shinohara, F.; Endo, M.; Sugazaki, M.; Echigo, S.; Rikiishi, H. Zebularine suppresses the apoptotic potential of 5-fluorouracil via camp/pka/creb pathway against human oral squamous cell carcinoma cells. Cancer Chemother. Pharmacol. 2009, 64, 223–232. [Google Scholar] [CrossRef] [PubMed]
- Kato, K.; Long, N.K.; Makita, H.; Toida, M.; Yamashita, T.; Hatakeyama, D.; Hara, A.; Mori, H.; Shibata, T. Effects of green tea polyphenol on methylation status of reck gene and cancer cell invasion in oral squamous cell carcinoma cells. Br. J. Cancer 2008, 99, 647–654. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Suzuki, M.; Sato, Y.; Echigo, S.; Rikiishi, H. Sequence-dependent interaction between cisplatin and histone deacetylase inhibitors in human oral squamous cell carcinoma cells. Int. J. Oncol. 2006, 28, 1233–1241. [Google Scholar] [CrossRef] [PubMed]
- Manikandan, M.; Deva Magendhra Rao, A.K.; Arunkumar, G.; Manickavasagam, M.; Rajkumar, K.S.; Rajaraman, R.; Munirajan, A.K. Oral squamous cell carcinoma: Microrna expression profiling and integrative analyses for elucidation of tumourigenesis mechanism. Mol. Cancer 2016, 15, 28. [Google Scholar] [CrossRef] [PubMed]
- Manasa, V.G.; Kannan, S. Impact of microrna dynamics on cancer hallmarks: An oral cancer scenario. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Saito, Y.; Liang, G.; Egger, G.; Friedman, J.M.; Chuang, J.C.; Coetzee, G.A.; Jones, P.A. Specific activation of microrna-127 with downregulation of the proto-oncogene bcl6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 2006, 9, 435–443. [Google Scholar] [CrossRef] [PubMed]
- Balch, C.; Yan, P.; Craft, T.; Young, S.; Skalnik, D.G.; Huang, T.H.; Nephew, K.P. Antimitogenic and chemosensitizing effects of the methylation inhibitor zebularine in ovarian cancer. Mol. Cancer Ther. 2005, 4, 1505–1514. [Google Scholar] [CrossRef] [PubMed]


| Gene | Mechanism | Locus | Epigenetic Modification | Ref |
|---|---|---|---|---|
| CDKN2A/p16 | Cell cycle, senescence | 9p21 | Hypermethylation | [73,74,90,91,92] |
| CDH1/E-cadherin | EMT, adhesion | 16q22.1 | Hypermethylation * | [73,74,75,77] |
| PTEN | Differentiation, survival, proliferation, invasion, apoptosis | 10q23.3 | Hypermethylation * | [84,89] |
| DAPK1 | Apoptosis | 9q34.1 | Hypermethylation | [93,94,95] |
| MGMT | DNA repair | 10q26 | Hypermethylation | [90,92,94,96] |
| RARB2 | Cell proliferation | 3p24 | Hypermethylation | [97,98] |
| RASSF1/2 | Cell cycle, apoptosis, and microtubule formation | 3p21.3/20p13 | Hypermethylation | [99,100] |
| APC | Cell proliferation | 5q22.2 | Hypermethylation | [64,65] |
| Survivin | Cell proliferation and apoptosis | 17q25 | Hypomethylation | [70,71,101] |
| MLH1 | DNA repair | 3p22.2 | Hypermethylation | [73,102,103] |
| p14(ARF) | Cell proliferation, division, angiogenesis | 9p21 | Hypermethylation | [104,105,106] |
| p15INK4B | Cell cycle | 9p21.3 | Hypermethylation | [104,107] |
| p16INK4A | Cell cycle, senescence | 9p21 | Hypermethylation | [73,74,90,91,92] |
| RARβ | Cell growth and differentiation | 3p24 | Hypermethylation * | [97,108] |
| miRNA | Type of Malignancy | Possible Clinical Utility | Expression | Reference |
|---|---|---|---|---|
| miR-21 | Head and neck squamous cell carcinoma, Oral Squamous cell carcinoma | Diagnosis/prognosis utility (detected also in plasma) | Upregulated | [117,118] |
| miR-375 | Oral squamous cell carcinoma | Associated with clinical parameters | Downregulated | [119] |
| miR-31 | Oral carcinoma | Non-invasive and early diagnosis tool (saliva) and also prognosis marker | Upregulated | [120] |
| miR-7 | Oral squamous cell carcinoma | Upregulated | [118] | |
| miR-27b | Oral squamous cell carcinoma | Saliva biomarker for OSCC | Upregulated | [121] |
| miR-125b | Oral squamous cell carcinoma | Therapeutic target | Downregulated | [122] |
| miR-155 | Oral squamous cell carcinoma | Prognosis value | Upregulated | [123,124] |
| miR-181 | Oral squamous cell carcinoma | Lymph node metastasis marker | Upregulated | [125] |
| miR-211 | Oral carcinoma | Poor prognosis marker | Upregulated | [126] |
| Additional upregulated miRNAs | miR-9*, miR-424, miR-7–1*, miR-15b, miR-9, miR-155, and miR-196a, miR-24, miR-18a, miR-221, miR-16, let-7b, | [118,121,127] | ||
| Additional downregulated miRNAs | miR-486-5p, miR-136, miR-147, miR-1250, miR-148a, miR-632, miR-646, miR-668, miR-877, miR-503, miR-220a, miR-323-5p, miR-223, miR-29a | [118,121,127] | ||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Irimie, A.I.; Ciocan, C.; Gulei, D.; Mehterov, N.; Atanasov, A.G.; Dudea, D.; Berindan-Neagoe, I. Current Insights into Oral Cancer Epigenetics. Int. J. Mol. Sci. 2018, 19, 670. https://doi.org/10.3390/ijms19030670
Irimie AI, Ciocan C, Gulei D, Mehterov N, Atanasov AG, Dudea D, Berindan-Neagoe I. Current Insights into Oral Cancer Epigenetics. International Journal of Molecular Sciences. 2018; 19(3):670. https://doi.org/10.3390/ijms19030670
Chicago/Turabian StyleIrimie, Alexandra Iulia, Cristina Ciocan, Diana Gulei, Nikolay Mehterov, Atanas G. Atanasov, Diana Dudea, and Ioana Berindan-Neagoe. 2018. "Current Insights into Oral Cancer Epigenetics" International Journal of Molecular Sciences 19, no. 3: 670. https://doi.org/10.3390/ijms19030670