A Three–MicroRNA Signature as a Potential Biomarker for the Early Detection of Oral Cancer
Abstract
:1. Introduction
2. Results
2.1. Characteristics of Study Subjects
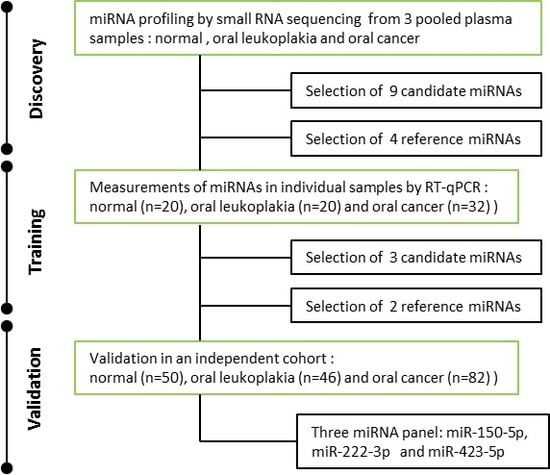
2.2. Expression Profiling of miRNAs by Small RNA-seq
2.3. Selection of Suitable Reference miRNAs
2.4. Investigation of the Six Candidate miRNAs in Training Phase
2.5. Validation of Three Significant miRNA with an Independent Cohort
2.6. Correlations between miRNA Signature and Clinical Parameters
2.7. Logistic Regression Analysis of miRNA Biomarkers
2.8. Diagnostic Performance of miRNA Signature
2.9. Bioinformatics Analysis of miR-222-3p, miR-423-5p, and miR-150-5p
3. Discussion
4. Materials and Methods
4.1. Clinical Samples
4.2. Small RNA Library Preparation and Sequencing
4.3. Small RNA Sequencing Analysis
4.4. miRNA Quantification by qRT-PCR Assays
4.5. Statistical Analysis
4.6. Bioinformatics Analysis
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| OSCC | Oral squamous cell carcinoma |
| OL | Oral leukoplakia |
| miRNAs | MicroRNAs |
| qRT-PCR | Quantitative real-time polymerase chain reaction |
| PMD | Potentially malignant disorders |
| ROC | Receiver operating characteristic |
| AUC | Area under curve |
| TCGA | The Cancer Genome Atlas |
Appendix A
| mir-222-3p | mir-423-5p | mir-150-5p | ||||
|---|---|---|---|---|---|---|
| Variables | ρ | p-Value | ρ | p-Value | ρ | p-Value |
| Age | 0.105 | NS | 0.037 | NS | −0.052 | NS |
| Gender | 0.039 | NS | 0.004 | NS | 0.103 | NS |
| Betel chewing status | −0.241 | 0.005 | −0.149 | NS | −0.099 | NS |
| Smoking status | −0.161 | NS | 0.034 | NS | 0.001 | NS |
| Alcohol status | −0.121 | NS | 0.086 | NS | −0.178 | NS |
| Clinical stage | −0.201 | 0.032 | −0.237 | 0.011 | −0.116 | NS |
| T stage | −0.220 | NS | −0.276 | 0.003 | −0.156 | NS |
| Lymph node metastasis | −0.222 | 0.018 | −0.220 | 0.019 | 0.012 | NS |
References
- Corso, G.D.; Villa, A.; Tarsitano, A.; Gohel, A. Current trends in oral cancer: A systematic review. Cancer Cell Microenviron. 2016, 3. [Google Scholar] [CrossRef]
- Lo, W.L.; Kao, S.Y.; Chi, L.Y.; Wong, Y.K.; Chang, R.C. Outcomes of oral squamous cell carcinoma in Taiwan after surgical therapy: Factors affecting survival. J. Oral Maxillofac. Surg. 2003, 61, 751–758. [Google Scholar] [CrossRef]
- Chen, Y.K.; Huang, H.C.; Lin, L.M.; Lin, C.C. Primary oral squamous cell carcinoma: An analysis of 703 cases in southern Taiwan. Oral Oncol. 1999, 35, 173–179. [Google Scholar] [CrossRef]
- Kao, S.Y.; Chu, Y.W.; Chen, Y.W.; Chang, K.W.; Liu, T.Y. Detection and screening of oral cancer and pre-cancerous lesions. J. Chin. Med. Assoc. 2009, 72, 227–233. [Google Scholar] [CrossRef]
- Jeng, J.H.; Chang, M.C.; Hahn, L.J. Role of areca nut in betel quid-associated chemical carcinogenesis: Current awareness and future perspectives. Oral Oncol. 2001, 37, 477–492. [Google Scholar] [CrossRef]
- Yeh, C.Y.; Lin, C.L.; Chang, M.C.; Chen, H.M.; Kok, S.H.; Chang, S.H.; Kuo, Y.S.; Hahn, L.J.; Chan, C.P.; Lee, J.J.; et al. Differences in oral habit and lymphocyte subpopulation affect malignant transformation of patients with oral precancer. J. Formos. Med. Assoc. 2016, 115, 263–268. [Google Scholar] [CrossRef] [PubMed]
- Neville, B.W.; Day, T.A. Oral cancer and precancerous lesions. CA Cancer J. Clin. 2002, 52, 195–215. [Google Scholar] [CrossRef] [PubMed]
- Mortazavi, H.; Baharvand, M.; Mehdipour, M. Oral potentially malignant disorders: An overview of more than 20 entities. J. Dent Res. Dent Clin. Dent Prospect. 2014, 8, 6–14. [Google Scholar]
- Zhang, B.; Pan, X.; Cobb, G.P.; Anderson, T.A. microRNAs as oncogenes and tumor suppressors. Dev. Biol. 2007, 302, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Gorenchtein, M.; Poh, C.F.; Saini, R.; Garnis, C. MicroRNAs in an oral cancer context—from basic biology to clinical utility. J. Dent. Res. 2012, 91, 440–446. [Google Scholar] [CrossRef] [PubMed]
- Cervigne, N.K.; Reis, P.P.; Machado, J.; Sadikovic, B.; Bradley, G.; Galloni, N.N.; Pintilie, M.; Jurisica, I.; Perez-Ordonez, B.; Gilbert, R.; et al. Identification of a microRNA signature associated with progression of leukoplakia to oral carcinoma. Hum. Mol. Genet. 2009, 18, 4818–4829. [Google Scholar] [CrossRef] [PubMed]
- Maimaiti, A.; Abudoukeremu, K.; Tie, L.; Pan, Y.; Li, X. MicroRNA expression profiling and functional annotation analysis of their targets associated with the malignant transformation of oral leukoplakia. Gene 2015, 558, 271–277. [Google Scholar] [CrossRef] [PubMed]
- Krysan, K.; Kusko, R.; Grogan, T.; O’Hearn, J.; Reckamp, K.L.; Walser, T.C.; Garon, E.B.; Lenburg, M.E.; Sharma, S.; Spira, A.E.; et al. PGE2-driven expression of c-Myc and oncomiR-17-92 contributes to apoptosis resistance in NSCLC. Mol. Cancer Res. 2014, 12, 765–774. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Ng, E.K.; Chong, W.W.; Jin, H.; Lam, E.K.; Shin, V.Y.; Yu, J.; Poon, T.C.; Ng, S.S.; Sung, J.J. Differential expression of microRNAs in plasma of patients with colorectal cancer: A potential marker for colorectal cancer screening. Gut 2009, 58, 1375–1381. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, K.; Yuan, Y.; Cho, J.H.; McClarty, S.; Baxter, D.; Galas, D.J. Comparing the MicroRNA spectrum between serum and plasma. PLoS ONE 2012, 7, e41561. [Google Scholar] [CrossRef] [PubMed]
- Messadi, D.V. Diagnostic aids for detection of oral precancerous conditions. Int. J. Oral Sci. 2013, 5, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Khan, R.S.; Khurshid, Z.; Asiri, F.Y.I. Advancing Point-of-Care (PoC) Testing Using Human Saliva as Liquid Biopsy. Diagnostics 2017, 7, 39. [Google Scholar] [CrossRef] [PubMed]
- Torrente-Rodriguez, R.M.; Campuzano, S.; Ruiz-Valdepenas Montiel, V.; Gamella, M.; Pingarron, J.M. Electrochemical bioplatforms for the simultaneous determination of interleukin (IL)-8 mRNA and IL-8 protein oral cancer biomarkers in raw saliva. Biosens. Bioelectron. 2016, 77, 543–548. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.C.; Chang, J.T.; Huang, Y.C.; Huang, C.C.; Chen, W.H.; Lee, L.Y.; Huang, B.S.; Chen, Y.J.; Li, H.F.; Cheng, A.J. Combined determination of circulating miR-196a and miR-196b levels produces high sensitivity and specificity for early detection of oral cancer. Clin. Biochem. 2015, 48, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.J.; Tsai, M.M.; Tu, H.F.; Lui, M.T.; Cheng, H.W.; Lin, S.C. miR-196a overexpression and miR-196a2 gene polymorphism are prognostic predictors of oral carcinomas. Ann. Surg. Oncol. 2013, 20, S406–S414. [Google Scholar] [CrossRef] [PubMed]
- Hung, P.S.; Liu, C.J.; Chou, C.S.; Kao, S.Y.; Yang, C.C.; Chang, K.W.; Chiu, T.H.; Lin, S.C. miR-146a enhances the oncogenicity of oral carcinoma by concomitant targeting of the IRAK1, TRAF6 and NUMB genes. PLoS ONE 2013, 8, e79926. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.J.; Kao, S.Y.; Tu, H.F.; Tsai, M.M.; Chang, K.W.; Lin, S.C. Increase of microRNA miR-31 level in plasma could be a potential marker of oral cancer. Oral Dis. 2010, 16, 360–364. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.J.; Lin, S.C.; Yang, C.C.; Cheng, H.W.; Chang, K.W. Exploiting salivary miR-31 as a clinical biomarker of oral squamous cell carcinoma. Head Neck 2012, 34, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Tiberio, P.; Callari, M.; Angeloni, V.; Daidone, M.G.; Appierto, V. Challenges in using circulating miRNAs as cancer biomarkers. Biomed Res. Int. 2015, 2015, 731479. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Lin, J.; Kong, D.; Huang, M.; Xu, C.; Kim, T.K.; Etheridge, A.; Luo, Y.; Ding, Y.; Wang, K. Current State of Circulating MicroRNAs as Cancer Biomarkers. Clin. Chem. 2015, 61, 1138–1155. [Google Scholar] [CrossRef] [PubMed]
- Kirschner, M.B.; Kao, S.C.; Edelman, J.J.; Armstrong, N.J.; Vallely, M.P.; van Zandwijk, N.; Reid, G. Haemolysis during sample preparation alters microRNA content of plasma. PLoS ONE 2011, 6, e24145. [Google Scholar] [CrossRef] [PubMed]
- Moldovan, L.; Batte, K.E.; Trgovcich, J.; Wisler, J.; Marsh, C.B.; Piper, M. Methodological challenges in utilizing miRNAs as circulating biomarkers. J. Cell Mol. Med. 2014, 18, 371–390. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Wang, X.; Veno, M.T.; Bakholdt, V.; Sorensen, J.A.; Krogdahl, A.; Sun, Z.; Gao, S.; Kjems, J. Circulating miRNAs as biomarkers for oral squamous cell carcinoma recurrence in operated patients. Oncotarget 2017, 8, 8206–8214. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.J.; Shen, W.G.; Liu, C.J.; Chen, Y.W.; Lu, H.H.; Tsai, M.M.; Lin, S.C. miR-221 and miR-222 expression increased the growth and tumorigenesis of oral carcinoma cells. J. Oral Pathol. Med. 2011, 40, 560–566. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Jiang, F.; Chen, X.; Liu, Z.; Ouyang, Y.; Zhao, W.; Yu, D. Downregulation of miR-221/222 by a microRNA sponge promotes apoptosis in oral squamous cell carcinoma cells through upregulation of PTEN. Oncol. Lett. 2016, 12, 4419–4426. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yu, J.; Jiang, L.; Wang, A.; Shi, F.; Ye, H.; Zhou, X. MicroRNA-222 regulates cell invasion by targeting matrix metalloproteinase 1 (MMP1) and manganese superoxide dismutase 2 (SOD2) in tongue squamous cell carcinoma cell lines. Cancer Genom. Proteom. 2009, 6, 131–139. [Google Scholar]
- Jiang, F.; Zhao, W.; Zhou, L.; Liu, Z.; Li, W.; Yu, D. MiR-222 targeted PUMA to improve sensitization of UM1 cells to cisplatin. Int. J. Mol. Sci. 2014, 15, 22128–22141. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Zhao, W.; Zhou, L.; Zhang, L.; Liu, Z.; Yu, D. miR-222 regulates the cell biological behavior of oral squamous cell carcinoma by targeting PUMA. Oncol. Rep. 2014, 31, 1255–1262. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.Y.; Bai, Y.N.; Luo, L.X.; Wu, H.; Zeng, Y. Expression of microRNA-150 targeting vascular endothelial growth factor-A is downregulated under hypoxia during liver regeneration. Mol. Med. Rep. 2013, 8, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Naruse, T.; Kawasaki, G.; Yanamoto, S.; Mizuno, A.; Umeda, M. Immunohistochemical study of VEGF expression in oral squamous cell carcinomas: Correlation with the mTOR-HIF-1alpha pathway. Anticancer Res 2011, 31, 4429–4437. [Google Scholar] [PubMed]
- Roy, R.; Singh, R.; Chattopadhyay, E.; Ray, A.; Sarkar, N.D.; Aich, R.; Paul, R.R.; Pal, M.; Roy, B. MicroRNA and target gene expression based clustering of oral cancer, precancer and normal tissues. Gene 2016, 593, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Brase, J.C.; Johannes, M.; Schlomm, T.; Falth, M.; Haese, A.; Steuber, T.; Beissbarth, T.; Kuner, R.; Sultmann, H. Circulating miRNAs are correlated with tumor progression in prostate cancer. Int. J. Cancer 2011, 128, 608–616. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Ren, C.; Han, J.; Ding, Y.; Du, J.; Dai, N.; Dai, J.; Ma, H.; Hu, Z.; Shen, H.; et al. A five-microRNA panel in plasma was identified as potential biomarker for early detection of gastric cancer. Br. J. Cancer 2014, 110, 2291–2299. [Google Scholar] [CrossRef] [PubMed]
- Cabibi, D.; Caruso, S.; Bazan, V.; Castiglia, M.; Bronte, G.; Ingrao, S.; Fanale, D.; Cangemi, A.; Calo, V.; Listi, A.; et al. Analysis of tissue and circulating microRNA expression during metaplastic transformation of the esophagus. Oncotarget 2016, 7, 47821–47830. [Google Scholar] [CrossRef] [PubMed]
- Jo, P.; Azizian, A.; Salendo, J.; Kramer, F.; Bernhardt, M.; Wolff, H.A.; Gruber, J.; Grade, M.; Beissbarth, T.; Ghadimi, B.M.; et al. Changes of Microrna Levels in Plasma of Patients with Rectal Cancer during Chemoradiotherapy. Int. J. Mol. Sci. 2017, 18, 1140. [Google Scholar] [CrossRef] [PubMed]
- Molina-Pinelo, S.; Suarez, R.; Pastor, M.D.; Nogal, A.; Marquez-Martin, E.; Martin-Juan, J.; Carnero, A.; Paz-Ares, L. Association between the miRNA signatures in plasma and bronchoalveolar fluid in respiratory pathologies. Dis. Markers 2012, 32, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Brunet Vega, A.; Pericay, C.; Moya, I.; Ferrer, A.; Dotor, E.; Pisa, A.; Casalots, A.; Serra-Aracil, X.; Oliva, J.C.; Ruiz, A.; et al. microRNA expression profile in stage III colorectal cancer: Circulating miR-18a and miR-29a as promising biomarkers. Oncol. Rep. 2013, 30, 320–326. [Google Scholar] [CrossRef] [PubMed]
- Pigati, L.; Yaddanapudi, S.C.; Iyengar, R.; Kim, D.J.; Hearn, S.A.; Danforth, D.; Hastings, M.L.; Duelli, D.M. Selective release of microRNA species from normal and malignant mammary epithelial cells. PLoS ONE 2010, 5, e13515. [Google Scholar] [CrossRef] [PubMed]
- Shiah, S.G.; Shieh, Y.S.; Chang, J.Y. The Role of Wnt Signaling in Squamous Cell Carcinoma. J. Dent. Res. 2016, 95, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Makarev, E.; Schubert, A.D.; Kanherkar, R.R.; London, N.; Teka, M.; Ozerov, I.; Lezhnina, K.; Bedi, A.; Ravi, R.; Mehra, R.; et al. In silico analysis of pathways activation landscape in oral squamous cell carcinoma and oral leukoplakia. Cell Death Discov. 2017, 3, 17022. [Google Scholar] [CrossRef] [PubMed]
- Kroh, E.M.; Parkin, R.K.; Mitchell, P.S.; Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods 2010, 50, 298–301. [Google Scholar] [CrossRef] [PubMed]








| Screening and Training Phase (n = 72) | Validation Phase (n = 178) | |||||
|---|---|---|---|---|---|---|
| Variables | Normal (%) | OL (%) | OSCC (%) | Normal (%) | OL (%) | OSCC (%) |
| Number | 20 | 20 | 32 | 50 | 46 | 82 |
| Age (mean ± SD) | 52.05 ± 12.78 | 52.20 ± 12.53 | 52.20 ± 9.03 | 52.86 ± 14.06 | 48.35 ± 12.11 | 53.79 ± 11.25 |
| Sex | ||||||
| Male | 20 (100.0) | 18 (90.0) | 31 (96.8) | 48 (96.0) | 44 (95.6) | 80 (97.5) |
| Female | 0 (0.0) | 2 (10.0) | 1 (3.2) | 2 (4.0) | 2 (4.4) | 2 (2.5) |
| Smoking | ||||||
| Non-smoker | 4 (20.0) | 6 (30.0) | 1 (3.2) | 2 (4.0) | 6 (13.0) | 7 (8.5) |
| Former smoker | 9 (45.0) | 11 (55.0) | 12 (37.5) | 15 (30.0) | 12 (26.1) | 16 (19.5) |
| Current smoker | 7 (35.0) | 3 (15.0) | 19 (59.3) | 33 (66.0) | 28 (60.9) | 59 (72.0) |
| BQ chewing | ||||||
| Non-BQ | 11 (55.0) | 6 (30.0) | 3 (9.4) | 27 (54.0) | 12 (26.1) | 7 (8.5) |
| Former BQ chewing | 6 (30.0) | 10 (50.0) | 25 (78.1) | 15 (30.0) | 28 (60.9) | 63 (76.8) |
| Current BQ-chewing | 3 (15.0) | 4 (20.0) | 4 (12.5) | 8 (16.0) | 6 (13.0) | 12 (14.7) |
| Alcohol consumption | ||||||
| Non-drinker | 5 (25.0) | 9 (45.0) | 11 (34.4) | 20 (40.0) | 20 (43.5) | 22 (26.8) |
| Former drinker | 10 (50.0) | 9 (45.0) | 13 (40.6) | 22 (44.0) | 23 (50.0) | 36 (43.9) |
| Current drinker | 5 (25.0) | 2 (10.0) | 8 (25.0) | 8 (16.0) | 3 (6.5) | 24 (29.3) |
| Stage | ||||||
| I | 14 (43.8) | 32 (39.0) | ||||
| II | 0 (0.0) | 15 (18.3) | ||||
| III | 0 (0.0) | 11 (13.4) | ||||
| IV | 18 (56.2) | 24 (29.3) | ||||
| T stage | ||||||
| T1 | 14 (43.8) | 33 (40.2) | ||||
| T2 | 3 (9.4) | 24 (29.3) | ||||
| T3 | 0 (0.0) | 4 (4.9) | ||||
| T4 | 15 (46.8) | 21 (25.6) | ||||
| N stage | ||||||
| N0 | 19 (59.4) | 60 (73.2) | ||||
| N1 | 5 (15.6) | 11 (13.4) | ||||
| N2 | 8 (25.0) | 11 (13.4) | ||||
| Univariate | Mulitivariate | |||
|---|---|---|---|---|
| Relative Risk | p-Value | Relative Risk | p-Value | |
| Model 1 | ||||
| OL | ||||
| miR-222-3p | 0.205 (0.123−0.344) | <0.001 | 0.212 (0.127−0.357) | <0.001 |
| miR-150-5p | 1.114 (1.027−1.210) | 0.010 | 1.124 (1.032−1.223) | 0.007 |
| miR-423-5p | 0.880 (0.673−1.150) | 0.349 | 0.897 (0.682−1.180) | 0.437 |
| miR panel a | 1.348 (1.233−1.474) | <0.001 | 1.361 (1.238−1.496) | <0.001 |
| OSCC | ||||
| miR-222-3p | 1.038 (0.858−1.256) | 0.699 | 1.114 (0.898−1.383) | 0.324 |
| miR-150-5p | 1.189 (1.083−1.306) | <0.001 | 1.198 (1.079−1.330) | 0.001 |
| miR-423-5p | 1.466 (1.182−1.817) | <0.001 | 1.599 (1.238−2.066) | <0.001 |
| miR panel b | 1.377 (1.198−1.584) | <0.001 | 1.386 (1.189−1.615) | <0.001 |
| Model 2 | ||||
| OSCC | ||||
| miR-222-3p | 2.915 (2.087−4.072) | <0.001 | 3.014 (2.102−4.321) | <0.001 |
| miR-150-5p | 1.038 (1.003−1.075) | 0.035 | 1.048 (1.007−1.091) | 0.020 |
| miR-423-5p | 1.581 (1.238−2.021) | <0.001 | 1.601 (1.236−2.075) | <0.001 |
| miR panel c | 1.455 (1.308−1.617) | <0.001 | 1.448 (1.292−1.623) | <0.001 |
| Term | No. of Genes | p-Value | Fold Enrichment |
|---|---|---|---|
| Wnt signaling pathway | 16 | 4.69 × 10−6 | 4.22 |
| Pathways in cancer | 26 | 6.07 × 10−5 | 2.41 |
| Hepatitis B | 14 | 1.55 × 10−4 | 3.51 |
| Axon guidance | 13 | 1.70 × 10−4 | 3.72 |
| Sphingolipid signaling pathway | 11 | 1.58 × 10−3 | 3.33 |
| HTLV-I infection | 17 | 1.66 × 10−3 | 2.42 |
| PI3K-Akt signaling pathway | 20 | 2.71 × 10−3 | 2.11 |
| Proteoglycans in cancer | 14 | 3.19 × 10−3 | 2.55 |
| FoxO signaling pathway | 11 | 3.57 × 10−3 | 2.99 |
| Rap1 signaling pathway | 14 | 4.79 × 10−3 | 2.42 |
| MicroRNAs in cancer | 17 | 4.84 × 10−3 | 2.17 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, Y.-A.; Weng, S.-L.; Yang, S.-F.; Chou, C.-H.; Huang, W.-C.; Tu, S.-J.; Chang, T.-H.; Huang, C.-N.; Jong, Y.-J.; Huang, H.-D. A Three–MicroRNA Signature as a Potential Biomarker for the Early Detection of Oral Cancer. Int. J. Mol. Sci. 2018, 19, 758. https://doi.org/10.3390/ijms19030758
Chang Y-A, Weng S-L, Yang S-F, Chou C-H, Huang W-C, Tu S-J, Chang T-H, Huang C-N, Jong Y-J, Huang H-D. A Three–MicroRNA Signature as a Potential Biomarker for the Early Detection of Oral Cancer. International Journal of Molecular Sciences. 2018; 19(3):758. https://doi.org/10.3390/ijms19030758
Chicago/Turabian StyleChang, Yi-An, Shun-Long Weng, Shun-Fa Yang, Chih-Hung Chou, Wei-Chih Huang, Siang-Jyun Tu, Tzu-Hao Chang, Chien-Ning Huang, Yuh-Jyh Jong, and Hsien-Da Huang. 2018. "A Three–MicroRNA Signature as a Potential Biomarker for the Early Detection of Oral Cancer" International Journal of Molecular Sciences 19, no. 3: 758. https://doi.org/10.3390/ijms19030758