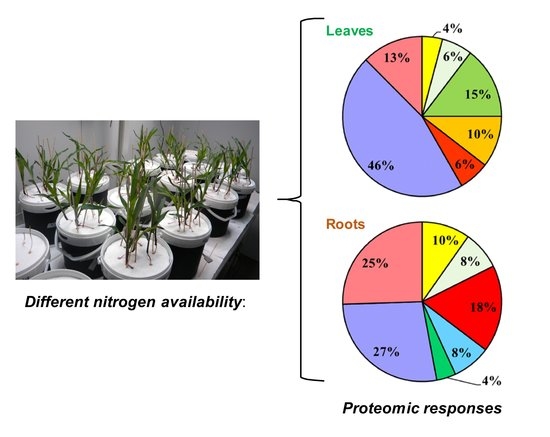
Time-Course of Metabolic and Proteomic Responses to Different Nitrate/Ammonium Availabilities in Roots and Leaves of Maize
Abstract
:1. Introduction
2. Results and Discussion
2.1. Plant Growth and Metabolic Status of Roots and Leaves
2.2. Comparative Proteomic Analyses in Roots and Leaves of Maize during Exposure to Different N Sources
2.3. Proteomic Changes Involved in Nitrogen (N) Assimilation and Amino Acid Metabolism
2.4. Changes in Proteins Involved in Photosynthesis, Energy, and Carbon Metabolism
2.5. Root Proteomic Changes Involved in Cell Water Homeostasis and Cell Wall Metabolism
2.6. Proteomic Changes Related to Protein Synthesis and Folding
2.7. Stress Responses and Other Functions
3. Materials and Methods
3.1. Plant Material and Nutritional Treatments
3.2. Determination of the Contents of Nitrate, Ammonium, Amino Acids, Sucrose and Reducing Sugars
3.3. Protein Extraction, Gel Electrophoresis and in-Gel Digestion
3.4. Mass Spectrometry Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| 2-ME | 2-Mercaptoethanol |
| CBB | Coomassie Brilliant Blue |
| DPs | Differentially abundant Proteins |
| FA | Formic acid |
| NUP | Number of unique peptides |
| OPA | o-Phthalaldehyde |
| SDS | Sodium dodecyl sulfate |
| SI | Spectrum intensity |
References
- Hawkesford, M.; Horst, W.; Kichey, T.; Lambers, H.; Schjoerring, J.; Møller, I.S.; White, P. Marschner’s Mineral Nutrition of Higher Plants, 3rd ed.; Elsevier/Academic Press: Amsterdam, The Netherlands, 2011; pp. 135–151. ISBN 978-0-12-384905-2. [Google Scholar]
- Miller, A.J.; Cramer, M.D. Root nitrogen acquisition and assimilation. Plant Soil 2004, 274, 1–36. [Google Scholar] [CrossRef]
- Andrews, M.; Raven, J.A.; Lea, P.J. Do plants need nitrate? The mechanisms by which nitrogen form affects plants. Ann. Appl. Biol. 2013, 163, 174–199. [Google Scholar] [CrossRef]
- Krapp, A. Plant nitrogen assimilation and its regulation: A complex puzzle with missing pieces. Curr. Opin. Plant Biol. 2015, 25, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Kiba, T.; Krapp, A. Plant nitrogen acquisition under low availability: Regulation of uptake and root architecture. Plant Cell Physiol. 2016, 57, 707–714. [Google Scholar] [CrossRef] [PubMed]
- Nunes-Nesi, A.; Fernie, A.R.; Stitt, M. Metabolic and signaling aspects underpinning the regulation of plant carbon nitrogen interactions. Mol. Plant 2010, 3, 973–996. [Google Scholar] [CrossRef] [PubMed]
- Masakapalli, S.K.; Kruger, N.J.; Ratcliffe, R.G. The metabolic flux phenotype of heterotrophic Arabidopsis cells reveals a complex response to changes in nitrogen supply. Plant J. 2013, 74, 569–582. [Google Scholar] [CrossRef] [PubMed]
- Andrews, M.; Lea, P.J.; Raven, J.A.; Azevedo, R.A. Nitrogen use efficiency. 3. Nitrogen fixation. Genes and costs. Ann. Appl. Biol. 2009, 155, 1–13. [Google Scholar] [CrossRef]
- Meyer, C.; Stitt, M. Nitrate reduction and signaling. In Plant Nitrogen; Lea, P.J., Morot-Gaudry, J.F., Eds.; Springer: Berlin, Germany, 2001; pp. 37–59. ISBN 3-540-67799-2. [Google Scholar]
- Britto, D.T.; Kronzucker, H.J. NH4+ toxicity in higher plants: A critical review. J. Plant Physiol. 2002, 159, 567–584. [Google Scholar] [CrossRef]
- Esteban, R.; Ariz, I.; Crus, C.; Moran, J.F. Review: Mechanisms of ammonium toxicity and the quest for tolerance. Plant Sci. 2016, 248, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Touraine, B. Nitrate uptake by roots—Transporters and root development. In Nitrogen Acquisition and Assimilation in Higher Plants; Amâncio, S., Stulen, I., Eds.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2004; pp. 1–34. ISBN 1-4020-2727-3. [Google Scholar]
- Hachiya, T.; Sakakibara, H. Interactions between nitrate and ammonium in their uptake, allocation, assimilation, and signalling in plants. J. Exp. Bot. 2017, 68, 2501–2512. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, J.A.; Vega, A.; Bouguyon, E.; Krouk, G.; Gojon, A.; Coruzzi, G.; Gutiérrez, R.A. Nitrate transport, sensing, and responses in plants. Mol. Plant 2016, 9, 837–856. [Google Scholar] [CrossRef] [PubMed]
- Hirel, B.; Le Gouis, J.; Ney, B.; Gallais, A. The challenge of improving nitrogen use efficiency in crop plants: Towards a more central role for genetic variability and quantitative genetics within integrated approaches. J. Exp. Bot. 2007, 58, 2369–2387. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Han, L.; Chen, F.; Bao, J.; Zhang, F.; Mi, G. Microarray analysis reveals early responsive genes possibly involved in localized nitrate stimulation of lateral root development in maize (Zea mays L.). Plant Sci. 2008, 175, 272–282. [Google Scholar] [CrossRef]
- Zamboni, A.; Astolfi, S.; Zuchi, S.; Pii, Y.; Guardini, K.; Tononi, P.; Varanini, Z. Nitrate induction triggers different transcriptional changes in a high and a low nitrogen use efficiency maize inbred line. J. Integr. Plant Biol. 2014, 56, 1080–1094. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pii, Y.; Alessandrini, M.; Dall’Osto, L.; Guardini, K.; Prinsi, B.; Espen, L.; Zamboni, A.; Varanini, Z. Time-resolved investigation of molecular components involved in the induction of NO3− High Affinity Transport system in maize roots. Front. Plant Sci. 2016, 7, 1657. [Google Scholar] [CrossRef] [PubMed]
- Amiour, N.; Imbaud, S.; Clément, G.; Agier, N.; Zivy, M.; Valot, B.; Balliau, T.; Armengaud, P.; Quilleré, I.; Cañas, R.; et al. The use of metabolomics integrated with transcriptomic and proteomic studies for identifying key steps involved in the control of nitrogen metabolism in crops such as maize. J. Exp. Bot. 2012, 63, 5017–5033. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prinsi, B.; Negri, A.S.; Pesaresi, P.; Cocucci, M.; Espen, L. Evaluation of protein pattern changes in roots and leaves of Zea mays plants in response to nitrate availability by two-dimensional gel electrophoresis analysis. BMC Plant Biol. 2009, 9, 113. [Google Scholar] [CrossRef] [PubMed]
- Plénet, D.; Lemaire, G. Relationships between dynamics of nitrogen uptake and dry matter accumulation in maize crops. Determination of critical N concentration. Plant Soil 2000, 216, 65–82. [Google Scholar] [CrossRef]
- Schortemeyer, M.; Stamp, P.; Feil, B. Ammonium tolerance and carbohydrate status in maize cultivars. Ann. Bot. 1997, 79, 25–30. [Google Scholar] [CrossRef]
- Andrews, M.; Raven, J.A.; Sprent, J.I. Environmental effects on dry matter partitioning between shoot and root of crop plants: Relations with growth and shoot protein concentration. Ann. Appl. Biol. 2001, 138, 57–68. [Google Scholar] [CrossRef]
- Cramer, M.D.; Lewis, O.A.M. The influence of NO3− and NH4+ on the carbon and nitrogen partitioning characteristics of wheat (Triticum aestivum L.) and maize (Zea mays L.) plants. Plant Soil 1993, 154, 289–300. [Google Scholar] [CrossRef]
- Kronzucker, H.J.; Glass, A.D.M.; Siddiqi, M.Y. Inhibition of nitrate uptake by ammonium in barley. Analysis of the component fluxes. Plant Physiol. 1999, 120, 283–291. [Google Scholar] [CrossRef] [PubMed]
- Cerezo, M.; Tillard, P.; Filleur, S.; Muños, S.; Daniel-Vedele, F.; Gojon, A. Major alterations of the regulation of root NO3− uptake are associated with the mutation of Nrt2.1 and Nrt2.2 genes in Arabidopsis. Plant Physiol. 2001, 127, 262–271. [Google Scholar] [CrossRef] [PubMed]
- Prinsi, B.; Espen, L. Mineral nitrogen sources differently affect root glutamine synthetase isoforms and amino acid balance among organs in maize. BMC Plant Biol. 2015, 15, 96. [Google Scholar] [CrossRef] [PubMed]
- Gilmore, J.M.; Washburn, M.P. Advances in shotgun proteomics and the analysis of membrane proteomes. J. Proteom. 2010, 73, 2078–2091. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Okamoto, M.; Xing, X.; Crawford, N.M. Microarray analysis of the nitrate response in Arabidopsis roots and shoots reveals over 1000 rapidly responding genes and new linkages to glucose, trehalose-6-phosphate, iron and sulphate metabolism. Plant Physiol. 2003, 132, 556–567. [Google Scholar] [CrossRef] [PubMed]
- Patterson, K.; Cakmak, T.; Cooper, A.; Larger, I.; Rasmusson, A.G.; Escobar, M.A. Distinct signalling pathways and transcriptome response signatures differentiate ammonium- and nitrate-supplied plants. Plant Cell Environ. 2010, 33, 1486–1501. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, A.; Oaks, A.; Jacquot, J.-P.; Vidal, J.; Gadal, P. An electron transport system in maize roots for reactions of glutamate synthase and nitrite reductase. Plant Physiol. 1985, 78, 374–378. [Google Scholar] [CrossRef] [PubMed]
- Redinbaugh, M.G.; Campbell, W.H. Higher plant responses to environmental nitrate. Physiol. Plant. 1991, 82, 640–650. [Google Scholar] [CrossRef]
- Cren, M.; Hirel, B. Glutamine synthetase in higher plants: Regulation of gene and protein expression from the organ to cell. Plant Cell Physiol. 1999, 40, 1187–1193. [Google Scholar] [CrossRef]
- Lea, P.J.; Sodek, L.; Parry, M.A.J.; Shewry, P.R.; Halford, N.G. Asparagine in plants. Ann. Appl. Biol. 2007, 150, 1–26. [Google Scholar] [CrossRef] [Green Version]
- Fritz, C.; Palacios-Rojas, N.; Feil, R.; Stitt, M. Regulation of secondary metabolism by the carbon-nitrogen status in tobacco: Nitrate inhibits large sectors of phenylpropanoid metabolism. Plant J. 2006, 46, 533–548. [Google Scholar] [CrossRef] [PubMed]
- Amunts, A.; Nelson, N. Functional organization of a plant Photosystem I: Evolution of a highly efficient photochemical machine. Plant Physiol. Biochem. 2008, 46, 228–237. [Google Scholar] [CrossRef] [PubMed]
- Allahverdiyeva, Y.; Suorsa, M.; Rossi, F.; Pavesi, A.; Kater, M.M.; Antonacci, A.; Tadini, L.; Pribil, M.; Schneider, A.; Wanner, G.; et al. Arabidopsis plants lacking PsbQ and PsbR subunits of the oxygen-evolving complex show altered PSII super-complex organization and short-term adaptive mechanisms. Plant J. 2013, 75, 671–684. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wollman, F.-A. State transitions reveal the dynamics and flexibility of the photosynthetic apparatus. EMBO J. 2001, 20, 3623–3630. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-B.; Lu, T.-C.; Wang, H.-X.; Shen, J.; Bu, T.-T.; Chao, Q.; Gao, Z.-F.; Zhu, X.-G.; Wang, Y.-F.; Wang, B.-C. Posttranslational modification of maize chloroplast pyruvate orthophosphate dikinase reveals the precise regulatory mechanism of its enzymatic activity. Plant Physiol. 2014, 165, 534–549. [Google Scholar] [CrossRef] [PubMed]
- Flügge, U.-I. Phosphate translocators in plastids. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1999, 50, 27–45. [Google Scholar] [CrossRef] [PubMed]
- Hammargren, J.; Sundström, J.; Johansson, M.; Bergman, P.; Knorpp, C. On the phylogeny, expression and targeting of plant nucleoside diphosphate kinases. Physiol. Plant. 2007, 129, 79–89. [Google Scholar] [CrossRef]
- Redinbaugh, M.G.; Campbell, W.H. Nitrate regulation of the oxidative pentose phosphate pathway in maize (Zea mays L.) root plastids: Induction of 6-phosphogluconate dehydrogenase activity, protein and transcript levels. Plant Sci. 1998, 134, 129–140. [Google Scholar] [CrossRef]
- Esposito, S.; Guerriero, G.; Vona, V.; Di Martino Rigano, V.; Carfagna, S.; Rigano, C. Glutamate synthase activities and protein changes in relation to nitrogen nutrition in barley: The dependence on different plastidic glucose-6P dehydrogenase isoforms. J. Exp. Bot. 2005, 56, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Fernie, A.R.; Carrari, F.; Sweetlove, L.J. Respiratory metabolism: Glycolysis, the TCA cycle and mitochondrial electron transport. Curr. Opin. Plant Biol. 2004, 7, 254–261. [Google Scholar] [CrossRef] [PubMed]
- Britto, D.T.; Kronzucker, H.J. Nitrogen acquisition, PEP carboxylase, and cellular pH homeostasis: New views on old paradigms. Plant Cell Environ. 2005, 28, 1396–1409. [Google Scholar] [CrossRef]
- Coskun, D.; Britto, D.V.; Kronzucker, H.J. The nitrogen-potassium intersections: Membranes, metabolism, and mechanism. Plant Cell Environ. 2017, 40, 2029–2041. [Google Scholar] [CrossRef] [PubMed]
- Coskun, D.; Britto, D.T.; Li, M.; Oh, S.; Kronzucker, H.J. Capacity and plasticity of potassium channels and high-affinity transporters in roots of barley and Arabidopsis. Plant Physiol. 2013, 162, 496–511. [Google Scholar] [CrossRef] [PubMed]
- Maurel, C.; Verdoucq, L.; Luu, D.-T.; Santoni, V. Plant aquaporins: Membrane channels with multiple integrated functions. Annu. Rev. Plant Biol. 2008, 59, 595–624. [Google Scholar] [CrossRef] [PubMed]
- Gorska, A.; Zwieniecka, A.; Holbrook, N.M.; Zwieniecki, M.A. Nitrate induction of root hydraulic conductivity in maize is not correlated with aquaporin expression. Planta 2008, 228, 989–998. [Google Scholar] [CrossRef] [PubMed]
- Loqué, D.; Ludewig, U.; Yuan, L.; von Wirén, N. Tonoplast intrinsic proteins AtTIP2;1 and AtTIP2;3 facilitate NH3 transport into the vacuole. Plant Physiol. 2005, 137, 671–680. [Google Scholar] [CrossRef] [PubMed]
- Stiefel, V.; Ruiz-Avila, L.; Raz, R.; Pilar Vallés, M.; Gómez, J.; Pagés, M.; Martinez-Izquierdo, J.A.; Ludevid, M.D.; Langdale, J.A.; Nelson, T.; Puigdomènech, P. Expression of a maize cell wall hydroxyproline-rich glycoprotein gene in early leaf and root vascular differentiation. Plant Cell 1990, 2, 785–793. [Google Scholar] [CrossRef] [PubMed]
- Held, B.M.; Wang, H.; John, I.; Syrkin Wurtele, E.S.; Colbert, J.T. An mRNA putatively coding for an O-methyltransferase accumulates preferentially in maize roots and is located predominantly in the region of the endodermis. Plant Physiol. 1993, 102, 1001–1008. [Google Scholar] [CrossRef] [PubMed]
- Scheible, W.R.; Morcuende, R.; Czechowski, T.; Fritz, C.; Osuna, D.; Palacio-Rojas, N.; Schindelasch, D.; Thimm, O.; Udvardi, M.K.; Stitt, M. Genome-wide reprogramming of primary and secondary metabolism, protein synthesis, cellular growth processes, ant the regulatory infrastructure of Arabidopsis in response to nitrogen. Plant Physiol. 2004, 136, 2483–2499. [Google Scholar] [CrossRef] [PubMed]
- Barakat, A.; Szick-Miranda, K.; Chang, I.-F.; Guyot, R.; Blanc, G.; Cooke, R.; Delseny, M.; Bailey-Serres, J. The organization of cytoplasmic ribosomal protein genes in the Arabidopsis genome. Plant Physiol. 2001, 127, 398–415. [Google Scholar] [CrossRef] [PubMed]
- Sormani, R.; Masclaux-Daubresse, C.; Daniel-Vedele, F.; Chardon, F. Transcriptional regulation of ribosome components are determined by stress according to cellular compartment in Arabidopsis thaliana. PLoS ONE 2011, 6, e28070. [Google Scholar] [CrossRef] [PubMed]
- Carroll, A.J. The Arabidopsis cytosolic ribosomal proteome: From form to function. Front. Plant Sci. 2013, 4, 32. [Google Scholar] [CrossRef] [PubMed]
- Schiene, C.; Fischer, G. Enzymes that catalyse the restructuring of proteins. Curr. Opin. Struct. Biol. 2000, 10, 40–45. [Google Scholar] [CrossRef]
- Marivet, J.; Frenso, P.; Burkard, G. Effects of abiotic stresses on cyclophilin gene expression in maize and bean and sequence analysis of bean cyclophilin cDNA. Plant Sci. 1992, 84, 171–178. [Google Scholar] [CrossRef]
- Lee, J.-P.; Palfrey, H.C.; Bindokas, V.P.; Ghadge, G.D.; Ma, L.; Miller, R.J.; Roos, R.P. The role of immunophilins in mutant superoxide dismutase-1-linked familial amylotrophic lateral sclerosis. Proc. Natl. Acad. Sci. USA 1999, 96, 3251–3256. [Google Scholar] [CrossRef] [PubMed]
- Colas des Francs-Small, C.; Ambard-Breteville, F.; Small, F.; Rémy, R. Identification of a major soluble protein in mitochondria from nonphotosynthetic tissues as NAD-dependent formate dehydrogenase. Plant Physiol. 1993, 102, 1171–1177. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hanson, A.D.; Gage, D.A.; Shachar-Hill, Y. Plant one-carbon metabolism and its engineering. Trends Plant Sci. 2000, 5, 206–213. [Google Scholar] [CrossRef]
- Lou, H.Q.; Gong, Y.L.; Fan, W.; Xu, J.M.; Liu, Y.; Cao, M.J.; Wang, M.-H.; Yang, J.L.; Zheng, S.J. A formate dehydrogenase confers tolerance to aluminum and low pH. Plant Physiol. 2016, 171, 294–305. [Google Scholar] [CrossRef] [PubMed]
- Porta, H.; Rocha-Sosa, M. Plant lipoxygenases. Physiological and molecular features. Plant Physiol. 2002, 130, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Shigeto, J.; Tsutsumi, Y. Diverse functions and reactions of class III peroxidases. New Phytol. 2016, 209, 1395–1402. [Google Scholar] [CrossRef] [PubMed]
- Cataldo, D.A.; Maroon, M.; Schrader, L.E.; Youngs, V.L. Rapid colorimetric determination of nitrate in plant tissue by nitration of salicylic acid. Commun. Soil Sci. Plant Anal. 1975, 6, 71–80. [Google Scholar] [CrossRef]
- Coskun, D.; Britto, D.T.; Jean, Y.-K.; Schulze, L.M.; Backer, A.; Kronzucker, H.J. Silver ions disrupt K+ homeostasis and cellular integrity in intact barley (Hordeum vulgare L.) roots. J. Exp. Bot. 2012, 63, 151–162. [Google Scholar] [CrossRef] [PubMed]
- Moore, S.; Stein, W.H. A modified ninhydrin reagent for the photometric determination of amino acids and related compounds. J. Biol. Chem. 1954, 211, 907–913. [Google Scholar] [PubMed]
- Nelson, N. A photometric adaptation of the Somogy method for the determination of glucose. J. Biol. Chem 1944, 153, 375–380. [Google Scholar]
- Mariotti, F.; Tomé, D.; Patureau Mirand, P. Converting nitrogen into protein—Beyond 6.25 and Jones’ factors. Crit. Rev. Food Sci. 2008, 48, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]







| Condition | Time (h) | Total N Plant−1 (mg g−1 FW) | Inorganic N/Organic N Roots | Inorganic N/Organic N Leaves |
|---|---|---|---|---|
| 0 | 3.37 ± 0.02 (a) | 0.50 ± 0.07 (bc) | 0.24 ± 0.03 (abc) | |
| Nitrate (n) | 6 | 3.73 ± 0.06 (ab) | 0.80 ± 0.01 (de) | 0.23 ± 0.01 (ab) |
| 30 | 4.25 ± 0.10 (cd) | 1.28 ± 0.10 (f) | 0.26 ± 0.02 (bc) | |
| 54 | 4.38 ± 0.10 (cd) | 0.91 ± 0.07 (e) | 0.34 ± 0.01 (c) | |
| Ammonium (a) | 6 | 4.13 ± 0.04 (bc) | 0.49 ± 0.08 (bc) | 0.28 ± 0.01 (bc) |
| 30 | 4.55 ± 0.03 (de) | 0.34 ± 0.03 (ab) | 0.21 ± 0.01 (ab) | |
| 54 | 4.50 ± 0.06 (cde) | 0.13 ± 0.01 (a) | 0.14 ± 0.01 (a) | |
| Nitrate + Ammonium (na) | 6 | 4.34 ± 0.14 (cd) | 0.74 ± 0.02 (cde) | 0.28 ± 0.03 (bc) |
| 30 | 4.90 ± 0.10 (e) | 0.74 ± 0.07 (cde) | 0.28 ± 0.02 (bc) | |
| 54 | 4.60 ± 0.06 (de) | 0.61 ± 0.04 (bcd) | 0.30 ± 0.04 (bc) |
| Parameter | Root | Leaf |
|---|---|---|
| n. of total peptides | 39,516 | 31,739 |
| Average of peptides per sample (±SE, n = 27) | 1464 ± 15 | 1176 ± 11 |
| Average of peptides per condition (±SE, n = 9) | 4391 ± 55 | 3527 ± 50 |
| Average of unique peptide per protein (±SE) | 6.8 ± 0.3 | 6.4 ± 0.4 |
| Average amino acid coverage % (±SE) | 22.5 ± 0.8 | 22.4 ± 0.9 |
| n. of identified proteins | 336 | 246 |
| n. of differentially accumulated proteins (%) | 51 (15%) | 48 (19%) |
| ID | Entry—Protein Name | FC | Differences | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| V | h (n) | h (a) | h (na) | |||||||||||
| t | s | i | 06 | 30 | 54 | 06 | 30 | 54 | 06 | 30 | 54 | |||
| Nitrogen assimilation | ||||||||||||||
| R74 | B6SY01—Ferredoxin-nitrite reductase | 7.57 | * | abc | d | cd | abc | ab | a | abc | bcd | abc | ||
| R64 | B4FR61—Glutamine synthetase (cytosol) | 2.60 | * | a | a | a | ab | c | c | a | a | cb | ||
| R189 | A0A1D6NFK0—Glutamate synthase 1 [NADH] chloroplastic | 8.54 | * | * | a | ab | a | a | ab | b | a | a | ab | |
| R103 | B5U8J8—Asparagine synthetase | n.d. | * | a | a | a | a | b | c | a | a | a | ||
| R68 | B4G043—Ferredoxin-NADP reductase | 5.48 | * | cb | cb | c | cb | ab | a | cb | ab | abc | ||
| Amino acid metabolism | ||||||||||||||
| R6 | B6U4D6—Phenylalanine ammonia-lyase | 2.88 | * | bc | abc | bc | bc | a | a | c | abc | ab | ||
| R107 | P80608—Cysteine synthase | 2.74 | * | a | bc | ab | ab | ab | c | a | bc | abc | ||
| R49 | A0A1D6FUX8—S-adenosylmethionine synthase | 2.16 | * | b | a | ab | ab | ab | ab | b | ab | ab | ||
| R96 | K7TSD2—Serine hydroxymethyltransferase | 2.08 | * | b | ab | ab | ab | ab | a | ab | a | a | ||
| Carbon metabolism | ||||||||||||||
| R207 | C0PMR3—Glucose-6-phosphate 1-dehydrogenase (cytosol) | 7.66 | * | ab | b | ab | a | ab | ab | ab | ab | ab | ||
| R85 | A0A1D6J424—Glucose-6-phosphate 1-dehydrogenase (chloroplast) | 224 | * | ab | b | ab | ab | a | a | ab | ab | ab | ||
| R57 | A0A1D6NVZ7—Transketolase 1 | 2.98 | * | * | ab | a | ab | ab | a | a | b | ab | a | |
| R231 | Q8L5G8—Phosphotransferase | 7.59 | * | ab | b | ab | a | ab | ab | a | ab | ab | ||
| R168 | A0A1D6NR86—Pyrophosphate-fructose 6-phosphate 1-phosphotransferase subunit beta | 2.85 | * | a | b | ab | a | a | a | a | b | ab | ||
| R243 | B4F820—Triosephosphate isomerase a | 3.67 | * | a | ab | abc | a | ab | c | abc | abc | bc | ||
| R19 | C0HHU2—2,3-bisphosphoglycerate-independent phosphoglycerate mutase 1 | 2.35 | * | a | ab | ab | a | ab | c | a | a | cb | ||
| R119 | A0A1D6H4C4—Citrate synthase | 2.87 | * | ab | ab | a | a | ab | b | a | ab | a | ||
| R28 | Q9SAZ6—Phosphoenolpyruvate carboxylase | 3.20 | * | * | a | ab | a | ab | ab | b | a | ab | ab | |
| Cell water homeostasis | ||||||||||||||
| R290 | B6T7A1—Voltage-gated potassium channel beta subunit | 2.21 | * | * | ab | c | bc | ab | ab | ab | a | abc | abc | |
| R127 | B6T634—Aquaporin PIP2.1 | 2.75 | * | ab | ab | b | ab | ab | a | ab | b | b | ||
| R193 | A0A1R3N4Y1—Aquaporin PIP2-5 | 5.52 | * | * | a | abc | c | ab | abc | ab | abc | abc | bc | |
| R285 | B6TNY0—Aquaporin TIP2.1 | 2.81 | * | * | ab | ab | b | ab | a | ab | ab | ab | ab | |
| Cell wall metabolism | ||||||||||||||
| R92 | A0A1D6IMH7—Hydroxyproline-rich glycoprotein family protein | 4.07 | * | b | a | a | ab | ab | ab | ab | a | ab | ||
| R154 | B6UD26—O-methyltransferase ZRP4 | 3.66 | * | ab | ab | ab | a | a | b | ab | ab | ab | ||
| Protein synthesis and folding | ||||||||||||||
| R58 | C0PCQ6—40S ribosomal protein S2-1 | 3.21 | * | c | a | ab | bc | abc | abc | bc | ab | abc | ||
| R255 | A0A1D6PYT7—Ribosomal protein S25 | 2.80 | * | bc | ab | ab | c | abc | a | abc | ab | ab | ||
| R309 | A0A1D6FKZ4—60S acidic ribosomal protein P2-5 | 6.66 | * | a | b | ab | ab | ab | a | ab | ab | ab | ||
| R87 | C4JA45—60S ribosomal protein L5-1 homolog b | 2.52 | * | a | b | b | ab | ab | b | ab | b | ab | ||
| R122 | B6T267—Ribosomal protein L15 | 18.7 | * | bc | ab | abc | c | abc | abc | abc | a | abc | ||
| R4 | B7ZZ42—Heat shock 70 kDa protein 3 | 2.25 | * | bc | a | abc | bc | abc | ab | c | a | ab | ||
| R100 | B6U0V6—Endoplasmin | 5.19 | * | ab | ab | ab | a | ab | a | ab | b | ab | ||
| R188 | B6SXW8—RuBisCO large subunit-binding protein sub. alpha | 3.92 | * | a | ab | ab | a | ab | a | a | b | ab | ||
| R297 | A0A1D6FAH0—T-complex protein 1 subunit epsilon | 4.66 | * | ab | ab | a | ab | ab | a | ab | b | ab | ||
| R131 | B4FZZ2—Peptidyl-prolyl cis-trans isomerase | 4.68 | * | c | a | ab | c | a | ab | bc | a | ab | ||
| R216 | A0A1D6LN79—Peptidyl-prolyl cis-trans isomerase | 15.7 | * | d | a | abc | cd | abcd | abcd | bcd | ab | abcd | ||
| R145 | A0A1D6F8L7—Coatomer subunit gamma | 3.43 | * | a | ab | ab | a | a | a | a | b | a | ||
| R205 | A0A1D6PJW1—Proteasome subunit alpha type | n.d. | * | * | a | a | a | b | a | a | a | a | a | |
| R230 | B4FB55—Ras-related protein ARA-3 | 2.40 | * | * | ab | b | b | a | ab | ab | ab | b | ab | |
| Stress response and other functions | ||||||||||||||
| R105 | C0P848—Formate dehydrogenase, mitochondrial | 14.2 | * | a | a | a | a | a | b | a | ab | ab | ||
| R99 | B8A2B4—Guanine nucleotide-binding protein beta subunit-like protein a | 2.22 | * | abc | a | c | ab | a | bc | bc | ab | bc | ||
| R112 | A5H452—Peroxidase 70 | 2.75 | * | c | a | abc | abc | abc | abc | bc | ab | ab | ||
| R133 | A0A1D6E530—Peroxidase | 3.56 | * | ab | b | ab | ab | ab | ab | a | ab | ab | ||
| R300 | B6SIU4—Peroxidase | n.d. | * | bc | ab | c | abc | ab | a | abc | abc | ab | ||
| R31 | C0PH85—Tubulin beta chain | 2.88 | * | * | d | cb | cb | c | ab | a | c | bc | abc | |
| R79 | Q6RW10—Allene oxide synthase | 2.27 | * | a | b | ab | a | ab | ab | ab | ab | ab | ||
| R225 | A0A1D6HSR3—Putative carboxylesterase 15 | n.d. | * | a | a | a | a | b | c | a | ab | b | ||
| R236 | A0A1D6QNT6—Peroxisomal fatty acid beta-oxidation multifunctional protein a | 3.86 | * | a | ab | ab | ab | ab | ab | a | b | ab | ||
| R271 | Q71RX2—Isopentenyl pyrophosphate isomerase | n.d. | * | * | ab | c | bc | a | abc | abc | a | bc | bc | |
| R272 | B6TKK2—Gibberellin receptor GID1L2 | n.d. | * | ab | ab | b | ab | ab | ab | a | a | ab | ||
| R301 | B6TP36—Prohibitin | 3.56 | * | ab | bc | abc | a | abc | abc | ab | c | abc | ||
| R318 | B6T7H0—Glutathione S-transferase 6 | n.d. | * | a | ab | b | a | a | a | a | a | a | ||
| ID | Entry—Protein Name | FC | Differences | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| V | h (n) | h (a) | h (na) | |||||||||||
| t | s | i | 06 | 30 | 54 | 06 | 30 | 54 | 06 | 30 | 54 | |||
| Nitrogen assimilation | ||||||||||||||
| L149 | B6SY01—Ferredoxin-nitrite reductase | 3.00 | * | a | ab | b | a | a | ab | ab | a | ab | ||
| L177 | B6TE43—Glutamine synthetase (chloroplast) | 3.92 | * | ab | ab | ab | ab | a | a | b | ab | a | ||
| Amino acid metabolism | ||||||||||||||
| L193 | B6TQ06—Aminomethyltransferase | 2.31 | * | ab | ab | b | ab | a | ab | ab | ab | ab | ||
| L22 | C0P5Y3—5-methyltetrahydropteroyltriglutamate -homocysteine methyltransferase 1 (MetE) | 3.57 | * | ab | a | ab | ab | ab | b | ab | ab | ab | ||
| L194 | A0A1D6KDZ0—Shikimate kinase family protein | 2.85 | * | abc | ab | c | ab | ab | c | abc | a | bc | ||
| Photosynthesis | ||||||||||||||
| L31 | P04966—Photosystem I P700 chlorophyll a apoprotein A1 | 2.48 | * | a | ab | b | a | b | ab | ab | b | ab | ||
| L85 | P04967—Photosystem I P700 chlorophyll a apoprotein A2 | 2.62 | * | a | bc | ab | ab | c | ab | ab | ab | ab | ||
| L132 | B6U534—Photosystem I reaction center subunit V | 3.85 | * | c | c | a | bc | c | a | bc | bc | ab | ||
| L196 | B4FLT7—Photosystem I H subunit1 | 2.99 | * | * | bcd | d | ab | abc | cd | a | ab | bcd | a | |
| L113 | B6SP64—Oxygen evolving enhancer protein 3 containing protein | 2.34 | * | a | a | b | a | a | b | a | a | b | ||
| L162 | B8A3D1—Pyruvate phosphate dikinase regulatory protein, chloroplastic a | 6.68 | * | a | a | ab | a | a | b | a | a | ab | ||
| L212 | B6TKB3—Triose phosphate/phosphate translocator, non-green plastid, chloroplast | n.d. | * | a | a | ab | a | a | b | a | a | b | ||
| Energy metabolism | ||||||||||||||
| L115 | B6T908—ATP synthase B chain | 8.19 | * | * | ab | abc | c | a | ab | abc | ab | abc | bc | |
| L225 | B6SP77—ATP synthase C chain | 77.8 | * | a | a | b | a | a | ab | a | a | ab | ||
| L155 | B6T168—ATP synthase epsilon chain | 3.09 | * | a | abc | bc | a | abc | bc | abc | ab | c | ||
| L130 | B4FK49—Nucleoside diphosphate kinase | 4.88 | * | * | a | ab | abc | abc | abc | bc | abc | ab | c | |
| L236 | C0HHC4—Nucleoside diphosphate kinase | 7.24 | * | a | a | b | a | a | b | a | a | b | ||
| Carbon metabolism | ||||||||||||||
| L192 | A0A1D6N8I0—2,3-bisphosphoglycerate-independent phosphoglycerate mutase 1 | 4.77 | * | ab | a | ab | ab | ab | ab | ab | ab | b | ||
| L191 | B8A0W7—Enolase 1 a | 13.1 | * | a | a | abc | ab | a | bc | a | a | c | ||
| L110 | B4FZU8—Malate dehydrogenase | 2.31 | * | a | a | b | a | a | b | a | a | b | ||
| Protein synthesis and folding | ||||||||||||||
| L69 | A0A0B4J303—Eukaryotic initiation factor4a | 3.01 | * | abc | a | abc | ab | ab | bc | ab | a | c | ||
| L111 | A0A1D6LAB8—RNA helicase4 | 3.98 | * | a | a | a | a | a | a | a | a | b | ||
| L205 | A0A1X7YHC0—30S ribosomal protein S2, chloroplastic a | n.d. | * | ab | ab | a | ab | ab | a | b | ab | a | ||
| L127 | C0PEC4—30S ribosomal protein S5 chloroplastic | 4.70 | * | ab | ab | b | ab | a | b | ab | ab | ab | ||
| L211 | P08530—30S ribosomal protein S8, chloroplastic | 3.27 | * | ab | a | bc | ab | ab | bc | ab | a | c | ||
| L137 | B4FR40—30S ribosomal protein S9 chloroplastic | n.d. | * | ab | a | a | ab | b | a | ab | ab | a | ||
| L145 | B6UGL6—40S ribosomal protein S5 | 2.27 | * | * | a | a | b | a | a | b | a | a | b | |
| L118 | A0A1D6P3R8—40S ribosomal protein S14 | 2.15 | * | ab | a | ab | ab | b | ab | a | ab | ab | ||
| L88 | B6SNQ7—40S ribosomal protein S16 | n.d. | * | a | a | ab | a | a | b | ab | a | ab | ||
| L171 | B6UF84—50S ribosomal protein L3 | 2.23 | * | ab | a | ab | ab | ab | ab | ab | ab | b | ||
| L195 | B6SK79—60S ribosomal protein L4 | 3.28 | * | ab | a | ab | ab | ab | ab | ab | b | b | ||
| L216 | B6SHW0—60S ribosomal protein L6 | 2.33 | * | a | a | ab | ab | a | ab | a | ab | b | ||
| L126 | A0A1D6GM13—60S ribosomal protein L10 a | 2.40 | * | ab | a | ab | ab | ab | ab | ab | a | b | ||
| L239 | B6TM00—60 ribosomal protein L14 | 2.66 | * | * | ab | a | bc | bc | bc | bc | ab | ab | c | |
| L240 | B6SJH2—60S ribosomal protein L17 | 2.65 | * | a | a | b | a | a | b | a | a | b | ||
| L15 | A0A1X7YIM9—RuBisCO large subunit-binding protein subunit alpha, chloroplastic a | 2.10 | * | c | a | bc | c | abc | c | c | ab | c | ||
| L19 | B7ZZ42—Heat shock 70 kDa protein 3 | 2.73 | * | ab | a | ab | ab | b | ab | b | ab | b | ||
| L64 | C3UZ63—HSP protein | 6.72 | * | abc | a | cb | abc | abc | bc | ab | abc | c | ||
| L70 | A0A1D6LIK1—Peptidyl-prolyl cis-trans isomerase | 2.04 | * | d | abcd | a | cd | abc | ab | bcd | abcd | a | ||
| L204 | B4FZZ2—Peptidyl-prolyl cis-trans isomerase | n.d. | * | b | a | a | c | a | a | bc | a | a | ||
| L243 | A0A1D6FAW5—Peptidyl-prolyl cis-trans isomerase | n.d. | * | b | a | a | b | a | a | b | a | a | ||
| L154 | B6TCN7—Threonine endopeptidase | 3.58 | * | a | ab | abc | a | a | c | ab | a | bc | ||
| Stress response and other functions | ||||||||||||||
| L76 | A1XCI5—Lipoxygenase | 4.44 | * | * | a | a | a | a | a | b | a | a | ab | |
| L189 | B6T2W9—Thylakoid lumenal 19 kDa protein | 2.84 | * | b | a | ab | a | ab | ab | ab | a | ab | ||
| L201 | B6SU36—Thylakoid lumenal 16.5 kDa protein | 10.3 | * | a | ab | b | ab | a | ab | ab | ab | ab | ||
| L143 | B6T7W8—Cortical cell-delineating protein | 2.57 | * | ab | a | b | ab | ab | b | ab | a | b | ||
| L180 | K7UGI3—Putative thioredoxin superfamily protein | 4.52 | * | abc | a | bcd | abcd | ab | cd | ab | a | d | ||
| L208 | C4J9Y2—Aluminum-induced protein homolog1 | 10.5 | * | ab | a | b | a | a | ab | ab | a | ab | ||
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Prinsi, B.; Espen, L. Time-Course of Metabolic and Proteomic Responses to Different Nitrate/Ammonium Availabilities in Roots and Leaves of Maize. Int. J. Mol. Sci. 2018, 19, 2202. https://doi.org/10.3390/ijms19082202
Prinsi B, Espen L. Time-Course of Metabolic and Proteomic Responses to Different Nitrate/Ammonium Availabilities in Roots and Leaves of Maize. International Journal of Molecular Sciences. 2018; 19(8):2202. https://doi.org/10.3390/ijms19082202
Chicago/Turabian StylePrinsi, Bhakti, and Luca Espen. 2018. "Time-Course of Metabolic and Proteomic Responses to Different Nitrate/Ammonium Availabilities in Roots and Leaves of Maize" International Journal of Molecular Sciences 19, no. 8: 2202. https://doi.org/10.3390/ijms19082202