Comparison of Reliable Reference Genes Following Different Hormone Treatments by Various Algorithms for qRT-PCR Analysis of Metasequoia
Abstract
1. Introduction
2. Results
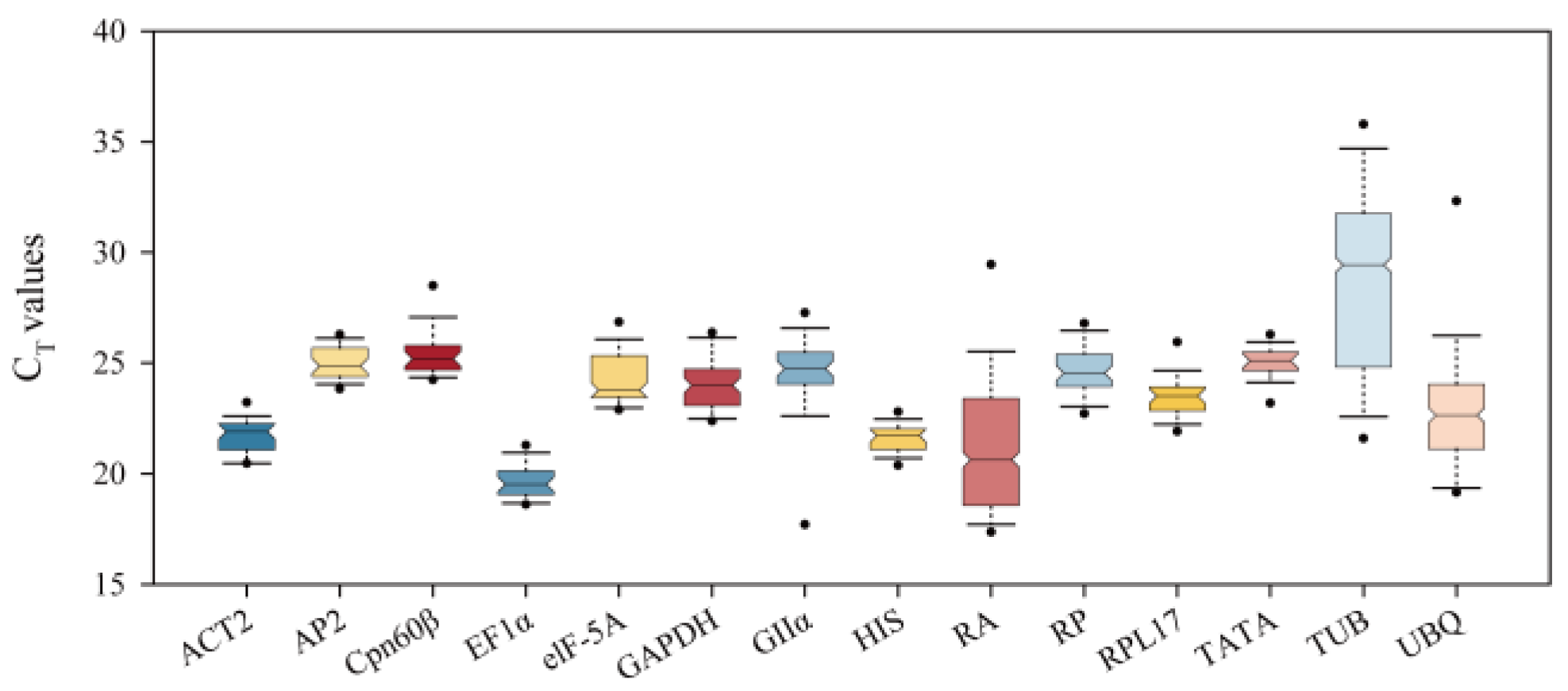
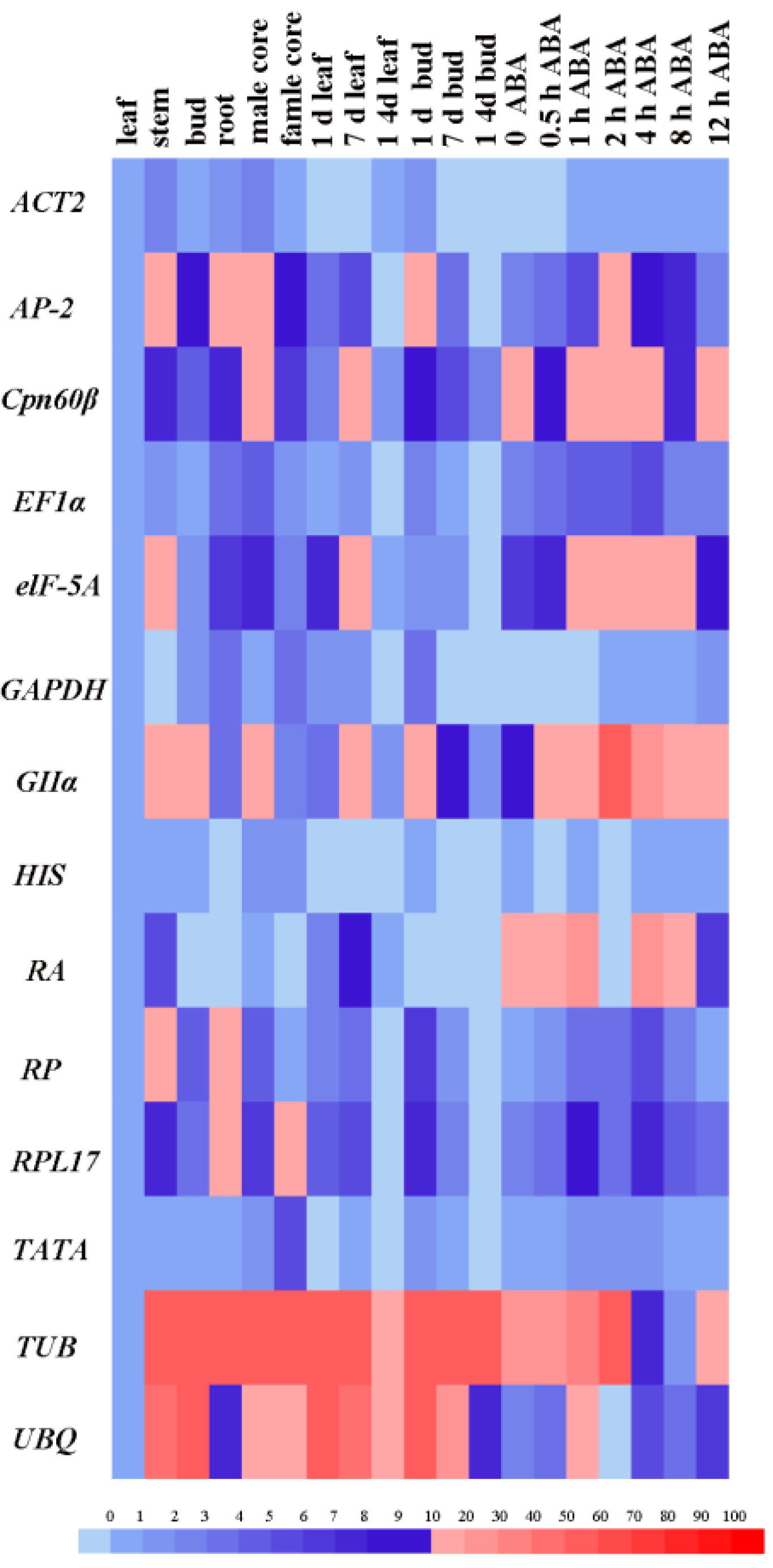
2.1. Primer Specificity and Expression Analysis of 14 Candidate Reference Genes and Two Target Genes
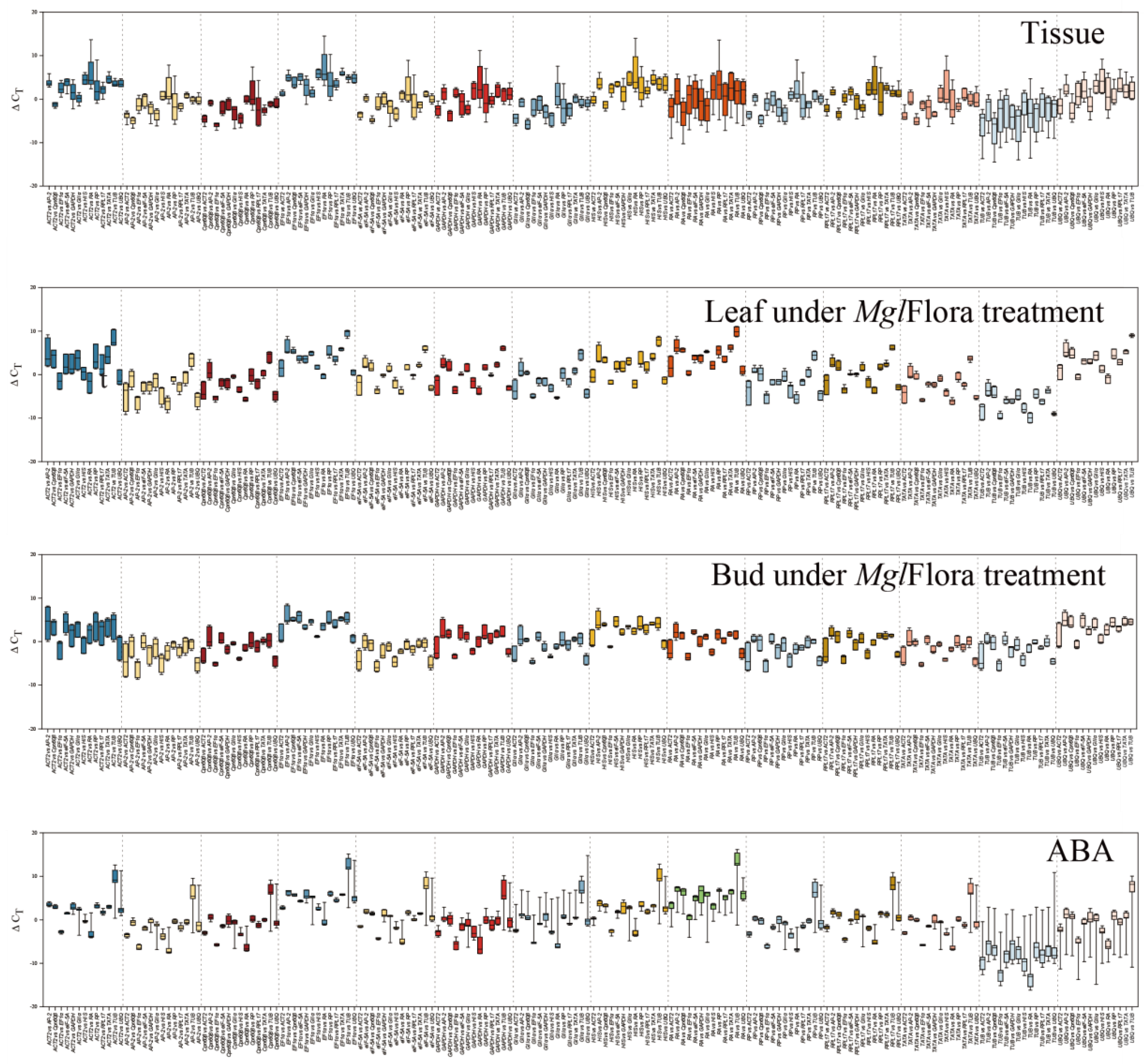
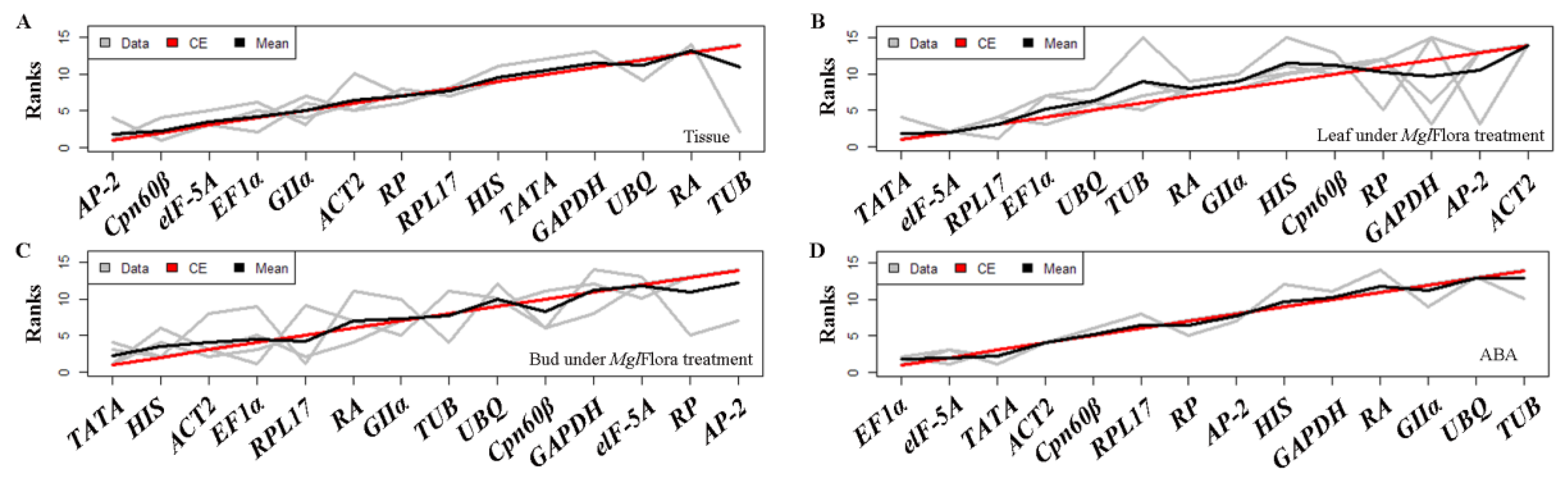
2.2. Analysis of Gene Expression Stability
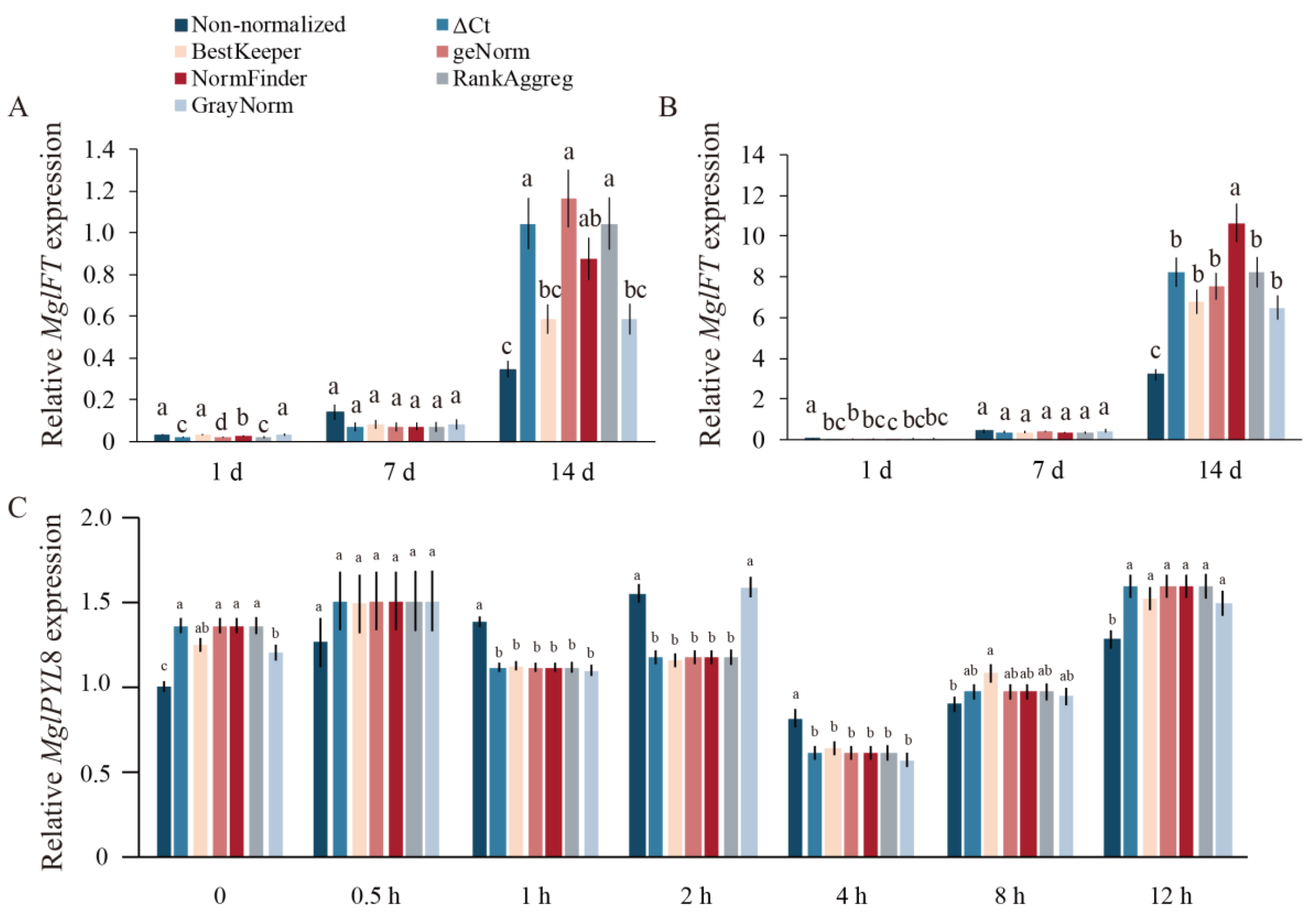
2.3. Validation of Selected Candidate Reference Genes
3. Discussion
4. Materials and Methods
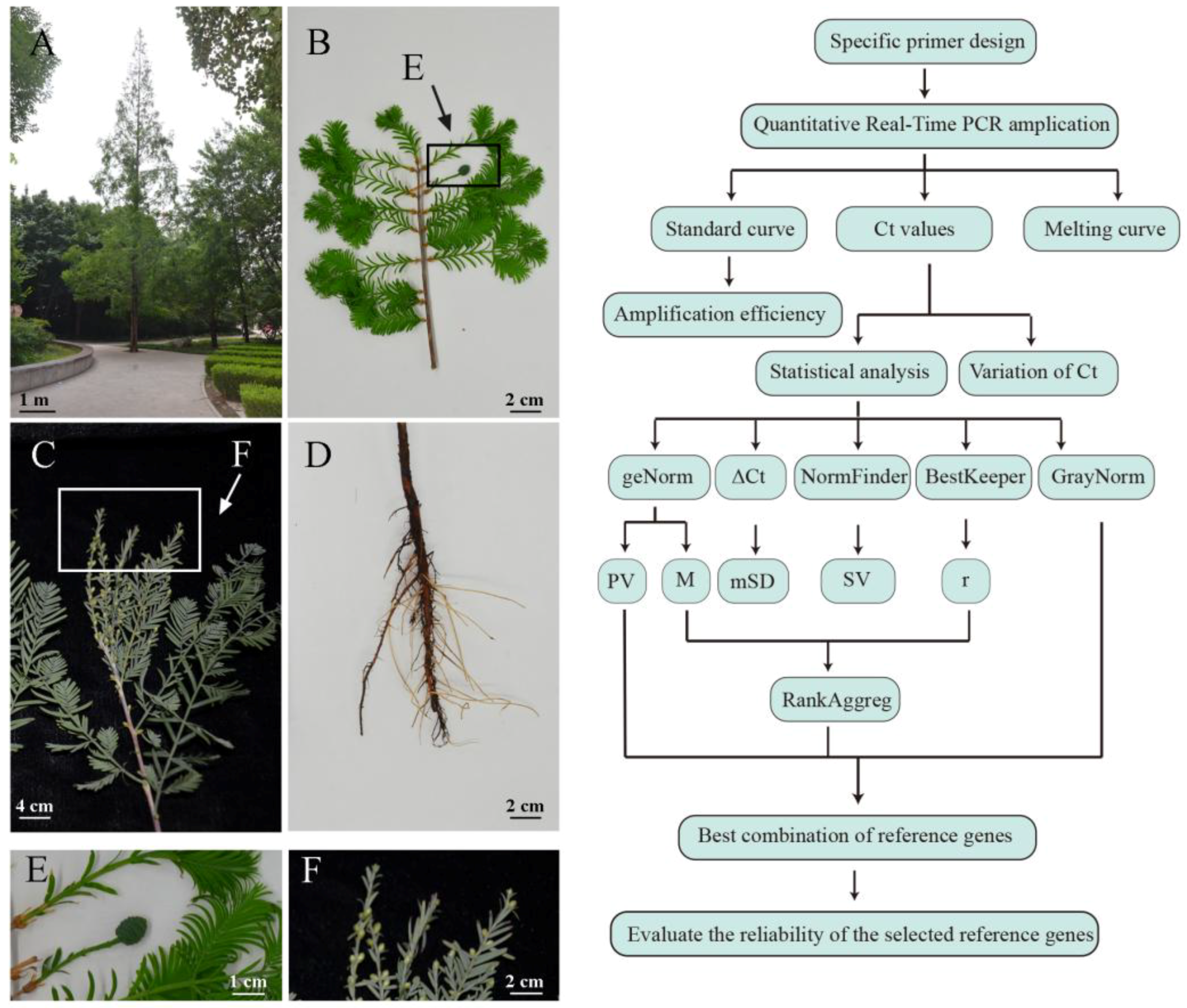
4.1. Hormone Treatment and Sample Collection
4.2. Selection of Candidate Reference Genes and Primer Design
4.3. Total RNA Isolation and cDNA Synthesis
4.4. qRT-PCR Conditions and Amplification Efficiency
4.5. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Huggett, J.; Dheda, K.; Bustin, S.; Zumla, A. Real-time RT-PCR normalisation; strategies and considerations. Genes Immun. 2005, 6, 279–284. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Bustin, S.A. Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): Trends and problems. J. Mol. Endocrinol. 2002, 29, 23–39. [Google Scholar] [CrossRef] [PubMed]
- Chao, J.; Yang, S.; Chen, Y.; Tian, W.-M. Evaluation of Reference Genes for Quantitative Real-Time PCR Analysis of the Gene Expression in Laticifers on the Basis of Latex Flow in Rubber Tree (Hevea brasiliensis Muell Arg). Front. Plant Sci. 2016, 7, 1149. [Google Scholar] [CrossRef] [PubMed]
- Yeap, W.C.; Jia, M.L.; Wong, Y.C.; Kulaveerasingam, H. Evaluation of suitable reference genes for qRT-PCR gene expression normalization in reproductive, vegetative tissues and during fruit development in oil palm. Plant Cell Tissue Organ Cult. 2014, 116, 55–66. [Google Scholar] [CrossRef]
- James, W.; Sophia, N.; Aneta, I.; Reena, N. Defining reference genes in Oryza sativa using organ, development, biotic and abiotic transcriptome datasets. BMC Plant Boil. 2010, 10, 56. [Google Scholar]
- Quackenbush, J. Microarray data normalization and transformation. Nat. Genet. 2002, 32, 496–501. [Google Scholar] [CrossRef]
- Czechowski, T.; Stitt, M.; Altmann, T.; Udvardi, M.K.; Scheible, W.R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 2005, 139, 5–17. [Google Scholar] [CrossRef]
- Wang, H.L.; Chen, J.; Tian, Q.; Wang, S.; Xia, X.; Yin, W. Identification and validation of reference genes for Populus euphratica gene expression analysis during abiotic stresses by quantitative real-time PCR. Physiol. Plant. 2014, 152, 529–545. [Google Scholar] [CrossRef]
- Wang, H.L.; Li, L.; Tang, S.; Yuan, C.; Tian, Q.Q.; Su, Y.; Li, H.G.; Zhao, L.; Yin, W.; Zhao, R.; et al. Evaluation of Appropriate Reference Genes for Reverse Transcription-Quantitative PCR Studies in Different Tissues of a Desert Poplar via Comparision of Different Algorithms. IJMS 2015, 16, 20468–20491. [Google Scholar] [CrossRef]
- Exposito-Rodriguez, M.; Borges, A.A.; Borges-Perez, A.; Perez, J.A. Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol. 2008, 8, 131. [Google Scholar] [CrossRef]
- Nicot, N.; Hausman, J.F.; Hoffmann, L.; Evers, D. Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J. Exp. Bot. 2005, 56, 2907–2914. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, L.A.; Breton, M.C.; Bastolla, F.M.; Camargo, S.D.; Margis, R.; Frazzon, J.; Pasquali, G. Reference Genes for the Normalization of Gene Expression in Eucalyptus Species. Plant Cell Physiol. 2012, 53, 405–422. [Google Scholar] [CrossRef] [PubMed]
- de Lima, J.C.; de Costa, F.; Fuller, T.N.; Rodrigues-Correa, K.C.D.; Kerber, M.R.; Lima, M.S.; Fett, J.P.; Fett-Neto, A.G. Reference Genes for qPCR Analysis in Resin-Tapped Adult Slash Pine as a Tool to Address the Molecular Basis of Commercial Resinosis. Front. Plant Sci. 2016, 7, 849. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Fu, Y.; Ban, L.; Wang, Z.; Feng, G.; Li, J.; Gao, H. Selection of reliable reference genes for quantitative real-time RT-PCR in alfalfa. Genes Genet. Syst. 2015, 90, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Sadritdinova, A.F.; Snezhkina, A.V.; Dmitriev, A.A.; Krasnov, G.S.; Astakhova, L.N.; Kudryavtsev, A.A.; Mel’nikova, N.V.; Speranskaya, A.S.; Darii, M.V.; Lakunina, V.A.; et al. A new reference gene, Ef1A, for quantitative real-time PCR assay of the starfish Asterias rubens pyloric ceca. Dokl. Biol. Sci. 2013, 452, 310–312. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.R.; Nam, H.Y.; Kim, S.U.; Kim, S.I.; Chang, Y.J. Normalization of reverse transcription quantitative-PCR with housekeeping genes in rice. Biotechnol. Lett. 2003, 25, 1869–1872. [Google Scholar] [CrossRef] [PubMed]
- Goidin, D.; Mamessier, A.; Staquet, M.J.; Schmitt, D.; Berthiervergnes, O. Ribosomal 18S RNA prevails over glyceraldehyde-3-phosphate dehydrogenase and beta-actin genes as internal standard for quantitative comparison of mRNA levels in invasive and noninvasive human melanoma cell subpopulations. Anal. Biochem. 2001, 295, 17–21. [Google Scholar] [CrossRef]
- Gutierrez, L.; Mauriat, M.; Pelloux, J.; Bellini, C.; van Wuytswinkel, O. Towards a systematic validation of references in real-time RT-PCR. Plant Cell 2008, 20, 1734–1735. [Google Scholar] [CrossRef]
- Gutierrez, L.; Mauriat, M.; Guenin, S.; Pelloux, J.; Lefebvre, J.F.; Louvet, R.; Rusterucci, C.; Moritz, T.; Guerineau, F.; Bellini, C.; et al. The lack of a systematic validation of reference genes: A serious pitfall undervalued in reverse transcription-polymerase chain reaction (RT-PCR) analysis in plants. Plant Biotechnol. J. 2008, 6, 609–618. [Google Scholar] [CrossRef]
- Yperman, J.; De Visscher, G.; Holvoet, P.; Flameng, W. Beta-actin cannot be used as a control for gene expression in ovine interstitial cells derived from heart valves. J. Heart Valve Dis. 2004, 13, 848–853. [Google Scholar]
- de Andrade, L.M.; Brito, M.D.; Peixoto, R.F.; Marchiori, P.E.R.; Nobile, P.M.; Martins, A.P.B.; Ribeiro, R.V.; Creste, S. Reference genes for normalization of qPCR assays in sugarcane plants under water deficit. Plant Methods 2017, 13, 28. [Google Scholar] [CrossRef]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, research0034-1. [Google Scholar] [CrossRef] [PubMed]
- Andersen, C.L.; Jensen, J.L.; Orntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Silver, N.; Best, S.; Jiang, J.; Thein, S.L. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR. BMC Mol. Biol. 2006, 7, 33. [Google Scholar] [CrossRef] [PubMed]
- Remans, T.; Keunen, E.; Bex, G.J.; Smeets, K.; Vangronsveld, J.; Cuypers, A. Reliable Gene Expression Analysis by Reverse Transcription-Quantitative PCR: Reporting and Minimizing the Uncertainty in Data Accuracy. Plant Cell 2014, 26, 3829–3837. [Google Scholar] [CrossRef] [PubMed]
- Pihur, V.; Datta, S.; Datta, S. RankAggreg, an R package for weighted rank aggregation. BMC Bioinform. 2009, 10, 62. [Google Scholar] [CrossRef]
- Brunsfeld, S.J.; Soltis, P.; Soltis, D.; Gadek, P.A.; Quinn, C.J.; Strenge, D.; Ranker, T. Phylogenetic Relationships Among the Genera of Taxodiaceae and Cupressaceae: Evidence from rbcL Sequences. Syst. Bot. 1994, 19, 253–262. [Google Scholar] [CrossRef]
- Hu, S.Y. The Metasequoia flora and its phytogeographic significance. J. Arnold Arbor. 1980, 61, 41–94. [Google Scholar] [CrossRef]
- Charles Nelson, E. Metasequoia glyptostroboides, the dawn redwood: Some Irish glosses on its discovery and introduction into cultivation. Curtis’s Bot. Mag. 2004, 15, 77–80. [Google Scholar] [CrossRef]
- Kunzmann, L.; Mai, D.H. The first record of fossil Metasequoia (Cupressaceae) from continental Europe. Rev. Palaeobot. Palynol. 2011, 164, 247–250. [Google Scholar] [CrossRef]
- Polman, J.E.; Michon, S.G.L.; Militz, H.; Helmink, A.T.F. The wood of Metasequoia glyptostroboides (Hu et Cheng) of Dutch origin. Holz Als Roh-Und Werkst. 1999, 57, 215–221. [Google Scholar] [CrossRef]
- Toth, J.; Hethelyi, E.B.; Haznagy-Radnai, E.; Boszormenyi, A.; Czigle, S. Living fossil volatile compounds analysis using SPME GC-MS: Examples of Sequoia, Sequoiadendron and Metasequoia. Planta Medica 2016, 82, P301. [Google Scholar] [CrossRef]
- Bajpai, V.K.; Rather, I.A.; Kang, S.C.; Park, Y.H. A Diterpenoid Taxoquinone from Metasequoia glyptostroboides with Pharmacological Potential. Indian J. Pharm. Educ. Res. 2016, 50, 458–464. [Google Scholar] [CrossRef]
- Longman, K.A. Initiation of Flowering on First Year Cuttings of Metasequoia glyptostroboides Hu and Cheng. Nature 1970, 227, 299. [Google Scholar] [CrossRef]
- Qiu, Z.; Wan, L.; Chen, T.; Wan, Y.; He, X.; Lu, S.; Wang, Y.; Lin, J. The regulation of cambial activity in Chinese fir (Cunninghamia lanceolata) involves extensive transcriptome remodeling. New Phytol. 2013, 199, 708–719. [Google Scholar] [CrossRef]
- Hu, Y.; Deng, T.; Chen, L.; Wu, H.; Zhang, S. Selection and Validation of Reference Genes for qRT-PCR in Cycas elongata. PLoS ONE 2016, 11, e0154384. [Google Scholar] [CrossRef] [PubMed]
- Behringer, D.; Zimmermann, H.; Ziegenhagen, B.; Liepelt, S. Differential Gene Expression Reveals Candidate Genes for Drought Stress Response in Abies alba (Pinaceae). PLoS ONE 2015, 10, e0124564. [Google Scholar] [CrossRef] [PubMed]
- Goncalves, S.; Cairney, J.; Maroco, J.; Oliveira, M.M.; Miguel, C. Evaluation of control transcripts in real-time RT-PCR expression analysis during maritime pine embryogenesis. Planta 2005, 222, 556–563. [Google Scholar] [CrossRef] [PubMed]
- Ranjan, A.; Ansari, S.A.; Srivastava, R.; Mantri, S.; Asif, M.H.; Sawant, S.V.; Tuli, R. A T9G mutation in the prototype TATA-box TCACTATATATAG determines nucleosome formation and synergy with upstream activator sequences in plant promoters. Plant Physiol. 2009, 151, 2174–2186. [Google Scholar] [CrossRef] [PubMed]
- Eissa, N.; Hussein, H.; Wang, H.; Rabbi, M.F.; Bernstein, C.N.; Ghia, J.E. Stability of Reference Genes for Messenger RNA Quantification by Real-Time PCR in Mouse Dextran Sodium Sulfate Experimental Colitis. PLoS ONE 2016, 11, e0156289. [Google Scholar] [CrossRef] [PubMed]
- Engdahl, E.; Dunn, N.; Fogdell-Hahn, A. Investigation of reference gene expression during human herpesvirus 6B infection indicates peptidylprolyl isomerase A as a stable reference gene and TATA box binding protein as a gene up-regulated by this virus. J. Virol. Methods 2016, 227, 47–49. [Google Scholar] [CrossRef] [PubMed]
- Bansal, R.; Mamidala, P.; Mian, M.A.; Mittapalli, O.; Michel, A.P. Validation of reference genes for gene expression studies in Aphis glycines (Hemiptera: Aphididae). J. Econ. Entomol. 2012, 105, 1432–1438. [Google Scholar] [CrossRef] [PubMed]
- Rudus, I.; Kepczynski, J. Reference gene selection for molecular studies of dormancy in wild oat (Avena fatua L.) caryopses by RT-qPCR method. PLoS ONE 2018, 13, e0192343. [Google Scholar] [CrossRef] [PubMed]
- Chang, E.; Shi, S.; Liu, J.; Cheng, T.; Xue, L.; Yang, X.; Yang, W.; Lan, Q.; Jiang, Z. Selection of reference genes for quantitative gene expression studies in Platycladus orientalis (Cupressaceae) Using real-time PCR. PLoS ONE 2012, 7, e33278. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, J.; Xu, L.; Lai, H.; Chen, Y.; Yang, Z.; Huang, B. Identification and Validation of Reference Genes for Seashore Paspalum Response to Abiotic Stresses. Int. J. Mol. Sci. 2017, 18, 1322. [Google Scholar] [CrossRef] [PubMed]
- Sang, J.; Han, X.; Liu, M.; Qiao, G.; Jiang, J.; Zhuo, R. Selection and validation of reference genes for real-time quantitative PCR in hyperaccumulating ecotype of Sedum alfredii under different heavy metals stresses. PLoS ONE 2013, 8, e82927. [Google Scholar] [CrossRef]
- Mallona, I.; Lischewski, S.; Weiss, J.; Hause, B.; Egea-Cortines, M. Validation of reference genes for quantitative real-time PCR during leaf and flower development in Petunia hybrida. BMC Plant Biol. 2010, 10, 4. [Google Scholar] [CrossRef]
- Wan, Q.; Chen, S.; Shan, Z.; Yang, Z.; Chen, L.; Zhang, C.; Yuan, S.; Hao, Q.; Zhang, X.; Qiu, D.; et al. Stability evaluation of reference genes for gene expression analysis by RT-qPCR in soybean under different conditions. PLoS ONE 2017, 12, e0189405. [Google Scholar] [CrossRef]
- Silveira, E.D.; Alves-Ferreira, M.; Guimaraes, L.A.; da Silva, F.R.; Carneiro, V.T. Selection of reference genes for quantitative real-time PCR expression studies in the apomictic and sexual grass Brachiaria brizantha. BMC Plant Biol. 2009, 9, 84. [Google Scholar] [CrossRef]
- Zhao, Y.; Liang, H.; Li, L.; Tang, S.; Han, X.; Wang, C.; Xia, X.; Yin, W. Digital gene expression analysis of male and female bud transition in Metasequoia reveals high activity of MADS-box transcription factors and hormone-mediated sugar pathways. Front. Plant Sci. 2015, 6, 467. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Thammannagowda, S.; Staton, M.; Tang, S.; Xia, X.; Yin, W.; Liang, H. An EST dataset for Metasequoia glyptostroboides buds: The first EST resource for molecular genomics studies in Metasequoia. Planta 2013, 237, 755–770. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.; Puryear, J.; Cairney, J. A simple and efficient method for isolating RNA from pine trees. Plant Mol. Boil. Rep. 1993, 11, 113–116. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pair-wise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef] [PubMed]

| Gene Abbreviation | Primer Sequence of Forward (5′–3′) | Primer Sequence of Reward (5′–3′) | Amplicon Length (bp) | Tm(°C) | E | Arabidopsis Ortholog No. |
|---|---|---|---|---|---|---|
| ACT2 | GACGCTTATGTTGGTGATGAGGC | GAGTCATCTTCTCTCTGTTTGCCTTAGG | 211 | 58.2 °C/58.4 ºC | 1.9807 | AT5G09810.1 |
| AP-2 | GGAAGTGTTCTCCGGTGCGATG | CAGTTCAAATTCTCCATCAGGTGGGAC | 252 | 60.3 °C/59.3 °C | 2.1419 | AT5G46630.2 |
| Cpn60β | GTGATCGCGCCAGAATGGCATC | CCGACCATAACTTTGGCAGCAGG | 196 | 61.1 °C/60.5 °C | 1.9454 | AT5G56500.1 |
| EF1α | GCTTGGGTGCTTGACAAGCTCAAG | CAGAGCATGTTCTCGGGTCTGTCC | 251 | 60.8 °C/61.2 °C | 1.9384 | AT5G60390.1 |
| elF-5A | GTCGGATGAGGAGCACCATTTTGATCAC | CAGTTGTGGGATGAAGGAACTATATCCTCG | 245 | 61.1 °C/59.7 °C | 2.0517 | AT1G13950.1 |
| GAPDH | GATGATGTCGAGCTCGTTGCAGTGAAC | GATTCAACCACATACTCTGCACCAACC | 224 | 61.3 °C/59.7 °C | 2.0367 | AT1G13440.1 |
| GIIα | CGGTCCCCAGGCTGTTAGTTTAGATG | CGTCGTCGACTCCTTGGAATGAGAG | 229 | 60.8 °C/60.6 °C | 1.9042 | AT5G63840.1 |
| HIS | CACAGATACCGTCCCGGAACTGTTG | GCTTCTGCAGCTTCCTGGAGAGC | 179 | 61.1 °C/62.2 °C | 2.0234 | AT4G40030.2 |
| RA | GATGAGTGCGGGAGAGCTTGAAAGTG | GATGAGTGCGGGAGAGCTTGAAAGTG | 138 | 61.4 °C/61.4 °C | 1.9410 | AT2G39730.1 |
| RP | GGTCACTGCCTCGTCGCAG | GCCTTCAGATCCACATCCAATGTGTG | 164 | 61 °C/59.9 °C | 1.8518 | AT3G22230.1 |
| RPL17 | GAGGCAGCCAATGGCACTCATC | CAACCTGGTTGAAGGTCTTCCCATTG | 167 | 60.9 °C/59.9 °C | 1.9946 | AT1G04270.1 |
| TATA | GGAAGGGAGTCAGCCTGTCGATCTG | GCACCTGTGCAGACCATCTTTCCTGAT | 229 | 62.5 °C/62.6 °C | 1.9278 | AT1G55520.1 |
| TUB | GTTTGAGGTTCCCTGGTCAGCTC | CTGCCATGTCTTGGATCAGCAGC | 211 | 59.8 °C/60.6 °C | 2.0385 | AT5G23860 |
| UBQ | CGGCCGTACTCTTGCCGAC | GGCCTTGACGTTGTCGATGGTG | 157 | 61 °C/60.7 °C | 1.9534 | AT5G20620.1 |
| FT | GTCACGAGATCCACTAACGACGGG | GCTTGAGCTCGCAGCCATTTTTG | 125 | 60.8 °C/59.7 °C | 1.8475 | AT1G65480.1 |
| PYL8 | CAGCGACAGCTAGCGAAGAGAG | CCATTGTACCTGGCCTCCCATC | 154 | 59.5 °C/59.8 °C | 1.9692 | AT5G53160.2 |
| Rank | Total | mSD | Tissue | mSD | LEAF of MglFlora | mSD | Bud of MglFlora | mSD | ABA | mSD |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | EF1α | 0.522 | Cpn60β | 1.41 | TATA | 0.75 | TATA | 0.74 | elF-5A | 1.18 |
| 2 | Cpn60β | 0.522 | EF1α | 1.43 | elF-5A | 0.77 | ACT2 | 0.78 | EF1α | 1.19 |
| 3 | TATA | 0.698 | elF-5A | 1.43 | RPL17 | 0.81 | EF1α | 0.83 | TATA | 1.19 |
| 4 | HIS | 0.761 | AP-2 | 1.44 | EF1α | 0.82 | HIS | 0.84 | ACT2 | 1.21 |
| 5 | RPL17 | 0.806 | ACT2 | 1.53 | GAPDH | 0.84 | RPL17 | 0.86 | Cpn60β | 1.26 |
| 6 | ACT2 | 0.884 | GIIα | 1.58 | UBQ | 0.87 | RA | 0.86 | RPL17 | 1.27 |
| 7 | RP | 0.949 | RPL17 | 1.6 | RA | 0.92 | GIIα | 0.89 | RP | 1.29 |
| 8 | elF-5A | 1.006 | RP | 1.66 | GIIα | 0.93 | TUB | 0.99 | AP-2 | 1.32 |
| 9 | AP-2 | 1.049 | HIS | 1.69 | TUB | 0.94 | UBQ | 1.02 | HIS | 1.38 |
| 10 | GAPDH | 1.118 | TATA | 1.75 | HIS | 1.09 | Cpn60β | 1.03 | GAPDH | 1.76 |
| 11 | GIIα | 1.267 | GAPDH | 1.92 | Cpn60β | 1.09 | GAPDH | 1.12 | RA | 2.4 |
| 12 | UBQ | 1.596 | UBQ | 2.37 | RP | 1.16 | elF-5A | 1.12 | GIIα | 2.8 |
| 13 | RA | 1.869 | RA | 3.93 | AP-2 | 1.68 | RP | 1.18 | UBQ | 3.79 |
| 14 | TUB | 2.189 | TUB | 4.01 | ACT2 | 2.23 | AP-2 | 1.8 | TUB | 4.32 |
| Rank | Total | r | Tissue | r | Leaf of MglFlora | r | Bud of MglFlora | r | ABA | r |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | HIS | 0.944 | AP-2 | 0.969 | HIS | 1.000 | Cpn60β | 1.000 | TATA | 0.989 |
| 2 | TATA | 0.913 | HIS | 0.960 | ACT2 | 0.996 | GIIα | 0.999 | Cpn60β | 0.964 |
| 3 | ACT2 | 0.889 | TATA | 0.927 | TUB | 0.994 | HIS | 0.998 | elF-5A | 0.936 |
| 4 | Cpn60β | 0.834 | EF1α | 0.927 | TATA | 0.99 | EF1α | 0.998 | EF1α | 0.923 |
| 5 | RPL17 | 0.818 | ACT2 | 0.903 | EF1α | 0.989 | TATA | 0.998 | HIS | 0.909 |
| 6 | EF1α | 0.800 | Cpn60β | 0.808 | Cpn60β | 0.975 | ACT2 | 0.997 | ACT2 | 0.845 |
| 7 | GAPDH | 0.750 | GAPDH | 0.788 | UBQ | 0.974 | GAPDH | 0.993 | RPL17 | 0.794 |
| 8 | AP-2 | 0.648 | GIIα | 0.777 | GIIα | 0.938 | RA | 0.985 | AP-2 | 0.687 |
| 9 | GIIα | 0.613 | RP | 0.776 | elF-5A | 0.931 | elF-5A | 0.984 | RP | 0.658 |
| 10 | RP | 0.588 | RPL17 | 0.763 | GAPDH | 0.902 | RPL17 | 0.977 | GAPDH | 0.491 |
| 11 | elF-5A | 0.503 | elF-5A | 0.442 | RA | 0.897 | TUB | 0.960 | RA | 0.452 |
| 12 | UBQ | 0.426 | TUB | 0.363 | RPL17 | 0.874 | UBQ | 0.956 | GIIα | 0.153 |
| 13 | TUB | 0.276 | UBQ | 0.158 | RP | 0.79 | RP | 0.908 | UBQ | 0.001 |
| 14 | RA | 0.138 | RA | 0.130 | AP-2 | 0.001 | AP-2 | 0.856 | TUB | 0.001 |
| Rank | Total | M | Tissue | M | Leaf of MglFlora | M | Bud of MglFlora | M | ABA | M |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | EF1α | 0.522 | AP-2 | 0.487 | RPL17 | 0.105 | EF1α | 0.19 | EF1α | 0.139 |
| 2 | Cpn60β | 0.522 | Cpn60β | 0.487 | elF-5A | 0.105 | HIS | 0.19 | TATA | 0.139 |
| 3 | TATA | 0.698 | elF-5A | 0.542 | GAPDH | 0.147 | ACT2 | 0.243 | elF-5A | 0.155 |
| 4 | HIS | 0.761 | EF1α | 0.622 | TATA | 0.284 | TATA | 0.294 | ACT2 | 0.195 |
| 5 | RPL17 | 0.806 | ACT2 | 0.692 | TUB | 0.377 | GIIα | 0.34 | Cpn60β | 0.227 |
| 6 | ACT2 | 0.884 | RP | 0.772 | UBQ | 0.406 | Cpn60β | 0.371 | RPL17 | 0.296 |
| 7 | RP | 0.949 | GIIα | 0.823 | EF1α | 0.472 | RA | 0.449 | RP | 0.342 |
| 8 | elF-5A | 1.006 | RPL17 | 0.88 | RA | 0.55 | GAPDH | 0.535 | AP-2 | 0.376 |
| 9 | AP-2 | 1.049 | HIS | 0.964 | GIIα | 0.592 | RPL17 | 0.627 | HIS | 0.428 |
| 10 | GAPDH | 1.118 | TATA | 1.018 | Cpn60β | 0.648 | UBQ | 0.694 | GAPDH | 0.564 |
| 11 | GIIα | 1.267 | GAPDH | 1.079 | HIS | 0.689 | TUB | 0.756 | RA | 0.816 |
| 12 | UBQ | 1.596 | UBQ | 1.258 | RP | 0.75 | elF-5A | 0.81 | GIIα | 1.113 |
| 13 | RA | 1.869 | RA | 1.645 | AP-2 | 0.87 | RP | 0.872 | UBQ | 1.48 |
| 14 | TUB | 2.189 | TUB | 1.983 | ACT2 | 1.065 | AP-2 | 1.005 | TUB | 1.885 |
| Rank | Total | SV | Tissue | SV | Leaf of MglFlora | SV | Bud of MglFlora | SV | ABA | SV |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | RPL17 | 0.428 | AP-2 | 0.197 | TATA | 0.152 | TATA | 0.233 | TATA | 0.070 |
| 2 | EF1α | 0.448 | Cpn60β | 0.282 | elF-5A | 0.210 | RPL17 | 0.366 | EF1α | 0.070 |
| 3 | Cpn60β | 0.478 | elF-5A | 0.352 | EF1α | 0.249 | ACT2 | 0.389 | elF-5A | 0.080 |
| 4 | RP | 0.582 | GIIα | 0.454 | RPL17 | 0.355 | RA | 0.435 | ACT2 | 0.085 |
| 5 | TATA | 0.652 | EF1α | 0.582 | UBQ | 0.365 | EF1α | 0.581 | Cpn60β | 0.120 |
| 6 | HIS | 0.670 | ACT2 | 0.683 | GAPDH | 0.415 | HIS | 0.613 | RPL17 | 0.177 |
| 7 | ACT2 | 0.782 | RP | 0.810 | TUB | 0.527 | GIIα | 0.663 | RP | 0.177 |
| 8 | AP-2 | 0.921 | RPL17 | 0.862 | RA | 0.561 | TUB | 0.673 | AP-2 | 0.184 |
| 9 | elF-5A | 0.948 | HIS | 1.038 | GIIα | 0.590 | UBQ | 0.697 | HIS | 0.398 |
| 10 | GAPDH | 1.186 | TATA | 1.206 | HIS | 0.773 | elF-5A | 0.841 | GAPDH | 1.031 |
| 11 | GIIα | 1.770 | GAPDH | 1.472 | Cpn60β | 0.824 | Cpn60β | 0.897 | RA | 2.072 |
| 12 | UBQ | 3.104 | UBQ | 1.793 | RP | 0.970 | GAPDH | 0.943 | GIIα | 2.474 |
| 13 | RA | 3.335 | RA | 3.730 | AP-2 | 1.621 | RP | 0.967 | UBQ | 3.726 |
| 14 | TUB | 3.886 | TUB | 3.839 | ACT2 | 2.180 | AP-2 | 1.748 | TUB | 4.255 |
| Algorithms | Tissue | Leaf of MglFlora Treatment | Bud of MglFlora Treatment | ABA |
|---|---|---|---|---|
| Delta CT | Cpn60β + EF1α + elF-5A | TATA + elF-5A + RPL17 | TATA + ACT2 +EF1α | elF-5A + TATA + EF1α |
| BestKeeper | HIS + EF1α + TATA | HIS + ACT2 + EF1α | Cpn60β + HIS + EF1α | TATA + EF1α + Cpn60β |
| NormFinder | AP-2 + Cpn60β + elF-5A | TATA + elF-5A + EF1α | TATA + RPL17 + ACT2 | TATA + EF1α + elF-5A |
| geNorm | AP-2 + Cpn60β + elF-5A | RPL17 + elF-5 + GAPDH | EF1α + HIS + ACT2 | EF1α + TATA + elF-5A |
| RankAggreg | AP-2 + Cpn60β + elF-5A | TATA + elF-5A + RPL17 | TATA + HIS + ACT2 | EF1α + elF-5A + TATA |
| GrayNorm | ACT2 + HIS + TATA | ACT2 + EFα + HIS | Cpn60β + GAPDH + HIS | EF1α + HIS + TATA |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, J.-J.; Han, S.; Yin, W.; Xia, X.; Liu, C. Comparison of Reliable Reference Genes Following Different Hormone Treatments by Various Algorithms for qRT-PCR Analysis of Metasequoia. Int. J. Mol. Sci. 2019, 20, 34. https://doi.org/10.3390/ijms20010034
Wang J-J, Han S, Yin W, Xia X, Liu C. Comparison of Reliable Reference Genes Following Different Hormone Treatments by Various Algorithms for qRT-PCR Analysis of Metasequoia. International Journal of Molecular Sciences. 2019; 20(1):34. https://doi.org/10.3390/ijms20010034
Chicago/Turabian StyleWang, Jing-Jing, Shuo Han, Weilun Yin, Xinli Xia, and Chao Liu. 2019. "Comparison of Reliable Reference Genes Following Different Hormone Treatments by Various Algorithms for qRT-PCR Analysis of Metasequoia" International Journal of Molecular Sciences 20, no. 1: 34. https://doi.org/10.3390/ijms20010034
APA StyleWang, J.-J., Han, S., Yin, W., Xia, X., & Liu, C. (2019). Comparison of Reliable Reference Genes Following Different Hormone Treatments by Various Algorithms for qRT-PCR Analysis of Metasequoia. International Journal of Molecular Sciences, 20(1), 34. https://doi.org/10.3390/ijms20010034




