Derived Polymorphic Amplified Cleaved Sequence (dPACS): A Novel PCR-RFLP Procedure for Detecting Known Single Nucleotide and Deletion–Insertion Polymorphisms
Abstract
:1. Introduction
2. Results
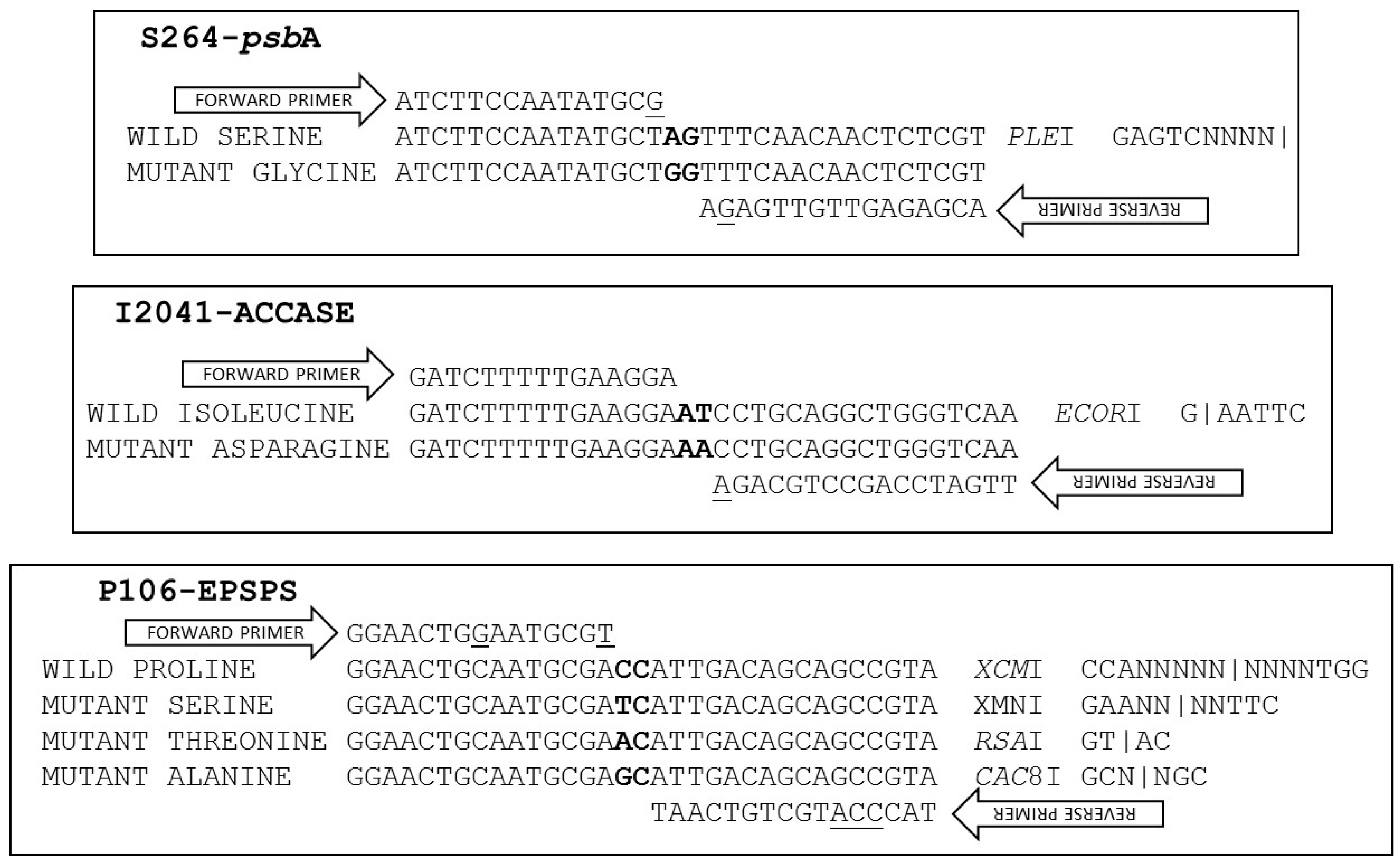
2.1. S264-psbA dPACS Assay
2.2. I2041-ACCase dPACS Assay
2.3. P106-EPSPS dPACS Assay
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Establishment of Derived Polymorphic Amplified Cleaved Sequence (dPACS) Assays
4.2.1. DNA Extraction
4.2.2. Nucleotide Sequences Around the SNPs of Interest
4.2.3. Development of the dPACS 1.0 Software and Selection of Distinguishing Primer/Enzyme Combinations
4.2.4. Three-Step PCR, Restriction Digestion and Gel Electrophoresis
4.3. Transferability of the dPACS Assays to a Wide Range of Weed Species
4.4. Genotyping Weed Populations Resistant to PSII, ACCase and EPSPS-Inhibiting Herbicides
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| SNP | single nucleotide polymorphism |
| DIP | deletion–insertion polymorphism |
| INDEL | insertion–deletion |
| BP | base pair |
References
- Brookes, A.J. The essence of snps. Gene 1999, 234, 177–186. [Google Scholar] [CrossRef]
- Venter, J.C.; Adams, M.D.; Myers, E.W.; Li, P.W.; Mural, R.J.; Sutton, G.G.; Smith, H.O.; Yandell, M.; Evans, C.A.; Holt, R.A.; et al. The sequence of the human genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Gurguis, C.I.; Zhou, J.J.; Zhu, Y.; Ko, E.-A.; Ko, J.-H.; Wang, T.; Zhou, T. Functional and structural consequence of rare exonic single nucleotide polymorphisms: One story, two tales. Genome Biol. Evol. 2015, 7, 2929–2940. [Google Scholar] [CrossRef]
- Marotta, C.; Wilson, J.; Forget, B.; Weissman, S. Human beta-globin messenger rna. Iii. Nucleotide sequences derived from complementary DNA. J. Biol. Chem. 1977, 252, 5040–5053. [Google Scholar] [PubMed]
- Saiki, R.K.; Chang, C.-A.; Levenson, C.H.; Warren, T.C.; Boehm, C.D.; Kazazian, H.H.J.; Erlich, H.A. Diagnosis of sickle cell anemia and β-thalassemia with enzymatically amplified DNA and nonradioactive allele-specific oligonucleotide probes. N. Engl. J. Med. 1988, 319, 537–541. [Google Scholar] [CrossRef] [PubMed]
- Tsui, L.-C. The spectrum of cystic fibrosis mutations. Trends Genet. 1992, 8, 392–398. [Google Scholar] [CrossRef]
- Zhou, Y.-A.; Ma, Y.-X.; Zhang, Q.-B.; Gao, W.-H.; Liu, J.-P.; Yang, J.-P.; Zhang, G.-X.; Zhang, X.-G.; Yu, L. Mutations of the phenylalanine hydroxylase gene in patients with phenylketonuria in Shanxi, China. Genet. Mol. Biol. 2012, 35, 709–713. [Google Scholar] [CrossRef] [Green Version]
- Khurana, E.; Fu, Y.; Chakravarty, D.; Demichelis, F.; Rubin, M.A.; Gerstein, M. Role of non-coding sequence variants in cancer. Nat. Rev. Genet. 2016, 17, 93–108. [Google Scholar] [CrossRef]
- Roy, B.; Haupt, L.M.; Griffiths, L.R. Review: Alternative splicing (as) of genes as an approach for generating protein complexity. Curr. Genom. 2013, 14, 182–194. [Google Scholar] [CrossRef]
- Wang, X.; Zhong, J.; Gao, Y.; Ju, Z.; Huang, J. A SNP in intron 8 of cd46 causes a novel transcript associated with mastitis in holsteins. BMC Genomics 2014, 15, 630. [Google Scholar] [CrossRef]
- Danjou, F.; Zoledziewska, M.; Sidore, C.; Steri, M.; Busonero, F.; Maschio, A.; Mulas, A.; Perseu, L.; Barella, S.; Porcu, E.; et al. Genome-wide association analyses based on whole-genome sequencing in sardinia provide insights into regulation of hemoglobin levels. Nat. Genet. 2015, 47, 1264–1271. [Google Scholar] [CrossRef] [PubMed]
- Zhi, S.; Li, Q.; Yasui, Y.; Banting, G.; Edge, T.A.; Topp, E.; McAllister, T.A.; Neumann, N.F. An evaluation of logic regression-based biomarker discovery across multiple intergenic regions for predicting host specificity in Escherichia coli. Mol. Phylogenet. Evol. 2016, 103, 133–142. [Google Scholar] [CrossRef] [PubMed]
- Chávez-Galarza, J.; Garnery, L.; Henriques, D.; Neves, C.J.; Loucif-Ayad, W.; Jonhston, J.S.; Pinto, M.A. Mitochondrial DNA variation of Apis mellifera iberiensis: Further insights from a large-scale study using sequence data of the tRNAleu-cox2 intergenic region. Apidologie 2017, 48, 533–544. [Google Scholar] [CrossRef]
- Montgomery, S.; Goode, D.; Kvikstad, E.; Albers, C.; Zhang, Z.; Mu, X.; Ananda, G.; Howie, B.; Karczewski, K.; Smith, K.; et al. The origin, evolution, and functional impact of short insertion-deletion variants identified in 179 human genomes. Genome Res. 2013, 23, 749–761. [Google Scholar] [CrossRef] [PubMed]
- Wright, A.V.; Nuñez, J.K.; Doudna, J.A. Biology and applications of crispr systems: Harnessing nature’s toolbox for genome engineering. Cell 2016, 164, 29–44. [Google Scholar] [CrossRef]
- Amin, S.; Forrester, N.; Norman, A.; Lux, A.; Vijayakumar, K. Novel 9 amino acid in-frame deletion in the ntrk1 tyrosine kinase domain in a patient with congenital insensitivity to pain with anhydrosis. Clin. Genet. 2017, 92, 559–560. [Google Scholar] [CrossRef] [PubMed]
- Pausch, H.; Venhoranta, H.; Wurmser, C.; Hakala, K.; Iso-Touru, T.; Sironen, A.; Vingborg, R.K.; Lohi, H.; Söderquist, L.; Fries, R.; et al. A frameshift mutation in armc3 is associated with a tail stump sperm defect in swedish red (Bos taurus) cattle. BMC Genet. 2016, 17, 49. [Google Scholar] [CrossRef]
- Chen, X.; Sullivan, P. Single nucleotide polymorphism genotyping: Biochemistry, protocol, cost and throughput. Pharmacogenomics J. 2003, 3, 77–96. [Google Scholar] [CrossRef]
- Mamotte, C.D.S. Genotyping of single nucleotide substitutions. Clin. Biochem. Rev. 2006, 27, 63–75. [Google Scholar]
- Kwok, P.-Y.; Chen, X. Detection of single nucleotide polymorphisms. Curr. Issues Mol. Biol. 2003, 5, 43–60. [Google Scholar]
- Ding, C.; Jin, S. High-throughput methods for SNP genotyping. In Single Nucleotide Polymorphisms: Methods and Protocols; Komar, A.A., Ed.; Humana Press: Totowa, NJ, USA, 2009; pp. 245–254. [Google Scholar]
- Malkki, M.; Petersdorf, E.W. Genotyping of single nucleotide polymorphisms by 5′ nuclease allelic discrimination. Methods in Molecular Biology 2012, 882, 173–182. [Google Scholar] [PubMed]
- Holland, P.M.; Abramson, R.D.; Watson, R.; Gelfand, D.H. Detection of specific polymerase chain reaction product by utilizing the 5’-3’ exonuclease activity of Thermus aquaticus DNA polymerase. Proc. Natl. Acad. Sci. USA 1991, 88, 7276–7280. [Google Scholar] [CrossRef] [PubMed]
- Voelkerding, K.V.; Dames, S.A.; Durtschi, J.D. Next-generation sequencing: From basic research to diagnostics. Clin. Chem. 2009, 55, 641–658. [Google Scholar] [CrossRef] [PubMed]
- Nyren, P.; Pettersson, B.; Uhlen, M. Solid phase DNA minisequencing by an enzymatic luminometric inorganic pyrophosphate detection assay. Anal. Biochem. 1993, 208, 171–175. [Google Scholar] [CrossRef] [PubMed]
- Makridakis, N.; Reichardt, J. Multiplex automated primer extension analysis: Simultaneous genotyping of several polymorphisms. Biotechniques 2001, 31, 1374–1380. [Google Scholar] [CrossRef] [PubMed]
- Bottema, C.; Sommer, S. PCR amplification of specific alleles: Rapid detection of known mutations and polymorphisms. Mutat. Res. 1993, 288, 93–102. [Google Scholar] [CrossRef]
- Ota, M.; Asamura, H.; Oki, T.; Sada, M. Restriction enzyme analysis of PCR products. In Single Nucleotide Polymorphisms: Methods and Protocols; Komar, A.A., Ed.; Humana Press: Totowa, NJ, USA, 2009; pp. 405–414. [Google Scholar]
- Kogan, S.C.; Doherty, M.; Gitschier, J. An improved method for prenatal diagnosis of genetic diseases by analysis of amplified DNA sequences. N. Engl. J. Med. 1987, 317, 985–990. [Google Scholar] [CrossRef]
- Neff, M.M.; Neff, J.D.; Chory, J.; Pepper, A.E. Dcaps, a simple technique for the genetic analysis of single nucleotide polymorphisms: Experimental applications in Arabidposis thaliana genetics. Plant. J. 1998, 14, 387–392. [Google Scholar] [CrossRef]
- Konieczny, A.; Ausubel, F.M. A procedure for mapping arabidopsis mutations using co-dominant ecotype-specific PCR-based markers. Plant. J. 1993, 4, 403–410. [Google Scholar] [CrossRef]
- Neff, M.M.; Turk, E.; Kalishman, M. Web-based primer design for single nucleotide polymorphism analysis. Trends Genet. 2002, 18, 613–615. [Google Scholar] [CrossRef]
- Powles, S.; Yu, Q. Evolution in action: Plants reistant to herbicides. Annu Rev. Plant. Biol 2010, 61, 317–347. [Google Scholar] [CrossRef] [PubMed]
- Corriveau, J.; Coleman, A. Rapid screening method to detect potential biparental inheritance of plastid DNA and results for over 200 angiosperm species. Am. J. Bot. 1988, 75, 1443–1458. [Google Scholar] [CrossRef]
- Kim, S.; Misra, A. SNP genotyping: Technologies and biomedical applications. Annu Rev. Biomed. Eng. 2007, 9, 289–320. [Google Scholar] [CrossRef] [PubMed]
- Mammadov, J.; Aggarwal, R.; Buyyarapu, R.; Kumpatla, S. SNP markers and their impact on plant breeding. Int J. Plant. Genomics 2012, 2012, 11. [Google Scholar] [CrossRef] [PubMed]
- Helyar, S.J.; Hemmer-Hansen, J.; Bekkevold, D.; Taylor, M.I.; Ogden, R.; Limborg, M.T.; Cariani, A.; Maes, G.E.; Diopere, E.; Carvalho, G.R.; et al. Application of SNPs for population genetics of nonmodel organisms: New opportunities and challenges. Mol. Ecol. Resour. 2011, 11, 123–136. [Google Scholar] [CrossRef] [PubMed]
- Southern, E.M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J. Mol. Biol. 1975, 98, 503–517. [Google Scholar] [CrossRef]
- Cottingham, K. Product review: Multiple choices for SNPs. Anal. Chem 2004, 76, 179 A–181 A. [Google Scholar] [CrossRef]
- Teumer, A.; Ernst, F.D.; Wiechert, A.; Uhr, K.; Nauck, M.; Petersmann, A.; Völzke, H.; Völker, U.; Homuth, G. Comparison of genotyping using pooled DNA samples (allelotyping) and individual genotyping using the affymetrix genome-wide human SNP array 6.0. BMC Genom. 2013, 14, 506. [Google Scholar] [CrossRef]
- Semagn, K.; Babu, R.; Hearne, S.; Olsen, M. Single nucleotide polymorphism genotyping using kompetitive allele specific PCR (kasp): Overview of the technology and its application in crop improvement. Mol. Breed. 2014, 33, 1–14. [Google Scholar] [CrossRef]
- Hidaka, A.; Sasazuki, S.; Matsuo, K.; Ito, H.; Charvat, H.; Sawada, N.; Shimazu, T.; Yamaji, T.; Iwasaki, M.; Inoue, M.; et al. Cyp1a1, gstm1 and gstt1 genetic polymorphisms and gastric cancer risk among japanese: A nested case–control study within a large-scale population-based prospective study. Int. J. Cancer 2016, 139, 759–768. [Google Scholar] [CrossRef]
- Arita, H.; Narita, Y.; Matsushita, Y.; Fukushima, S.; Yoshida, A.; Takami, H.; Miyakita, Y.; Ohno, M.; Shibui, S.; Ichimura, K. Development of a robust and sensitive pyrosequencing assay for the detection of idh1/2 mutations in gliomas. Brain Tumor Pathol. 2015, 32, 22–30. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Henderson, G.; Sun, X.; Cox, F.; Janssen, P.H.; Guan, L.L. Taxonomic assessment of rumen microbiota using total rna and targeted amplicon sequencing approaches. Front. Microbiol. 2016, 7, 987. [Google Scholar] [CrossRef] [PubMed]
- Griffin, T.J.; Smith, L.M. Single-nucleotide polymorphism analysis by maldi–tof mass spectrometry. Trends Biotechnol. 2000, 18, 77–84. [Google Scholar] [CrossRef]
- Muir, P.; Li, S.; Lou, S.; Wang, D.; Spakowicz, D.J.; Salichos, L.; Zhang, J.; Weinstock, G.M.; Isaacs, F.; Rozowsky, J.; et al. The real cost of sequencing: Scaling computation to keep pace with data generation. Genome Biol. 2016, 17, 53. [Google Scholar] [CrossRef] [PubMed]
- Tandon, D.; Dewangan, J.; Srivastava, S.; Garg, V.K.; Rath, S.K. Mirna genetic variants: As potential diagnostic biomarkers for oral cancer. Pathol Res. Pract. In press. [CrossRef] [PubMed]
- Gamble, T.; Castoe, T.A.; Nielsen, S.V.; Banks, J.L.; Card, D.C.; Schield, D.R.; Schuett, G.W.; Booth, W. The discovery of xy sex chromosomes in a boa and python. Curr. Biol. 2017, 27, 2148–2153. [Google Scholar] [CrossRef] [PubMed]
- Patiño, L.; Cruz, M.; Martínez, P.; Cedeño-Escobar, V. Using PCR-RFLP for sexing of the endangered galápagos petrel (pterodroma phaeopygia). Genet. Mol. Res. 2013, 12, 4760–4767. [Google Scholar] [CrossRef]
- Fontecha, G.; Garcia, K.; Rueda, M.; Sosa-Ochoa, W.; Sanchez, A.; Leiva, B. A PCR-RFLP method for the simultaneous differentiation of three entamoeba species. Exp. Parasitol. 2015, 151–152, 80–83. [Google Scholar] [CrossRef]
- Rohit, A.; Maiti, B.; Shenoy, S.; Karunasagar, I. Polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) for rapid diagnosis of neonatal sepsis. Indian J. Med. Res. 2016, 143, 72–78. [Google Scholar]
- Riaz, M.; Mahmood, Z.; Javed, M.; Javed, I.; Shahid, M.; Abbas, M.; Ehtisham-ul-Haque, S. Drug resistant strains of Mycobacterium tuberculosis identified through PCR-RFLP from patients of Central Punjab, Pakistan. Int. J. Immunopathol. Pharmacol. 2016, 29, 433–449. [Google Scholar] [CrossRef]
- Tanaka, K.; Stevens, C.J.; Iwasaki, S.; Akashi, Y.; Yamamoto, E.; Dung, T.P.; Nishida, H.; Fuller, D.Q.; Kato, K. Seed size and chloroplast DNA of modern and ancient seeds explain the establishment of japanese cultivated melon (Cucumis melo L.) by introduction and selection. Genet. Resour Crop. Evol. 2016, 63, 1237–1254. [Google Scholar] [CrossRef]
- Liu, H.; Jiao, J.; Liang, X.; Liu, J.; Meng, H.; Chen, S.; Li, Y.; Cheng, Z. Map-based cloning, identification and characterization of the w gene controlling white immature fruit color in cucumber (Cucumis sativus L.). Theor. Appl. Genet. 2016, 129, 1247–1256. [Google Scholar] [CrossRef] [PubMed]
- Busi, R.; Michel, S.; Powles, S.B.; Délye, C. Gene flow increases the initial frequency of herbicide resistance alleles in unselected Lolium rigidum populations. Agric. Ecosyst. Environ. 2011, 142, 403–409. [Google Scholar] [CrossRef]
- Shimono, Y.; Shimono, A.; Oguma, H.; Konuma, A.; Tominaga, T. Establishment of lolium species resistant to acetolactate synthase-inhibiting herbicide in and around grain-importation ports in japan. Weed Res. 2015, 55, 101–111. [Google Scholar] [CrossRef]
- Délye, C.; Causse, R.; Michel, S. Genetic basis, evolutionary origin and spread of resistance to herbicides inhibiting acetolactate synthase in common groundsel (Senecio vulgaris). Pest Manag. Sci. 2016, 72, 89–102. [Google Scholar] [CrossRef] [PubMed]
- Kapusi, E.; Corcuera-Gómez, M.; Melnik, S.; Stoger, E. Heritable genomic fragment deletions and small indels in the putative engase gene induced by crispr/cas9 in barley. Front. Plant. Sci. 2017, 8, 540. [Google Scholar] [CrossRef]
- Shin, H.Y.; Wang, C.; Lee, H.K.; Yoo, K.H.; Zeng, X.; Kuhns, T.; Yang, C.M.; Mohr, T.; Liu, C.; Hennighausen, L. Crispr/cas9 targeting events cause complex deletions and insertions at 17 sites in the mouse genome. Nat. Commun. 2017, 8, 15464. [Google Scholar] [CrossRef]
- Delye, C.; Pernin, F.; Michel, S. ‘Universal’ PCR assays detecting mutations in acetyl-coenzyme a carboxylase or acetolactate synthase that endow herbicide resistance in grass weeds. Weed Res. 2011, 51, 353–362. [Google Scholar] [CrossRef]
- Kaundun, S.S.; Dale, R.P.; Bailly, G.C. Molecular basis of resistance to herbicides inhibiting acetolactate synthase in two rigid ryegrass (Lolium rigidum) populations from australia. Weed Sci 2012, 60, 172–178. [Google Scholar] [CrossRef]
- Di, H.; Liu, X.; Wang, Q.; Weng, J.; Zhang, L.; Li, X.; Wang, Z. Development of SNP-based dcaps markers linked to major head smut resistance quantitative trait locus qhs2.09 in maize. Euphytica 2015, 202, 69–79. [Google Scholar] [CrossRef]
- Kirii, E.; Goto, T.; Yoshida, Y.; Yasuba, K.; Tanaka, Y. Non-pungency in a japanese chili pepper landrace (capsicum annuum) is caused by a novel loss-of-function pun1 allele. Hort J. 2017, 86, 61–69. [Google Scholar] [CrossRef]
- Klesiewicz, K.; Nowak, P.; Karczewska, E.; Skiba, I.; Wojtas-Bonior, I.; Sito, E.; Budak, A. PCR-RFLP detection of point mutations a2143g and a2142g in 23s rrna gene conferring resistance to clarithromycin in Helicobacter pylori strains. Acta Biochim Pol. 2014, 61, 311–315. [Google Scholar] [CrossRef] [PubMed]
- Massa, D.; Krenz, B.; Gerhards, R. Target-site resistance to als-inhibiting herbicides in Apera spica-venti populations is conferred by documented and previously unknown mutations. Weed Res. 2011, 51, 294–303. [Google Scholar] [CrossRef]
- Whaley, C.M.; Wilson, H.P.; Westwood, J.H. A new mutation in plant als confers resistance to five classes of als-inhibiting herbicides. Weed Sci. 2007, 55, 83–90. [Google Scholar] [CrossRef]
- Yuan, G.; Liu, W.; Bi, Y.; Du, L.; Guo, W.; Wang, J. Molecular basis for resistance to accase-inhibiting herbicides in pseudosclerochloa kengiana populations. Pest Bichem. Physiol. 2015, 119, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Kaundun, S.S.; Bailly, G.; Dale, R.; Hutchings, S.-J.; Mcindoe, E. A novel w1999s mutation and non-target site resistance impact on acetyl-coa carboxylase inhibiting herbicides to varying degrees in a UK Lolium multiflorum population. PLoS ONE 2013, 8, e58012. [Google Scholar] [CrossRef] [PubMed]
- Wuerffel, R.; Young, J.; Matthews, J.; Young, B. Characterization of ppo-inhibitor–resistant waterhemp (Amaranthus tuberculatus) response to soil-applied ppo-inhibiting herbicides. Weed Sci 2015, 63, 511–521. [Google Scholar] [CrossRef]
- Giacomini, D.A.; Umphres, A.M.; Nie, H.; Mueller, T.C.; Steckel, L.E.; Young, B.G.; Scott, R.C.; Tranel, P.J. Two new ppx2 mutations associated with resistance to ppo-inhibiting herbicides in Amaranthus palmeri. Pest Manag. Sci. 2017, 73, 1559–1563. [Google Scholar] [CrossRef]
- Gryson, N. Effect of food processing on plant DNA degradation and PCR-based gmo analysis: A review. Anal. Bioanal. Chem. 2010, 396, 2003–2022. [Google Scholar] [CrossRef]
- Fulton, T.L.; Stiller, M. PCR amplification, cloning, and sequencing of ancient DNA. In Ancient DNA: Methods and Protocols; Shapiro, B., Hofreiter, M., Eds.; Humana Press: Totowa, NJ, USA, 2012; pp. 111–119. [Google Scholar]
- Schrader, C.; Schielke, A.; Ellerbroek, L.; Johne, R. PCR inhibitors – occurrence, properties and removal. J. Appl. Microbiol. 2012, 113, 1014–1026. [Google Scholar] [CrossRef]
- Lv, P.; Zhou, X.; You, J.; Ye, B.; Zhang, Y. Extraction of trace amount of severely degraded DNA. Z Naturforsch C. 2009, 64, 581–589. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Zhao, M.; Yao, H.; Yang, P.; Xin, T.; Li, B.; Sun, W.; Chen, S. Rapidly discriminate commercial medicinal pulsatilla chinensis (bge.) regel from its adulterants using its2 barcoding and specific PCR-RFLP assay. Sci. Rep. 2017, 7, 40000. [Google Scholar] [CrossRef] [PubMed]
- Razzak, M.A.; Hamid, S.B.A.; Ali, M.E. A lab-on-a-chip-based multiplex platform to detect potential fraud of introducing pig, dog, cat, rat and monkey meat into the food chain. Food Addit. Contam. Part. A 2015, 32, 1902–1913. [Google Scholar] [CrossRef] [PubMed]
- Doosti, A.; Ghasemi Dehkordi, P.; Rahimi, E. Molecular assay to fraud identification of meat products. J. Food Sci. Technol. 2014, 51, 148–152. [Google Scholar] [CrossRef] [PubMed]
- Shukla, A.; Devine, M.D. Chapter 9 - basis of crop selectivity and weed resistance to triazine herbicides. In The Triazine Herbicides; Elsevier: San Diego, CA, USA, 2008; pp. 111–118. [Google Scholar]
- Kaundun, S.S. Resistance to acetyl-coa carboxylase-inhibiting herbicides. Pest Manag. Sci. 2014, 70, 1405–1417. [Google Scholar] [CrossRef] [PubMed]
- Kaundun, S.S.; Zelaya, I.; Dale, R.; Lycett, A.; Carter, P.; Sharples, P.; McIndoe, E. Importance of the p106s target site mutation in conferring resistance to glyphosate in an Eleusine indica biotype from the philippines. Weed Sci. 2008, 56, 637–646. [Google Scholar] [CrossRef]
- Han, H.; Yu, Q.; Widderick, M.J.; Powles, S.B. Target-site epsps pro-106 mutations: Sufficient to endow glyphosate resistance in polyploid Echinochloa colona? Pest Manag. Sci. 2016, 72, 264–271. [Google Scholar] [CrossRef] [PubMed]
- Perez-Jones, A.; Mallory-Smith, C. Biochemical mechanisms and molecular basis of evolved glyphosate resistance in weed species. In Glyphosate Resistance in Crops and Weeds; John Wiley & Sons: Hoboken, NJ, USA, 2010; pp. 119–140. [Google Scholar]
- Sammons, R.; Gaines, T. Glyphosate resistance: State of knowledge. Pest Manag. Sci. 2014, 70, 1367–1377. [Google Scholar] [CrossRef] [PubMed]
- Bell, M.S.; Hager, A.G.; Tranel, P.J. Multiple resistance to herbicides from four site-of-action groups in waterhemp (Amaranthus tuberculatus). Weed Sci. 2013, 61, 460–468. [Google Scholar] [CrossRef]
- Baucom, R. The remarkable repeated evolution of herbicide resistance. Am. J. Bot 2016, 103, 181–183. [Google Scholar] [CrossRef] [Green Version]
- Preston, C. Herbicide resistance: Target site mutations. In Weedy and Invasive Plant Genomics; Wiley-Blackwell: Ames, IA, USA, 2009; pp. 127–148. [Google Scholar]
- Bettini, P.; McNally, S.; Sevignac, M.; Darmency, H.; Gasquez, J.; Dron, M. Atrazine resistance in Chenopodium album: Low and high levels of resistance to the herbicide are related to the same chloroplast psba gene mutation. Plant. Physiol. 1987, 84, 1442–1446. [Google Scholar] [CrossRef] [PubMed]
- Thiel, H.; Kluth, C.; Varrelmann, M. A new molecular method for the rapid detection of a metamitron-resistant target site in Chenopodium album. Pest Manag. Sci. 2010, 66, 1011–1017. [Google Scholar] [CrossRef] [PubMed]
- Tian, X.; Darmency, H. Rapid bidirectional allele-specific PCR identification for triazine resistance in higher plants. Pest Manag. Sci. 2006, 62, 531–536. [Google Scholar] [CrossRef] [PubMed]
- Ayyadevara, S.; Thaden, J.J.; Shmookler Reis, R.J. Discrimination of primer 3′-nucleotide mismatch by taq DNA polymerase during polymerase chain reaction. Anal. Biochem. 2000, 284, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Goloubinoff, P.; Edelman, M.; Hallick, R.B. Chloroplast-coded atrazine resistance in Solanum nigrum: Psba loci from susceptible and resistant biotypes are isogenic except for a single codon change. Nucleic Acids Res. 1984, 12, 9489–9496. [Google Scholar] [CrossRef]
- Mechant, E.; De Marez, T.; Hermann, O.; Bulcke, R. Resistance of Chenopodium album to photosystem ii-inhibitors. Commun. Agric. Appl. Biol. Sci. 2008, 73, 913–917. [Google Scholar]
- Robertson, M.J.; Holland, J.F.; Cawley, S.; Potter, T.D.; Burton, W.; Walton, G.H.; Thomas, G. Growth and yield differences between triazine-tolerant and non-triazine-tolerant cultivars of canola. Aust J. Agric. Res. 2002, 53, 643–651. [Google Scholar] [CrossRef]
- Yu, Q.; Collavo, A.; Zheng, M.-Q.; Owen, M.; Sattin, M.; Powles, S.B. Diversity of acetyl-coenzyme a carboxylase mutations in resistant lolium populations: Evaluation using clethodim. Plant. Physiol. 2007, 145, 547–558. [Google Scholar] [CrossRef]
- Délye, C.; Zhang, X.Q.; Chalopin, C.; Michel, S.; Powles, S.B. An isoleucine residue within the carboxyl-transferase domain of multidomain acetyl-coenzyme a carboxylase is a major determinant of sensitivity to aryloxyphenoxypropionate but not to cyclohexanedione inhibitors. Plant. Physiol. 2003, 132, 1716–1723. [Google Scholar] [CrossRef]
- Kwok, S. Procedures to minimize PCR-product carry-over. In PCR Protocols: A Guide to Methods and Applications, Innis, M.; Gelfand, D.H., Sninsky, J., White, T., Eds.; Academic Press: Cambridge, MA, USA, 1990; pp. 142–145. [Google Scholar]
- Yu, Q.; Jalaludin, A.; Han, H.; Chen, M.; Sammons, R.D.; Powles, S.B. Evolution of a double amino acid substitution in the 5-enolpyruvylshikimate-3-phosphate synthase in Eleusine indica conferring high-level glyphosate resistance. Plant. Physiol. 2015, 167, 1440–1447. [Google Scholar] [CrossRef]
- Molin, W.; Wright, A.; Nandula, V. Glyphosate-resistant goosegrass from mississippi. Agronomy 2013, 3, 474–487. [Google Scholar] [CrossRef]
- Kaundun, S.S.; Dale, R.; Zelaya, I.; Dinelli, G.; Maroti, I.; Mcindoe, E.; Cairns, A. A novel p106l mutation in the epsps and an unknown mechanism(s) act additively to confer resistance to glyphosate in a south african Lolium rigidum population. J. Agric. Food Chem. 2011, 59, 3227–3233. [Google Scholar] [CrossRef] [PubMed]
- Chatham, L.A.; Wu, C.; Riggins, C.W.; Hager, A.G.; Young, B.G.; Roskamp, G.K.; Tranel, P.J. Epsps gene amplification is present in the majority of glyphosate-resistant illinois waterhemp (Amaranthus tuberculatus) populations. Weed Technol. 2015, 29, 48–55. [Google Scholar] [CrossRef]
- Heap, I.M. International Survey of Herbicide-resistant Weeds. 2018. Available online: Http://www.Weedscience.Com (accessed on 22 May 2019).
- Chen, J.; Huang, H.; Zhang, C.; Wei, S.; Huang, Z.; Chen, J.; Wang, X. Mutations and amplification of epsps gene confer resistance to glyphosate in goosegrass (Eleusine indica). Planta 2015, 242, 859–868. [Google Scholar] [CrossRef] [PubMed]
- Gaines, T.; Zhang, W.; Wang, D.; Bukuna, B.; Chisholm, S.; Shaner, D.; Nissen, S.; Patzoldt, W.; Tranel, P.; Culpepper, A.; et al. Gene amplification confers glyphosate resistance in Amaranthus palmeri. Proc. Natl. Acad. Sci. USA 2010, 107, 1029–1034. [Google Scholar] [CrossRef] [PubMed]
- Molin, W.T.; Wright, A.A.; VanGessel, M.J.; McCloskey, W.B.; Jugulam, M.; Hoagland, R.E. Survey of the genomic landscape surrounding the 5-enolpyruvylshikimate-3-phosphate synthase (epsps) gene in glyphosate-resistant Amaranthus palmeri from geographically distant populations in the USA. Pest Manag. Sci. 2018, 74, 1109–1117. [Google Scholar] [CrossRef]
- Küpper, A.; Borgato, E.A.; Patterson, E.L.; Gonçalves Netto, A.; Nicolai, M.; de Carvalho, S.J.P.; Nissen, S.J.; Gaines, T.A.; Christoffoleti, P.J. Multiple resistance to glyphosate and acetolactate synthase inhibitors in palmer amaranth (Amaranthus palmeri) identified in brazil. Weed Sci. 2017, 65, 317–326. [Google Scholar] [CrossRef]
- Chahal, P.S.; Varanasi, V.K.; Jugulam, M.; Jhala, A.J. Glyphosate-resistant palmer amaranth (Amaranthus palmeri) in nebraska: Confirmation, epsps gene amplification, and response to post corn and soybean herbicides. Weed Technol. 2017, 31, 80–93. [Google Scholar] [CrossRef]
- De Carvalho, L.B.; Alves, P.L.d.C.A.; González-Torralva, F.; Cruz-Hipolito, H.E.; Rojano-Delgado, A.M.; De Prado, R.; Gil-Humanes, J.; Barro, F.; Luque de Castro, M.D. Pool of resistance mechanisms to glyphosate in Digitaria insularis. J. Agric. Food Chem. 2012, 60, 615–622. [Google Scholar] [CrossRef]
- Konishi, T.; Shinohara, K.; Yamada, K.; Sasaki, Y. Acetyl-coa carboxylase in higher plants: Most plants other than gramineae have both the prokaryotic and the eukaryotic forms of this enzyme. Plant. Cell Physiol. 1996, 37, 2. [Google Scholar] [CrossRef]
- Délye, C.; Duhoux, A.; Pernin, F.; Riggins, C.W.; Tranel, P.J. Molecular mechanisms of herbicide resistance. Weed Sci. 2015, 63, 91–115. [Google Scholar] [CrossRef]
- Alarcón-Reverte, R.; García, A.; Watson, S.B.; Abdallah, I.; Sabaté, S.; Hernández, M.J.; Dayan, F.E.; Fischer, A.J. Concerted action of target-site mutations and high epsps activity in glyphosate-resistant junglerice (Echinochloa colona) from california. Pest Manag. Sci. 2015, 71, 996–1007. [Google Scholar] [CrossRef] [PubMed]




| Species | Population | Origin | dPACS and Sanger Sequencing Assays | Genotypic Frequencies as Revealed by Sequencing and dPACS Assays |
|---|---|---|---|---|
| Amaranthus retroflexus | ArD1 | USA | S264G psbA | 100% mutant G264 |
| ArD2 | Switzerland | S264G psbA | 100% wild S264 | |
| ArD3 | USA | S264G psbA | 100% wild S264 | |
| ArD4 | Switzerland | S264G psbA | 100% mutant G264 | |
| ArD5 | Switzerland | S264G psbA | 100% wild S264 | |
| ArD6 | Switzerland | S264G psbA | 100% mutant G264 | |
| Lolium multiflorum | LmD1 | UK | I2041N ACCase | 25%II2041, 50%IN2041, 25%NN2041 |
| LmD2 | UK | I2041N ACCase | 100% II2041 | |
| LmD3 | UK | I2041N ACCase | 100% II2041 | |
| LmD4 | UK | I2041N ACCase | 100% II2041 | |
| LmD5 | UK | I2041N ACCase | 100% NN2041 | |
| LmD6 | UK | I2041N ACCase | 12.5% II2041, 87.5% NN2041 | |
| Eleusine indica | EiD1 | Philippines | P106S/T/A EPSPS | 100% SS106 |
| EiD2 | Malaysia | P106S/T/A EPSPS | 100% SS106 | |
| EiD3 | Ecuador | P106S/T/A EPSPS | 100 % AA106 | |
| EiD4 | Malaysia | P106S/T/A EPSPS | 100 % PP106 | |
| EiD5 | Malaysia | P106S/T/A EPSPS | 50% PP106, 25% PS106, 25% SS106 | |
| EiD6 | Malaysia | P106S/T/A EPSPS | 25% PP106, 75% SS106 |
| Species | Population | Origin | dPACS Assay | Genotypic Frequencies |
|---|---|---|---|---|
| Chenopodium album | CaR1 | Poland | S264G psbA | 100% G264 |
| Solanum nigrum | SnR1 | Germany | S264G psbA | 100% G264 |
| SnR2 | Unknown | S264G psbA | 100% G264 | |
| Setaria viridis | SvR1 | USA | I2041N ACCase | 100% II2041 |
| SvR2 | USA | I2041N ACCase | 87.5% II2041, 12.5% IN2041 | |
| SvR3 | Canada | I2041N ACCase | 100% II2041 | |
| SvR4 | USA | I2041N ACCase | 100% II2041 | |
| SvR5 | Canada | I2041N ACCase | 100% II2041 | |
| SvR6 | USA | I2041N ACCase | 100% II2041 | |
| Eleusine indica | EiR1 | Malaysia | I2041N ACCase | 100% II2041 |
| EiR2 | Philippines | I2041N ACCase | 87.5% II2041, 12.5 % NN2041 | |
| E2R3 | Malaysia | I2041N ACCase | 87.5% II2041, 12.5 % NN2041 | |
| EiR4 | Ecuador | I2041N ACCase | 100% II2041 | |
| EiR5 | Malaysia | I2041N ACCase | 100% II2041 | |
| EiR6 | Malaysia | I2041N ACCase | 75% II2041, 12.5% IN2041, 12.5% NN2041 | |
| Amaranthus palmeri | ApR1 | USA | P106S/T/A EPSPS | 100% PP106 |
| ApR2 | USA | P106S/T/A EPSPS | 100% PP106 | |
| ApR3 | USA | P106S/T/A EPSPS | 100% PP106 | |
| ApR4 | USA | P106S/T/A EPSPS | 100% PP106 | |
| ApR5 | USA | P106S/T/A EPSPS | 100% PP106 | |
| ApR6 | USA | P106S/T/A EPSPS | 100% PP106 | |
| Digitaria insularis | DiR1 | Brazil | P106S/T/A EPSPS | 100% PS106 |
| DiR2 | Brazil | P106S/T/A EPSPS | 100% PS106 | |
| DiR3 | Brazil | P106S/T/A EPSPS | 100% PS106 | |
| DiR4 | Brazil | P106S/T/A EPSPS | 100% PT106 | |
| DiR5 | Brazil | P106S/T/A EPSPS | 100% PS106 | |
| DiR6 | Brazil | P106S/T/A EPSPS | 100% PS106 |
| Primer ID | Primer Length (bp) | Primer Sequence 5’–3’ | Target Codon and Amino | PCR Product, Restriction Enzyme and Digest | |
|---|---|---|---|---|---|
| Sequencing primers | SeqFw-psbA-A.retroflexus | 25 | AAGGAAGCTTTTCTGATGGTATGCC | 264 psbA | 426 bp; codons 165–307 |
| SeqRv-psbA-A.retroflexus | 25 | CTACAGATTGGTTGAAGTTGAAACC | |||
| SeqFw-ACCase-L. multiflorum | 22 | TGGCAGAGCAAAACTTGGAGGG | 2041 ACCase | 468 bp; codons 1954–2110 | |
| SeqRv-ACCase-L. multiflorum | 23 | CTGAACTTGATCTCAATCAACCC | |||
| SeqFw-EPSPS-E. indica | 22 | GGAACAACTGTGGTGGATAACC | 106 EPSPS | 492 or 498 bp; codons 39–171 | |
| SeqRv-EPSPS-E. indica | 20 | CTTGCCACCAGGTAGCCCTC | |||
| dPACS primers | Fw-PsbA-A.retroflexus | 40 | TGCTTCATGGTTACTTTGGTCGATTGATCTTCCAATATGCG | S264 psbA | PleI (GAGTCN4|) |
| Rv-PsbA-A.retroflexus | 41 | AAGCAGCTAAGAAAAAGTGTAAAGAACGAGAGTTGTTGAGA | 83 bp undigested; 48/35 bp digested | ||
| Fw-ACCase-L. multiflorum | 35 | GCTTCTCTGGTGGGCAAAGAGACCTTTTTGAAGGA | I2041 ACCase | EcorI (G|AATTC) | |
| Rv-ACCase-L. multiflorum | 55 | GGCAGGCAGATTATATGTCCTAAGGTTCTCAACAATTGTTGATCCAGCCTGCAGA | 92 bp undigested; 58/34 bp digested | ||
| Fw-EPSPS-E. indica | 40 | GGTGCAGCTCTTCTTGGGGAATGCTGGAACTGGAATGCGT | P106 EPSPS | XcmI (CCA(NNNNN|NNNN)TGG | |
| Fw-EPSPS-E. indica | 40 | AGTTGCATTTCCTCCAGCAGCAGTTACCCATGCTGTCAAT | 82 bp undigested; 47/35 bp digested | ||
| Rv-EPSPS-E. indica | 40 | GGTGCAGCTCTTCTTGGGGAATGCTGGAACTGGAATGCGT | S106 EPSPS | XmnI (GAANN|NNTTC) | |
| Rv-EPSPS-E. indica | 40 | AGTTGCATTTCCTCCAGCAGCAGTTACCCATGCTGTCAAT | 82 bp undigested; 45/37 bp digested | ||
| Fw-EPSPS-E. indica | 40 | GGTGCAGCTCTTCTTGGGGAATGCTGGAACTGGAATGCGT | T106 EPSPS | RsaI (GT|AC) | |
| Rv-EPSPS-E. indica | 40 | AGTTGCATTTCCTCCAGCAGCAGTTACCCATGCTGTCAAT | 82 bp undigested; 42/40 bp digested | ||
| Fw-EPSPS-E. indica | 40 | GGTGCAGCTCTTCTTGGGGAATGCTGGAACTGGAATGCGT | A106 EPSPS | Cac8I (GCN|NGC) | |
| Rv-EPSPS-E. indica | 40 | AGTTGCATTTCCTCCAGCAGCAGTTACCCATGCTGTCAAT | 82 bp undigested; 43/39 bp digested | ||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaundun, S.S.; Marchegiani, E.; Hutchings, S.-J.; Baker, K. Derived Polymorphic Amplified Cleaved Sequence (dPACS): A Novel PCR-RFLP Procedure for Detecting Known Single Nucleotide and Deletion–Insertion Polymorphisms. Int. J. Mol. Sci. 2019, 20, 3193. https://doi.org/10.3390/ijms20133193
Kaundun SS, Marchegiani E, Hutchings S-J, Baker K. Derived Polymorphic Amplified Cleaved Sequence (dPACS): A Novel PCR-RFLP Procedure for Detecting Known Single Nucleotide and Deletion–Insertion Polymorphisms. International Journal of Molecular Sciences. 2019; 20(13):3193. https://doi.org/10.3390/ijms20133193
Chicago/Turabian StyleKaundun, Shiv Shankhar, Elisabetta Marchegiani, Sarah-Jane Hutchings, and Ken Baker. 2019. "Derived Polymorphic Amplified Cleaved Sequence (dPACS): A Novel PCR-RFLP Procedure for Detecting Known Single Nucleotide and Deletion–Insertion Polymorphisms" International Journal of Molecular Sciences 20, no. 13: 3193. https://doi.org/10.3390/ijms20133193





