Overexpression of Acyl-ACP Thioesterases, CpFatB4 and CpFatB5, Induce Distinct Gene Expression Reprogramming in Developing Seeds of Brassica napus
Abstract
:1. Introduction
2. Results
2.1. Seed FA Profiles (mol%) and 100-Seed Weights Showed
Different Patterns Depending on the Transgene Expressed
2.2. Transcriptome Data Summary
2.3. CpFatB4 and CpFatB5 Were Differentially Expressed during Seed Development
2.4. CpFatB4 Expression under the Napin Promoter Affected Napin Promoter Activity
2.5. CpFatB4 Expression Resulted in an Increase in the Overall FatB/FatA Ratio, but a Clear Decrease in Endogenous FatB/FatA Ratio
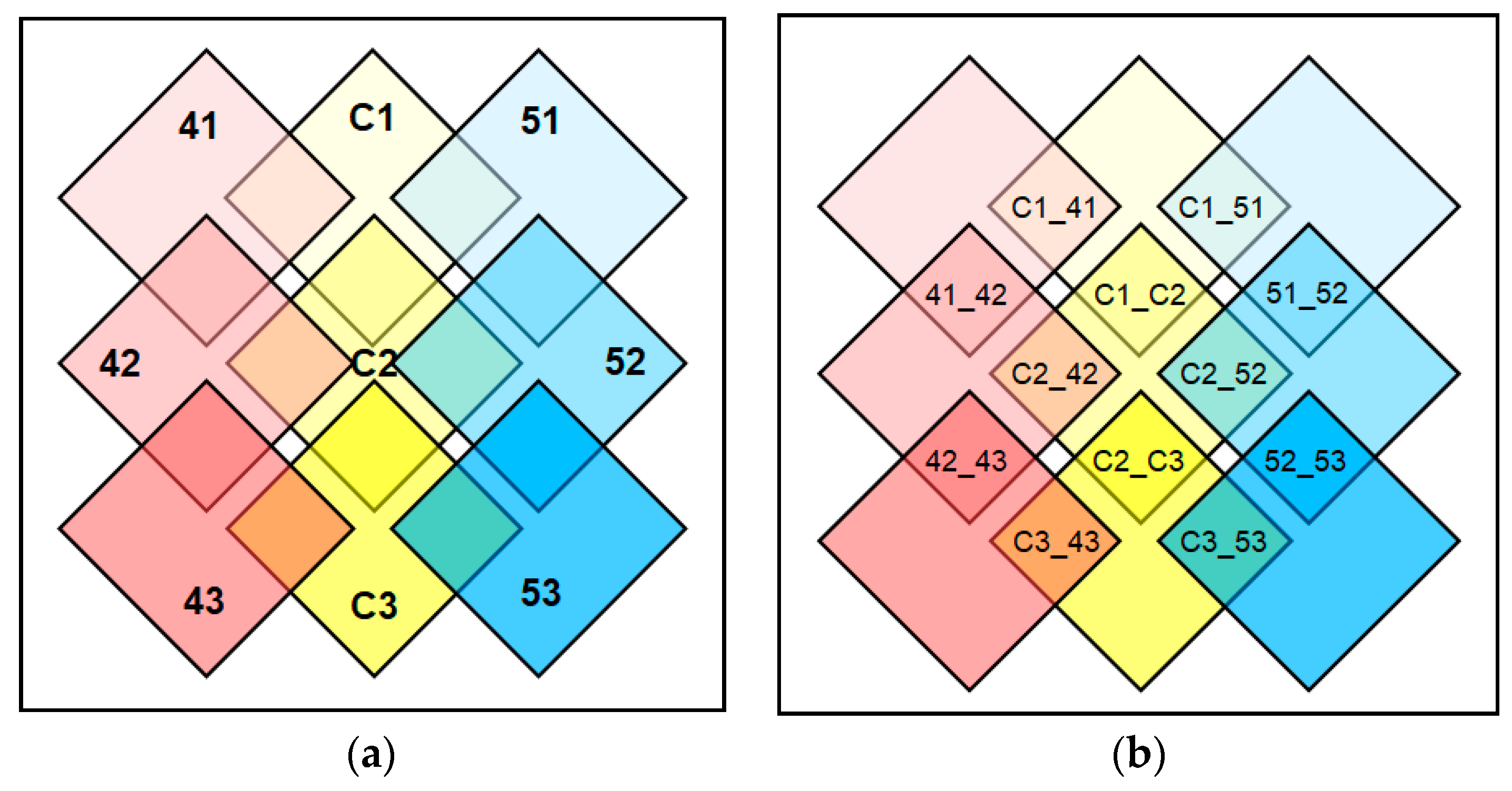
2.6. Depending on Developmental Stages or Genotypes, the Expressed Genes Showed Overlapping yet Distinct Expression Patterns
2.7. Among the Top 20 DEGs, Similarities Were Found in DEGs by Developmental Stages, but No Common Gene Was Identified in DEGs by Genotypes
2.8. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genome (KEGG) Analyses of DEGs Showed Gene Enrichment in Some Categories, Such as “Fatty Acid Biosynthetic Process” and “Glycolysis”, in CpFatB4 at 45 DAF
2.9. Lipid Metabolism DEGs with the Strongest Expression Changes Were Different between CpFatB4 and CpFatB5
2.10. Plastidial FA Synthesis Pathway Was Activated by CpFatB4 Overexpression, but TAG Synthesis Was Not Strongly Affected
2.11. RNA-Seq Results Were Validated by RT-qPCR
3. Discussion
3.1. Whole-Pod Transcriptomes in Transgenic B. napus Showed Similar Developmental Gene Expression Changes to Those of the Control
3.2. The Transcriptome Analyses Provided Comprehensive Gene Expression Changes Caused by 16:0 or 10:0/12:0 FA Accumulation in Seeds of B. napus
3.3. DEGs Detected in CpFatB5 Suggest the Modulation of Organellar Gene Expression Responding to Medium-Chain FA Accumulation
4. Materials and Methods
4.1. Plant Materials
4.2. FA Analysis
4.3. RNA Samples for RNA-Seq and Analysis of DEGs
4.4. Illumina Sequencing, Data Processing, Reads Mapping, and Gene Annotation
4.5. DEG Analysis and Gene Annotation
4.6. GO and KEGG Analysis
4.7. Real-Time Quantitative PCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| FA | Fatty acid |
| ACP | Acyl carrier protein |
| FatB | Fatty acyl-ACP thioesterase B |
| DAF | Days after flowering |
References
- United States Department of Agriculture Foreign Agricultural Service. Available online: https://apps.fas.usda.gov/psdonline/circulars/oilseeds.pdf (accessed on 1 February 2019).
- Sakhno, L.O. Variability in the fatty acid composition of rapeseed oil: Classical breeding and biotechnology. Cytol. Genet. 2010, 44, 389–397. [Google Scholar] [CrossRef]
- Mancini, A.; Imperlini, E.; Nigro, E.; Montagnese, C.; Daniele, A.; Orrù, S.; Buono, P. Biological and nutritional properties of palm oil and palmitic acid: Effects on health. Molecules 2015, 20, 17339–17361. [Google Scholar] [CrossRef] [PubMed]
- The Economist. Available online: https://www.economist.com/briefing/2010/06/24/the-other-oil-spill (accessed on 6 February 2019).
- Bhalla, P.L.; Singh, M.B. Agrobacterium-mediated transformation of Brassica napus and Brassica oleracea. Nat. Protoc. 2008, 3, 181–189. [Google Scholar] [CrossRef] [PubMed]
- Gupta, M.; Bhaskar, P.B.; Sriram, S.; Wang, P.H. Integration of omics approaches to understand oil/protein content during seed development in oilseed crops. Plant Cell Rep. 2017, 36, 637–652. [Google Scholar] [CrossRef] [PubMed]
- Mason, A.S.; Higgins, E.E.; Snowdon, R.J.; Batley, J.; Stein, A.; Werner, C.; Parkin, I.A.P. A user guide to the Brassica 60K Illumina Infinium™ SNP genotyping array. Theor. Appl. Genet. 2017, 130, 621–633. [Google Scholar] [CrossRef] [PubMed]
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.; Tang, H.; Wang, X.; Chiquet, J.; Belcram, H.; Tong, C.; Samans, B.; et al. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 2014, 345, 950–953. [Google Scholar] [CrossRef] [PubMed]
- Troncoso-Ponce, M.A.; Kilaru, A.; Cao, X.; Durrett, T.P.; Fan, J.; Jensen, J.K.; Thrower, N.A.; Pauly, M.; Wilkerson, C.; Ohlrogge, J.B. Comparative deep transcriptional profiling of four developing oilseeds. Plant J. 2011, 68, 1014–1027. [Google Scholar] [CrossRef] [Green Version]
- Roh, K.H.; Park, J.-S.; Kim, J.-B.; Kim, H.U.; Lee, K.R.; Kim, S.H. Gene expression profiling of oilseed rape embryos using microarray analysis. J. Appl. Biol. Chem. 2012, 55, 227–234. [Google Scholar] [CrossRef]
- Xu, H.M.; Kong, X.D.; Chen, F.; Huang, J.X.; Lou, X.Y.; Zhao, J.Y. Transcriptome analysis of Brassica napus pod using RNA-Seq and identification of lipid-related candidate genes. BMC Genom. 2015, 16, 858. [Google Scholar] [CrossRef]
- Chen, J.; Tan, R.K.; Guo, X.J.; Fu, Z.L.; Wang, Z.; Zhang, Z.Y.; Tan, X.L. Transcriptome analysis comparison of lipid biosynthesis in the leaves and developing seeds of Brassica napus. PLoS ONE 2015, 10, e0126250. [Google Scholar] [CrossRef]
- Wan, H.; Cui, Y.; Ding, Y.; Mei, J.; Dong, H.; Zhang, W.; Wu, S.; Liang, Y.; Zhang, C.; Li, J.; et al. Time-series analysis of transcriptome and proteomes reveal molecular networks underlying oil accumulation in Canola. Front. Plant Sci. 2017, 7, 2007. [Google Scholar] [CrossRef] [PubMed]
- Stoll, C.; Luhs, W.; Zarhloul, M.K.; Friedt, W. Genetic modification of saturated fatty acids in oilseed rape (Brassica napus). Eur. J. Lipid Sci. Technol. 2005, 107, 244–248. [Google Scholar] [CrossRef]
- Voelker, T.A.; Hayes, T.R.; Cranmer, A.C.; Davies, H.M. Genetic engineering of a quantitative trait: Metabolic and genetic parameters influencing the accumulation of laurate in rapeseed. Plant J. 1996, 9, 229–241. [Google Scholar] [CrossRef]
- Knutzon, D.S.; Hayes, T.R.; Wyrick, A.; Xiong, H.; Maelor Davies, H.; Voelker, T.A. Lysophosphatidic acid acyltransferase from coconut endosperm mediates the insertion of laurate at the sn-2 position of triacylglycerols in lauric rapeseed oil and can increase total laurate levels. Plant Physiol. 1999, 120, 739–746. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.-Y.; Hammerlindl, J.; Forseille, L.; Zhang, H.; Smith, M.A. Simultaneous over-expressing of an acyl-ACP thioesterase (FatB) and silencing of acyl-acyl carrier protein desaturase by artificial microRNAs increases saturated fatty acid levels in Brassica napus seeds. Plant Biotechnol. J. 2014, 12, 624–637. [Google Scholar] [CrossRef]
- Jones, A.; Davies, H.M.; Voelker, T.A. Palmitoyl-acyl carrier protein (ACP) thioesterase and the evolutionary origin of plant acyl-ACP thioesterases. Plant Cell 1995, 7, 359–371. [Google Scholar] [CrossRef]
- Bonaventure, G.; Salas, J.J.; Pollard, M.R.; Ohlrogge, J.B. Disruption of the FATB gene in arabidopsis demonstrates an essential role of saturated fatty acids in plant growth. Plant Cell 2003, 15, 1020–1033. [Google Scholar] [CrossRef]
- Huang, J.; Zhang, T.; Zhang, Q.; Chen, M.; Wang, Z.; Zheng, B.; Xia, G.; Yang, X.; Huang, C.; Huang, Y. The mechanism of high contents of oil and oleic acid revealed by transcriptomic and lipidomic analysis during embryogenesis in Carya cathayensis Sarg. BMC Genom. 2016, 17, 113. [Google Scholar] [CrossRef]
- Graham, S.A.; Knapp, S.J. Cuphea: A new plant source of medium-chain fatty acids. Crit. Rev. Food Sci. Nutr. 1989, 28, 139–173. [Google Scholar] [CrossRef]
- Graham, S.A.; Freudenstein, J.V.; Luker, M. A phylogenetic study of Cuphea (Lythraceae) based on morphology and nuclear rDNA ITS sequences. Syst. Bot. 2006, 31, 764–778. [Google Scholar] [CrossRef]
- Dehesh, K.; Jones, A.; Knutzon, D.S.; Voelker, T.A. Production of high levels of 8:0 and 10:0 fatty acids in transgenic canola by overexpression of ChFatB2, a thioesterase cDNA from Cuphea hookeriana. Plant J. 1996, 9, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Thompson, A.E.; Kleiman, R. Effect of seed maturity on seed oil, fatty acid and crude protein content of eight Cuphea species. J. Am. Oil Chem. Soc. 1988, 65, 139–146. [Google Scholar] [CrossRef]
- Krishnakumar, V.; Contrino, S.; Cheng, C.Y.; Belyaeva, I.; Ferlanti, E.S.; Miller, J.R.; Vaughn, M.W.; Micklem, G.; Town, C.D.; Chan, A.P. ThaleMine: A warehouse for arabidopsis data integration and discovery. Plant Cell Physiol. 2017, 58, e4. [Google Scholar] [CrossRef] [PubMed]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef] [PubMed]
- Crouch, M.L.; Sussex, I.M. Development and storage-protein synthesis in Brassica napus L. embryos in vivo and in vitro. Planta 1981, 153, 64–74. [Google Scholar] [CrossRef] [PubMed]
- Loader, N.M.; Woolner, E.M.; Hellyer, A.; Slabas, A.R.; Safford, R. Isolation and characterization of two Brassica napus embryo acyl-ACP thioesterase cDNA clones. Plant Mol. Biol. 1993, 23, 769–778. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016, 44, D457–D462. [Google Scholar] [CrossRef]
- Jing, F.; Cantu, D.C.; Tvaruzkova, J.; Chipman, J.P.; Nikolau, B.J.; Yandeau-Nelson, M.; Reilly, P.J. Phylogenetic and experimental characterization of an acyl-ACP thioesterase family reveals significant diversity in enzymatic specificity and activity. BMC Biochem. 2011, 12, 44. [Google Scholar] [CrossRef]
- Bennett, E.J.; Roberts, J.A.; Wagstaff, C. The role of the pod in seed development: Strategies for manipulating yield. New Phytol. 2011, 190, 838–853. [Google Scholar] [CrossRef]
- Huang, K.L.; Zhang, M.L.; Ma, G.J.; Wu, H.; Wu, X.M.; Ren, F.; Li, X.B. Transcriptome profiling analysis reveals the role of silique in controlling seed oil content in Brassica napus. PLoS ONE 2017, 12, e0179027. [Google Scholar] [CrossRef]
- Deng, W.; Yan, F.; Zhang, X.; Tang, Y.; Yuan, Y. Transcriptional profiling of canola developing embryo and identification of the important roles of BnDof5.6 in embryo development and fatty acids synthesis. Plant Cell Physiol. 2015, 56, 1624–1640. [Google Scholar] [CrossRef] [PubMed]
- Slocombe, S.P.; Cummins, I.; Jarvis, R.P.; Murphy, D.J. Nucleotide sequence and temporal regulation of a seed-specific Brassica napus cDNA encoding a stearoyl-acyl carrier protein (ACP) desaturase. Plant Mol. Biol. 1992, 20, 151–155. [Google Scholar] [CrossRef] [PubMed]
- Slocombe, S.P.; Piffanelli, P.; Fairbairn, D.; Bowra, S.; Hatzopoulos, P.; Tsiantis, M.; Murphy, D.J. Temporal and tissue-specific regulation of a Brassica napus stearoyl-acyl carrier protein desaturase gene. Plant Physiol. 1994, 104, 1167–1176. [Google Scholar] [CrossRef] [PubMed]
- Andre, C.; Haslam, R.P.; Shanklin, J. Feedback regulation of plastidic acetyl-CoA carboxylase by 18:1-acyl carrier protein in Brassica napus. Proc. Natl. Acad. Sci. USA 2012, 109, 10107–10112. [Google Scholar] [CrossRef] [PubMed]
- Eccleston, V.S.; Ohlrogge, J.B. Expression of lauroyl-acyl carrier protein thioesterase in Brassica napus seeds induces pathways for both fatty acid oxidation and biosynthesis and implies a set point for triacylglycerol accumulation. Plant Cell 1998, 10, 613–622. [Google Scholar] [CrossRef]
- Madoka, Y.; Tomizawa, K.; Mizoi, J.; Nishida, I.; Nagano, Y.; Sasaki, Y. Chloroplast transformation with modified accD operon increases acetyl-CoA carboxylase and causes extension of leaf longevity and increase in seed yield in tobacco. Plant Cell Physiol. 2002, 43, 1518–1525. [Google Scholar] [CrossRef]
- Bates, P.D.; Johnson, S.R.; Cao, X.; Li, J.; Nam, J.W.; Jaworski, J.G.; Ohlrogge, J.B.; Browse, J. Fatty acid synthesis is inhibited by inefficient utilization of unusual fatty acids for glycerolipid assembly. Proc. Natl. Acad. Sci. USA 2014, 111, 1204–1209. [Google Scholar] [CrossRef] [Green Version]
- Guan, M.; Li, X.; Guan, C. Microarray analysis of differentially expressed genes between Brassica napus strains with high- and low-oleic acid contents. Plant Cell Rep. 2012, 31, 929–943. [Google Scholar] [CrossRef]
- Moire, L.; Rezzonico, E.; Goepfert, S.; Poirier, Y. Impact of unusual fatty acid synthesis on futile cycling through beta-oxidation and on gene expression in transgenic plants. Plant Physiol. 2004, 134, 432–442. [Google Scholar] [CrossRef]
- Lee, K.R.; Kim, E.H.; Roh, K.H.; Kim, J.B.; Kang, H.C.; Go, Y.S.; Suh, M.C.; Kim, H.U. High-oleic oilseed rapes developed with seed-specific suppression of FAD2 gene expression. J. Appl. Biol. Chem. 2016, 59, 669–676. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [PubMed]
- Yi, X.; Du, Z.; Su, Z. PlantGSEA: A gene set enrichment analysis toolkit for plant community. Nucleic Acids Res. 2013, 41, W98–W103. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Wu, G.; Cao, Y.; Wu, Y.; Xiao, L.; Li, X.; Lu, C. Breeding response of transcript profiling in de eloping seeds of Brassica napus. BMC Mol. Biol. 2009, 10, 49. [Google Scholar] [CrossRef] [PubMed]








| Mol % | Youngsan | CpFatB4 | CpFatB5 |
|---|---|---|---|
| 10:0 | 0.0 | 0.1 | 2.1 ± 0.3 |
| 12:0 | 0.0 | 0.1 | 1.0 ± 0.1 |
| 14:0 | 0.2 | 1.0 ± 0.1 | 0.6 |
| 16:0 | 5.3 ± 0.1 | 22.7 ± 1.7 | 9.0 ± 0.2 |
| 18:0 | 2.3 ± 0.1 | 3.4 ± 0.1 | 2.5 ± 0.1 |
| 18:1 | 67.8 ± 0.7 | 49.9±1.8 | 61.8 ± 0.7 |
| 18:2 | 16.6 ± 0.6 | 15.4 ± 0.7 | 15.7 ± 0.6 |
| 18:3 | 5.3 ± 0.4 | 4.7 ± 0.2 | 4.9 ± 0.2 |
| 20:0 | 0.8 ± 0.0 | 1.2 ± 0.0 | 0.8 ± 0.1 |
| 20:1 | 1.3 ± 0.0 | 1.0 ± 0.0 | 1.2 ± 0.0 |
| 22:1 | 0.3 ± 0.0 | 0.5 ± 0.0 | 0.4 ± 0.0 |
| Percentage of saturated fatty acids | 8.7 | 28.4 | 16.0 |
| 100-seed weight (mg) | 227.4 ± 9.9 | 250.1 ± 9.3 | 216.1 ± 8.9 |
| Type and Name of Comparisons | DEG Numbers | Up-Regulated DEG Numbers | Down-Regulated DEG Numbers | ||||
|---|---|---|---|---|---|---|---|
| Total | Lipid Metabolism Genes (%) | Total | Lipid Metabolism Genes (%) | Total | Lipid Metabolism Genes (%) | ||
| Stage | C1_C2 | 2550 | 250 (9.8) | 1688 | 159 (9.4) | 862 | 91 (10.6) |
| C2_C3 | 4642 | 267 (5.8) | 2504 | 100 (4.0) | 2138 | 167 (7.8) | |
| 41_42 | 2486 | 244 (9.8) | 1829 | 172 (9.4) | 657 | 72 (11.0) | |
| 42_43 | 2445 | 207 (8.5) | 1028 | 72 (7.0) | 1417 | 135 (9.5) | |
| 51_52 | 2575 | 216 (8.4) | 1692 | 141 (8.3) | 883 | 75 (8.5) | |
| 52_53 | 4813 | 264 (5.5) | 1753 | 64 (3.7) | 3060 | 200 (6.5) | |
| Line | C1_41 | 366 | 10 (2.7) | 174 | 0 (0) | 192 | 10 (5.2) |
| C2_42 | 45 | 0 (0) | 24 | 0 (0) | 21 | 0 (0) | |
| C3_43 | 1847 | 105 (5.7) | 676 | 83 (12.3) | 1171 | 22 (1.9) | |
| C1_51 | 289 | 6 (2.1) | 155 | 0 (0) | 134 | 6 (4.5) | |
| C2_52 | 112 | 6 (5.4) | 12 | 1 (8.3) | 100 | 5 (5.0) | |
| C3_53 | 1365 | 48 (3.5) | 324 | 3 (0.9) | 1041 | 45 (4.3) | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nam, J.-W.; Yeon, J.; Jeong, J.; Cho, E.; Kim, H.B.; Hur, Y.; Lee, K.-R.; Yi, H. Overexpression of Acyl-ACP Thioesterases, CpFatB4 and CpFatB5, Induce Distinct Gene Expression Reprogramming in Developing Seeds of Brassica napus. Int. J. Mol. Sci. 2019, 20, 3334. https://doi.org/10.3390/ijms20133334
Nam J-W, Yeon J, Jeong J, Cho E, Kim HB, Hur Y, Lee K-R, Yi H. Overexpression of Acyl-ACP Thioesterases, CpFatB4 and CpFatB5, Induce Distinct Gene Expression Reprogramming in Developing Seeds of Brassica napus. International Journal of Molecular Sciences. 2019; 20(13):3334. https://doi.org/10.3390/ijms20133334
Chicago/Turabian StyleNam, Jeong-Won, Jinouk Yeon, Jiseong Jeong, Eunyoung Cho, Ho Bang Kim, Yoonkang Hur, Kyeong-Ryeol Lee, and Hankuil Yi. 2019. "Overexpression of Acyl-ACP Thioesterases, CpFatB4 and CpFatB5, Induce Distinct Gene Expression Reprogramming in Developing Seeds of Brassica napus" International Journal of Molecular Sciences 20, no. 13: 3334. https://doi.org/10.3390/ijms20133334





