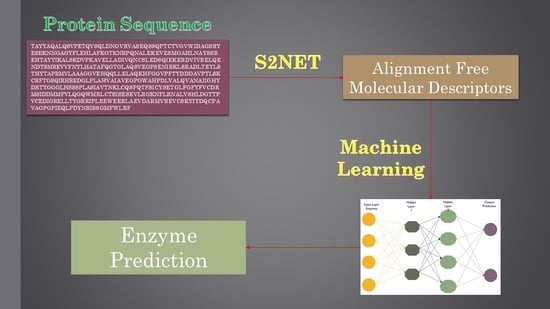
1. Introduction
By the late 1950s, the International Union of Biochemistry and Molecular Biology foresaw the need for unique nomenclature for enzymes. In those years, the number of known enzymes had grown very rapidly and, because of the absence of general guidelines, the nomenclature of the enzymes was getting out of hand. In some cases, enzymes with similar names were catalyzing different reactions, while conversely different names were given to the same or similar enzymes. Due to this, during the third International Congress of Biochemistry in Brussels in August 1955, the General Assembly of the International Union of Biochemistry (IUB) decided to establish an International Commission in charge of developing a nomenclature for enzymes. In 1961, the IUB finally released the first version of the Enzyme Classification (EC) and Nomenclature List. This nomenclature was based on assigning a four number code to enzymes with the following meaning: (i) the first number identifies the main enzyme class; (ii) the second digit indicates the subclass; (iii) the third number denotes the sub-subclass; and (iv) the fourth digit is the serial number of the enzyme in its sub-subclass. Six enzyme classes were identified, with the classification based on the type of reaction catalyzed: oxidoreductases (EC 1), transferases (EC 2), hydrolases (EC 3), lyases (EC 4), isomerases (EC 5) and ligases (EC 6) [
1]. Although several revisions have been made to the 1961 version, the six classes identified have not received any change. However, in August 2018, a new class was added. This new class contains the translocases (EC 7), and was added to describe those enzymes catalyzing the movement of ions or molecules across membranes or their separation within membranes. For this reason, some enzymes which had previously been classified in other classes—EC 3.6.3 for example—were now included in the EC 7 class.
Predicting enzyme classes or protein function using bioinformatic tools is still a key goal in bioinformatics and computational biology due to both the prohibitive costs and the time-consuming nature of wet-lab-based functional identification procedures. In point of fact, there are more than four thousand sequences whose function remains unknown so far and this number is still growing [
2]. The problem is that our ability to assign a specific function to a sequence is far lower than our ability to isolate and identify sequences. For this reason, significant efforts have been devoted to developing reliable methods able to predict protein function.
Several methodological strategies and tools have been proposed to classify enzymes based on different approaches [
3,
4,
5,
6,
7,
8,
9,
10]. The Basic Local Alignment Search Tool (BLAST) [
11] is likely to be one of the most powerful and used tools which finds regions of similarity between biological sequences. The program compares nucleotide or protein sequences to sequence databases and calculates their statistical significance. However, as is the case with all methods, these procedures may fail under certain conditions. In some cases, enzymes with a sequence similarity higher than 90% may belong to different enzyme families and, thus, have different EC annotations [
12,
13,
14]. On the other hand, some enzymes which share the same first EC number may have a sequence similarity below 30%. Some authors have described this situation well and highlighted the need to develop alignment-free methods, which may be used in a complementary way [
15,
16]. Other relevant tools based on sequence similarity are the UniProtKB database [
17], the Kyoto Encyclopedia of Genes and Genomes (KEGG) [
18], the PEDANT protein database [
19], DEEPre [
20], ECPred [
21] and EzyPred [
22]. DEEPre is a three-level EC number predictor, which predicts whether an input protein sequence is an enzyme, and its main class and subclass if it is. This method is based on a dataset of 22,198 sequences achieving an overall accuracy of more than 90%. ECPred is another enzymatic function prediction tool based on an ensemble of machine learning classifiers. The creators of this tool developed it using a dataset of approximately 245,000 proteins, achieving score classifications in the 6 EC classes and subclasses like the ones reported by DEEPre. EzyPred is a top-down approach for predicting enzyme classes and subclasses. This model was developed using a 3-layer predictor using the ENZYME [
23] dataset (approximately 9800 enzymes when the model was developed), which was able to achieve an overall accuracy above 86%. Other relevant methods with similar classification scores have also been reported [
10,
15,
20,
24,
25]. All these methods have proved to be robust; however, they are all outdated since they cannot predict the EC 7 classification, and should therefore be updated in accordance with the new EC class.
In light of what has been referred to so far, the major target of this work was to develop an alignment-free strategy using machine learning (ML) methods to predict the first two digits of the seven EC classes. Previous ML methods have used alignment-free numerical parameters to quantify information about the 2D or 3D structure of proteins [
26,
27,
28,
29]. Specifically, Graham, Bonchev, Marrero-Ponce, and others [
30,
31,
32,
33,
34] used Shannon’s entropy measures to quantify relevant structural information about molecular systems. In addition, González-Díaz et al. [
35,
36,
37] introduced so-called Markov–Shannon entropies (
θk) to codify the structural information of large bio-molecules and complex bio-systems or networks. For comparative purposes, we developed different linear and non-linear models, including a linear discriminant analysis (LDA) and various types of artificial neural networks (ANNs). In addition, we focused our work on performing an efficient feature selection (FS). Nowadays, there are several software packages or tools that may be used to calculate thousands of molecular descriptors (MDs). As a result, a proper FS method is essential to develop robust and reliable quantitative structure–activity relationship (QSAR) models. This is particularly the case when using ANNs, since QSAR models developed with a large set of MDs are really complex, vulnerable to overfitting and difficult to obtain a mechanistic interpretation from [
38,
39].
3. Discussion
The main aim of this study was to develop a new QSAR-ML model able to predict enzyme subclasses considering the new and recently introduced EC class 7. We retrieved from the Protein Data Bank (PDB) more than 26,000 enzyme and 55,000 non-enzyme sequences in order to build up our dataset. All of the enzyme sequences belonged to one of the 7 main classes and 65 subclasses. The EC 7 class was introduced just few months ago and, due to this, all of the current models do not include this new enzyme class. As a result, the classification or prediction such models are performing may be misleading. Hence, the development of new models which are capable of predicting all enzyme classes and subclasses—including the EC 7 class—are of utmost importance. In view of this, we developed a new machine learning model able to discriminate between enzymes and non-enzymes. In addition, the model was capable of assigning enzymes to a specific enzyme subclass. We generated linear and non-linear models using alignment-free variables to find the best model to predict EC classes and subclasses. The results of the linear model were impressive since with only four MDs the model could discriminate between enzymes and non-enzymes, as well as assign a specific EC class and subclass to each enzyme sequence. We checked the accuracy and robustness of the model and the results clearly indicate that the model is reliable. Regarding the validation, we performed a classical cross-validation procedure using 30% of the dataset. This led to almost the same results for the training and validation sets, indicating once more the robustness of the model and approach.
Although the accuracy of the derived LDA model was near 100%, we decided to further test our approach by developing some neural network models, which usually improve LDA results. To the best of our knowledge, an MLP is generally considered the best ANN algorithm and, in this case, had the potential to improve our linear model. As previously reported, the MLP was able to perfectly discriminate between enzymes and non-enzymes, in addition to assigning each enzyme sequence to a specific subclass. It is also remarkable that the best model only needed nine neurons in the hidden layer. This low number of neurons, considering the number of sequences and variables, suggest that the model is not suffering from an overfitting problem. Mechanistic interpretation of ANN models is always a challenging task since these models do not lead to simple linear equations. A sensitivity analysis may then be used to analyze the influence of each MD on the model. For the ANN model, we carried out such an analysis to evaluate the weight of each variable in the model. This analysis is also useful for identifying redundant variables in models, assisting in their eliminatation to avoid an unlikely overfitting problem. In the case of the ANN model, we identified that the same four variables used in the LDA model were able to perfectly discriminate between enzymes and non-enzymes and assign each enzyme sequence to a specific subclass.
Finally, we also tested RBF models, which afforded results that were worse than the MLP models. In fact, the general accuracy was lower when compared to the MLP models, which usually need less neurons to achieve greater accuracy.
5. Conclusions
Developing new, reliable, and robust methods for predicting protein function and enzyme class and subclasses is a key goal for theoreticians, especially in light of the recently introduced EC 7 class. In this work, we developed linear and non-linear models using an alignment-free approach to discriminate between enzymes and non-enzymes, as well as assign each enzyme sequence to a specific EC class. The best LDA model showed an overall accuracy of 98.63%, which is considered a remarkable result. However, we decided to explore further and develop some non-linear models using two different algorithms: MLP and RBF. While the latter was unable to improve the results of the LDA model, the MLP model was able to achieve an overall accuracy of 100%. This means that it was able to perfectly discriminate between enzymes and non-enzymes and identify the EC class of each enzyme.