Identifying Metabolic Perturbations and Toxic Effects of Rac-Metalaxyl and Metalaxyl-M in Mice Using Integrative NMR and UPLC-MS/MS Based Metabolomics
Abstract
:1. Introduction
2. Results
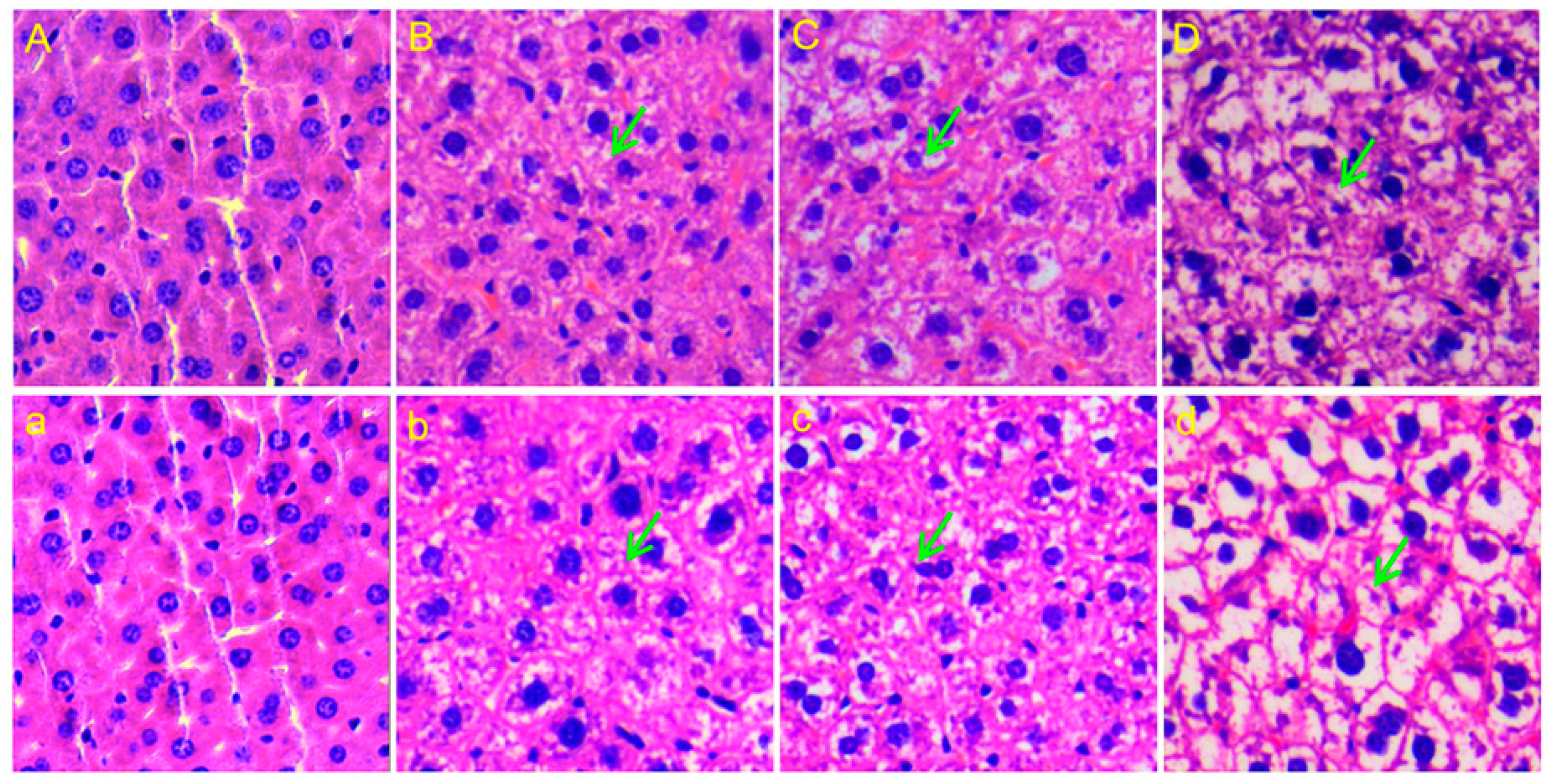
2.1. Body Weight and Histopathology
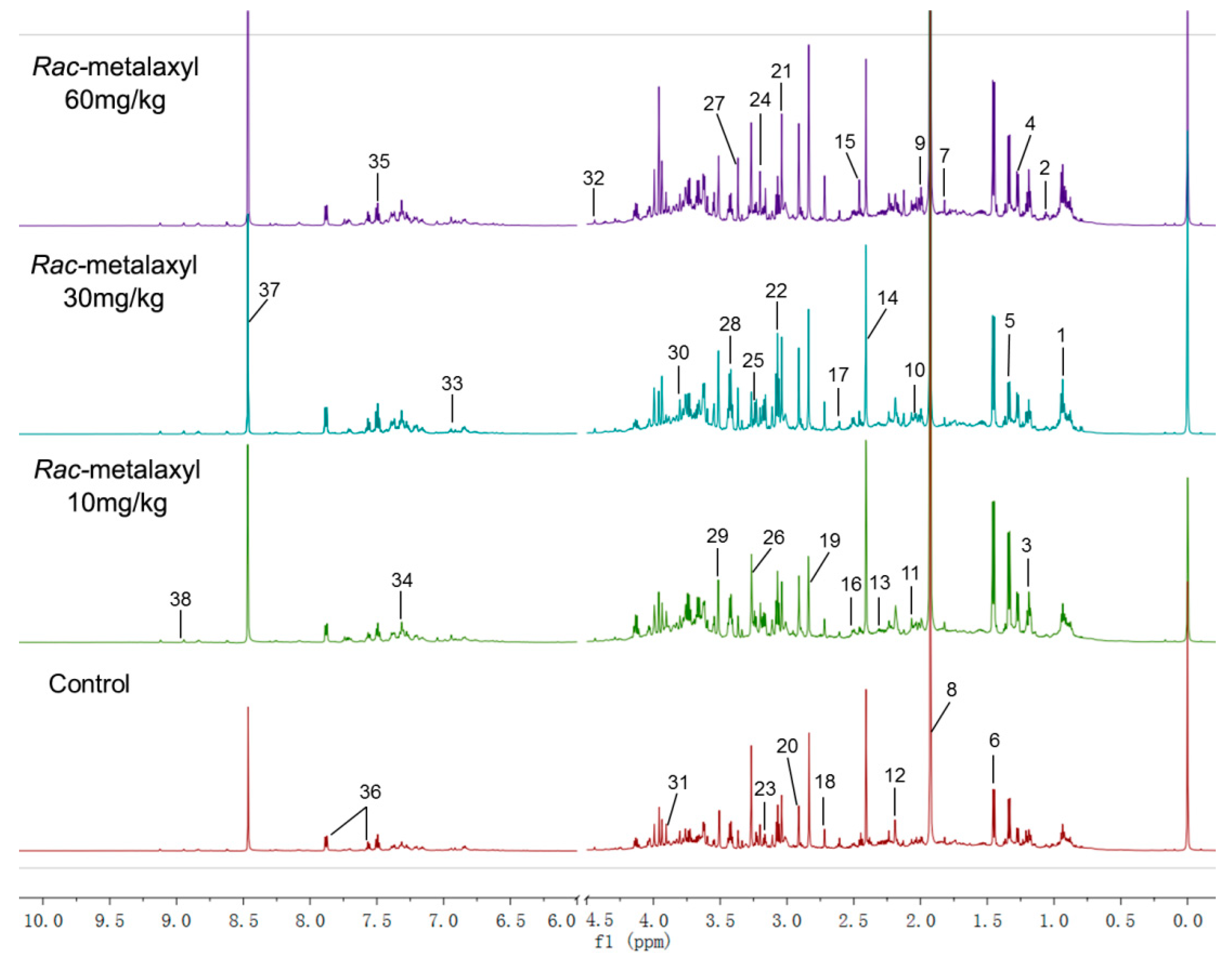
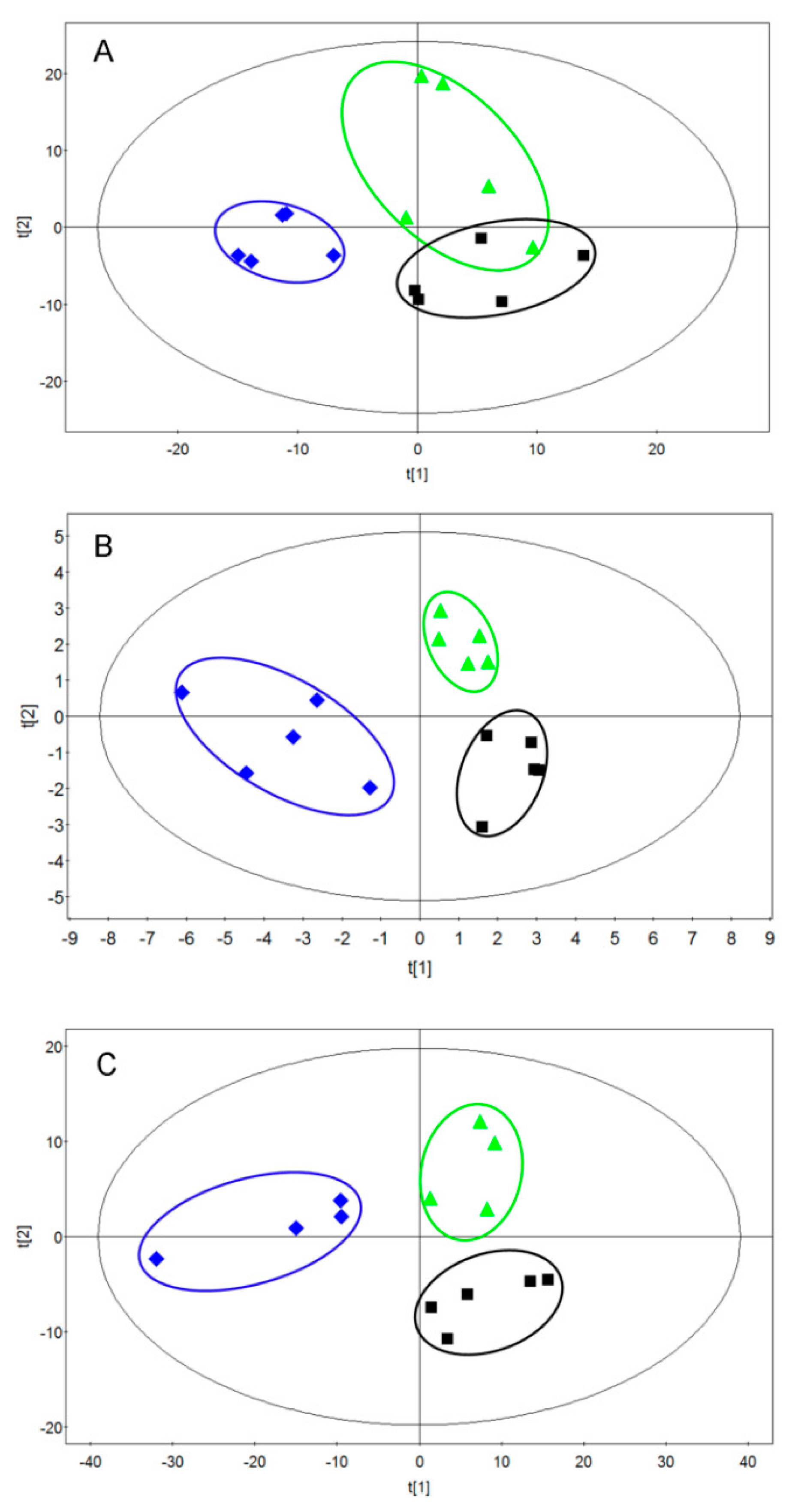
2.2. Metabolomic Alterations
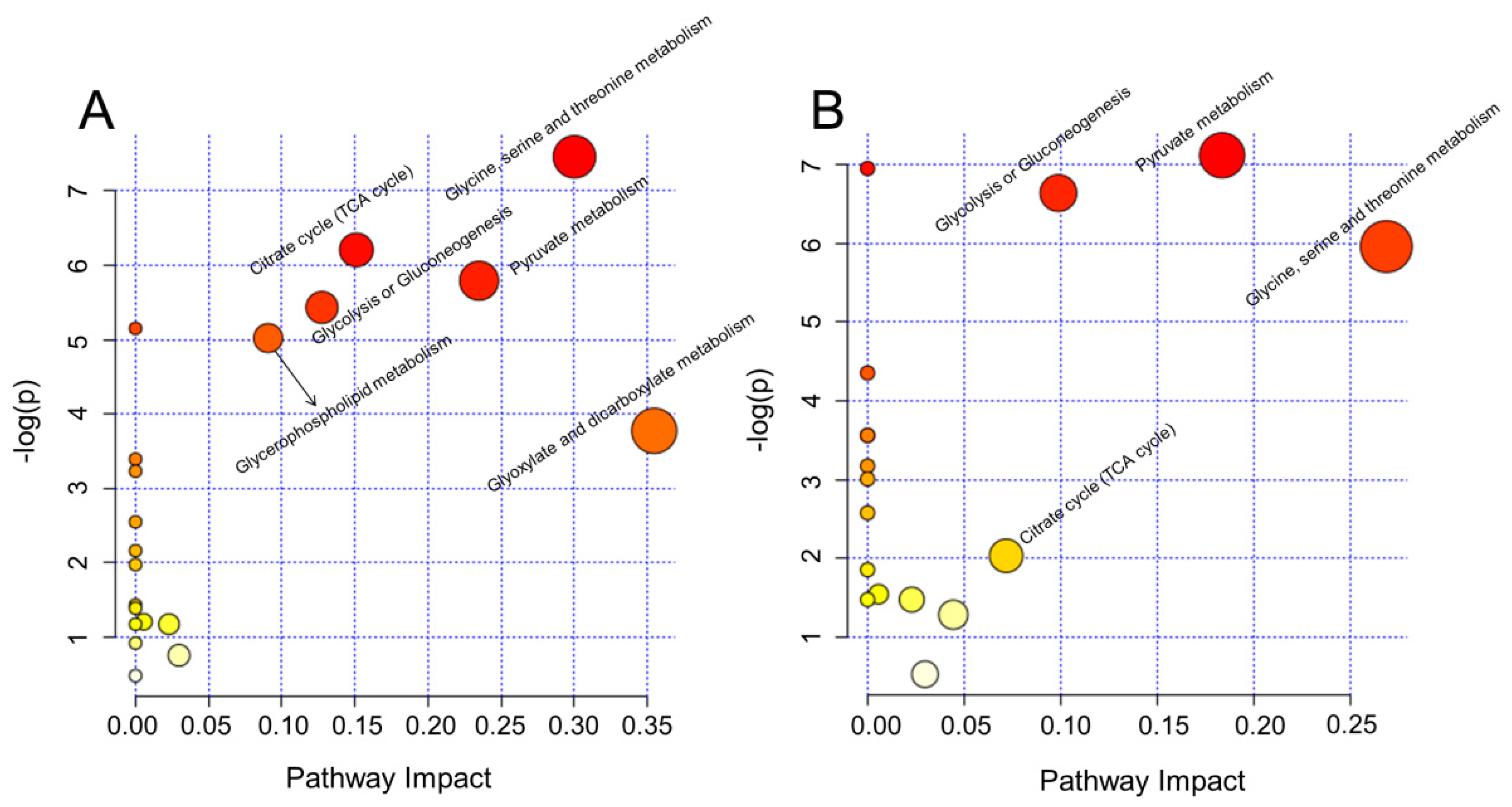
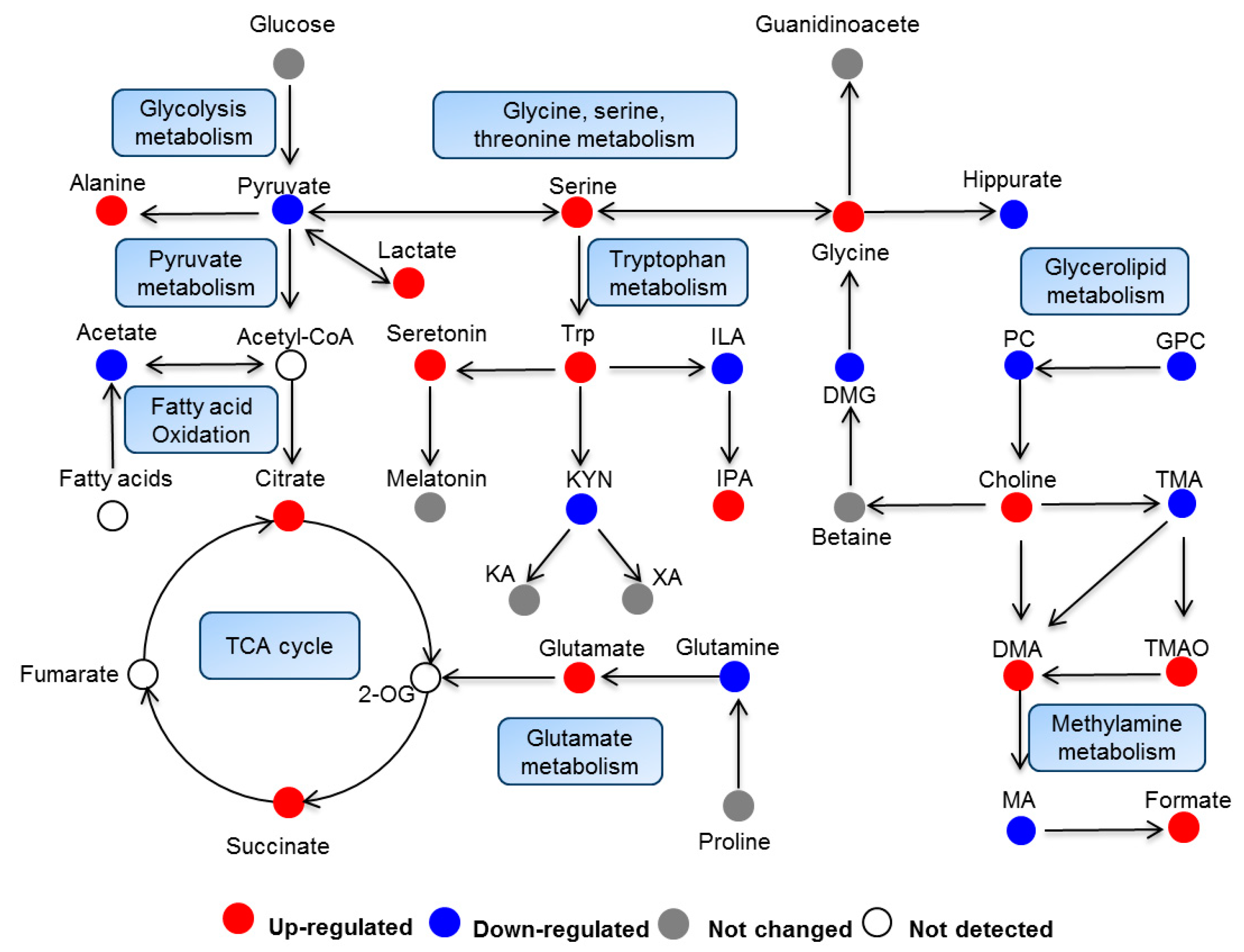
2.3. Biological Pathway Changes
2.4. Serum Amino Acids Changes
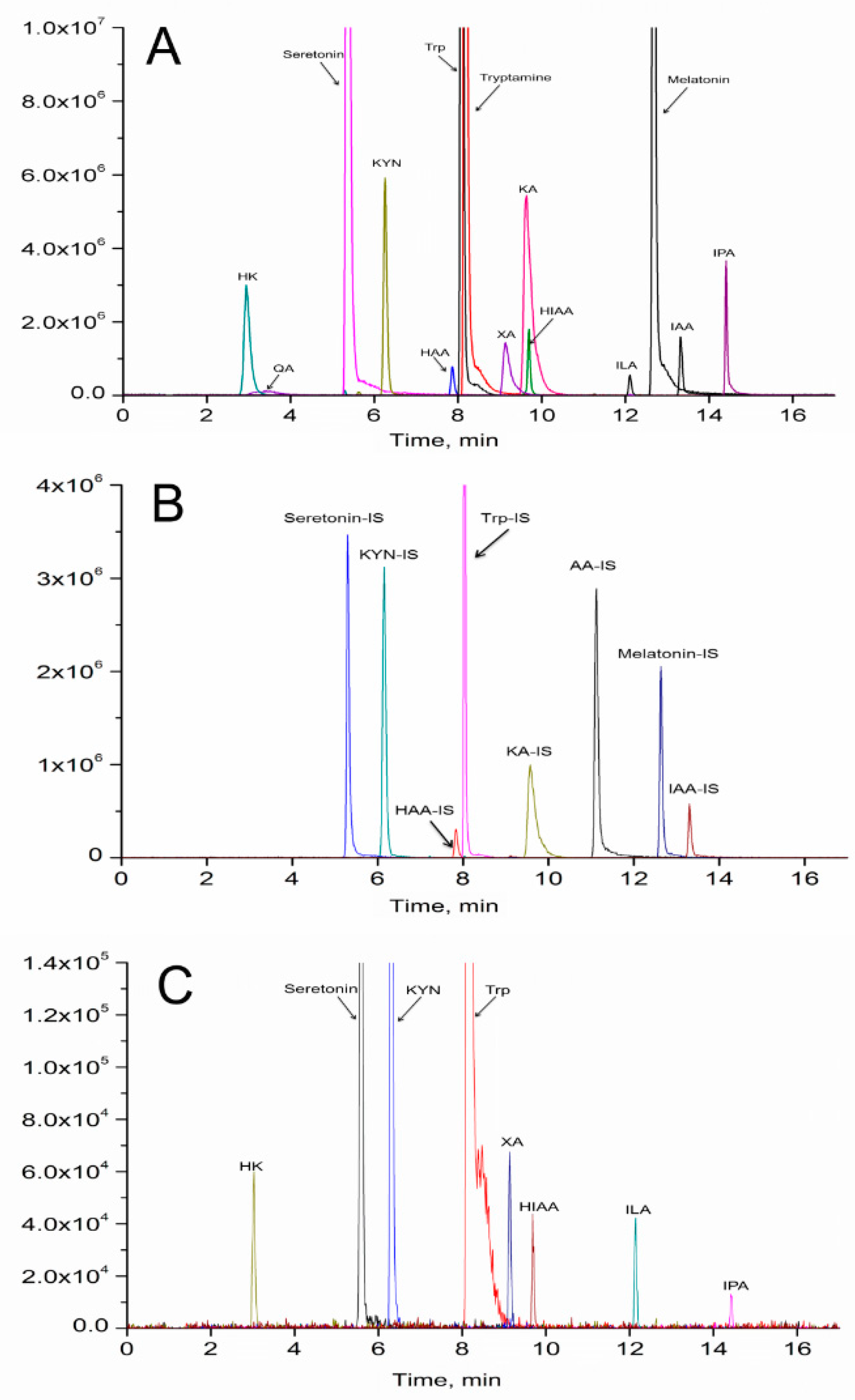
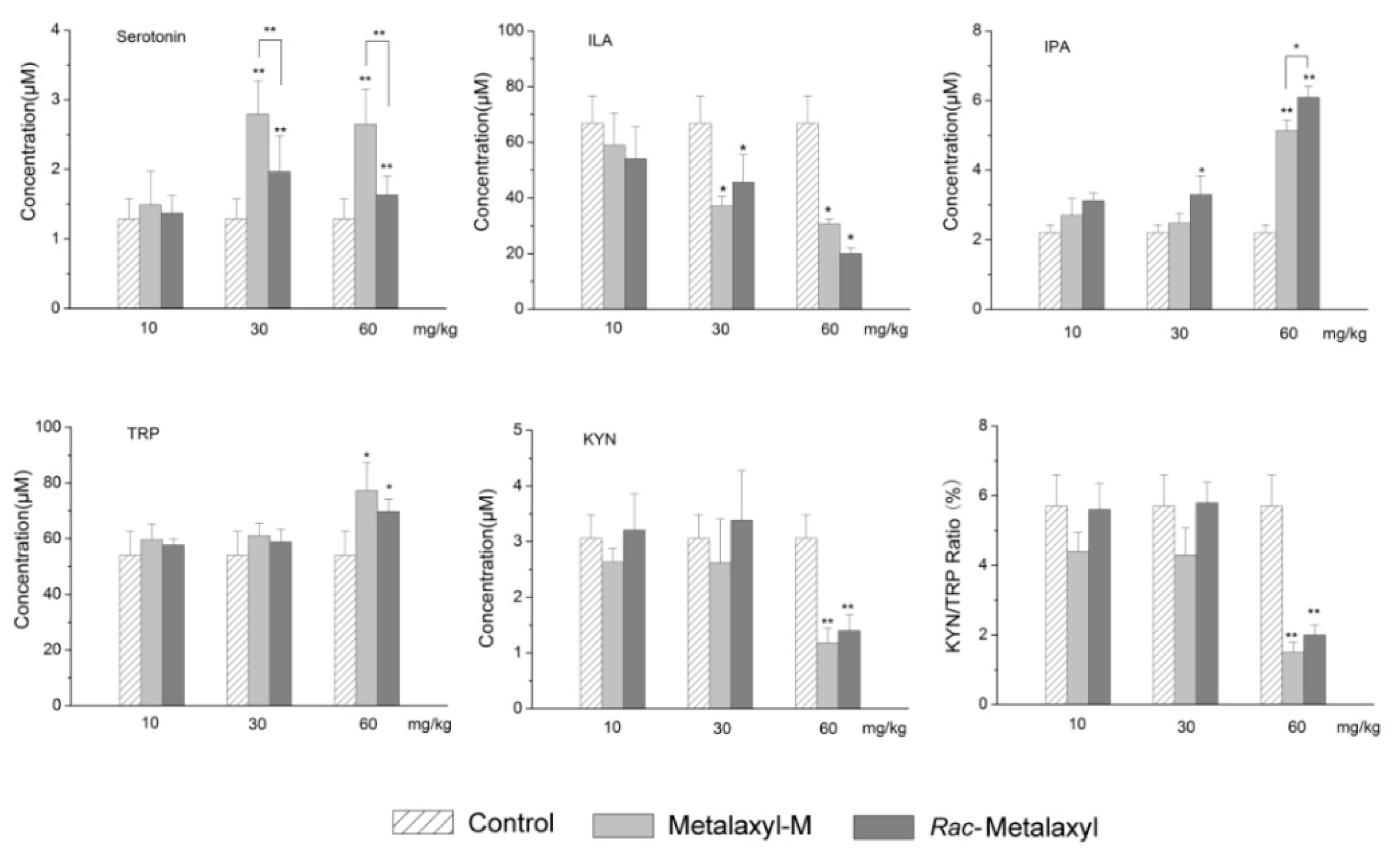
2.5. Tryptophan Metabolites Changes
3. Discussion
3.1. Body Weight Changes and Liver Damage
3.2. Energy Metabolism
3.3. Lipid Metabolism
3.4. Gut Microbiota Metabolism
3.5. Amino Acids Metabolism
3.6. Tryptophan Metabolism
4. Materials and Methods
4.1. Chemicals and Materials
4.2. Animals and Animal Treatments
4.3. Histopathology
4.4. Metabolomic Profilings
4.5. UPLC-MS/MS Analysis
4.6. Biological Pathway Analysis
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| UPLC-MS/MS | Ultra performance liquid chromatography-tandem mass spectrometry |
| NMR | Nuclear magnetic resonance |
| PLS-DA | Partial least-squares discriminant analysis |
| LOQ | Limit of quantitation |
| TCA | Tricarboxylic acid |
| COA | Acetyl-coenzyme A |
| MA | Methylamine |
| DMA | Dimethylamine |
| TMA | Trimethylamine |
| DMG | Dimethylglycine |
| PC | Phosphorylcholine |
| GPC | Glycerophosphocholine |
| TMAO | Trimethylamine N-oxided |
References
- Tomlin, C.D.S. The Pesticide Manual, 15th ed.; British Crop Protection Council: Farnham, UK, 2009. [Google Scholar]
- Di, S.; Liu, T.; Diao, J.; Zhou, Z. Enantioselective bioaccumulation and degradation of sediment-associated metalaxyl enantiomers in Tubifex tubifex. J. Agric. Food Chem. 2013, 61, 4997–5002. [Google Scholar] [CrossRef] [PubMed]
- Abass, K.; Reponen, P.; Jalonen, J.; Pelkonen, O. In vitro metabolism and interactions of the fungicide metalaxyl in human liver preparations. Environ. Toxicol. Pharmacol. 2007, 23, 39–47. [Google Scholar] [CrossRef] [PubMed]
- Lu, H. Stereoselectivity in drug metabolism. Expert Opin. Drug Metab. Toxicol. 2007, 3, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Buerge, I.I.; Poiger, T.; Muller, M.D.; Buser, H.R. Enantioselective degradation of metalaxyl in soils: Chiral preference changes with soil pH. Environ. Sci. Technol. 2003, 37, 2668–2674. [Google Scholar] [CrossRef]
- Blaser, H.-U.; Hanreich, R.; Schneider, H.-D.; Spindler, F.; Steinacher, B. The Chiral Switch of Metolachlor: The Development of a Large-Scale Enantioselective Catalytic Process; Wiley-VCH Verlag GmbH & Co. KGaA: Weinheim, Germany, 2004; pp. 55–70. [Google Scholar]
- Buser, H.-R.; Muller, M.D.; Poiger, T.; Balmer, M.E. Environmental Behavior of the Chiral Acetamide Pesticide Metalaxyl:Enantioselective Degradation and Chiral Stability in Soil. Environ. Sci. Technol. 2001, 36, 221–226. [Google Scholar] [CrossRef]
- Hrelia, P.; Maffei, F.; Fimognari, C.; Vigagni, F.; Cantelli-Forti, G. Cytogenetic effects of Metalaxyl on human and animal chromosomes. Mutat. Res. 1996, 369, 81–86. [Google Scholar] [CrossRef]
- Paolini, M.; Mesirca, R.; Pozzetti, L.; Sapone, A.; Cantelli-Forti, G. Biomarkers of effect in evaluating metalaxyl cocarcinogenesis. Selective induction of murine CYP 3A isoform. Mutat. Res. 1996, 361, 157–164. [Google Scholar] [CrossRef]
- Sakr, S.A.; Badawy, G.M. Effect of ginger (Zingiber officinale R.) on metiram-inhibited spermatogenesis and induced apoptosis in albino mice. J. Appl. Pharm. Sci. 2011, 1, 131–136. [Google Scholar]
- Bundy, J.G.; Davey, M.P.; Viant, M.R. Environmental metabolomics: A critical review and future perspectives. Metabolomics 2009, 5, 3–21. [Google Scholar] [CrossRef]
- Garcia-Sevillano, M.A.; Garcia-Barrera, T.; Gomez-Ariza, J.L. Environmental metabolomics: Biological markers for metal toxicity. Electrophoresis 2015, 36, 2348–2365. [Google Scholar] [CrossRef]
- Wang, P.; Ng, Q.; Zhang, B.; Wei, Z.; Hassan, M.; He, Y.; Ong, C.N. Employing multi-omics to elucidate the hormetic response against oxidative stress exerted by nC60 on Daphnia pulex. Environ. Pollut. 2019, 251, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Ong, E.S.; Chor, C.F.; Zou, L.; Ong, C.N. A multi-analytical approach for metabolomic profiling of zebrafish (Danio rerio) livers. Mol. Biosyst. 2009, 5, 288–298. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Ng, Q.X.; Zhang, H.; Zhang, B.; Ong, C.N.; He, Y. Metabolite changes behind faster growth and less reproduction of Daphnia similis exposed to low-dose silver nanoparticles. Ecotoxicol. Environ. Saf. 2018, 163, 266–273. [Google Scholar] [CrossRef]
- Zhang, Y.; Huang, K.; Deng, Y.; Zhao, Y.; Wu, B.; Xu, K.; Ren, H. Evaluation of the toxic effects of municipal wastewater effluent on mice using omic approaches. Environ. Sci. Technol. 2013, 47, 9470–9477. [Google Scholar] [CrossRef]
- Xu, H.D.; Wang, J.S.; Li, M.H.; Liu, Y.; Chen, T.; Jia, A.Q. 1H-NMR based metabolomics approach to study the toxic effects of herbicide butachlor on goldfish (Carassius auratus). Aquat. Toxicol. 2015, 159, 69–80. [Google Scholar] [CrossRef]
- Gao, Y.; Lu, Y.; Huang, S.; Gao, L.; Liang, X.; Wu, Y.; Wang, J.; Huang, Q.; Tang, L.; Wang, G.; et al. Identifying early urinary metabolic changes with long-term environmental exposure to cadmium by mass-spectrometry-based metabolomics. Environ. Sci. Technol. 2014, 48, 6409–6418. [Google Scholar] [CrossRef]
- Wang, X.; Qiu, J.; Xu, P.; Zhang, P.; Wang, Y.; Zhou, Z.; Zhu, W. Rapid metabolite discovery, identification, and accurate comparison of the stereoselective metabolism of metalaxyl in rat hepatic microsomes. J. Agric. Food Chem. 2015, 63, 754–760. [Google Scholar] [CrossRef]
- Wang, X.; Zhu, W.; Qiu, J.; Zhang, P.; Wang, Y.; Zhou, Z. Enantioselective metabolism and toxic effects of metalaxyl on primary hepatocytes from rat. Environ. Sci. Pollut. Res. Int. 2016, 23, 18649–18656. [Google Scholar] [CrossRef]
- Zhang, P.; Zhu, W.; Qiu, J.; Wang, D.; Wang, X.; Wang, Y.; Zhou, Z. Evaluating the enantioselective degradation and novel metabolites following a single oral dose of metalaxyl in mice. Pestic. Biochem. Physiol. 2014, 116, 32–39. [Google Scholar] [CrossRef]
- Zhang, R.; Zhou, Z. Effects of the Chiral Fungicides Metalaxyl and Metalaxyl-M on the Earthworm Eisenia fetida as Determined by (1)H-NMR-Based Untargeted Metabolomics. Molecules 2019, 24, 1293. [Google Scholar] [CrossRef]
- Gu, J.; Ji, C.; Yue, S.; Shu, D.; Su, F.; Zhang, Y.; Xie, Y.; Liu, W.; Zhao, M. Enantioselective Effects of Metalaxyl Enantiomers in Adolescent Rat Metabolic Profiles Using NMR-Based Metabolomics. Environ. Sci. Technol. 2018, 52, 5438–5447. [Google Scholar] [CrossRef] [PubMed]
- An, Y.; Xu, W.; Li, H.; Lei, H.; Zhang, L.; Hao, F.; Duan, Y.; Yan, X.; Zhao, Y.; Wu, J.; et al. High-fat diet induces dynamic metabolic alterations in multiple biological matrices of rats. J. Proteome Res. 2013, 12, 3755–3768. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; An, Y.; Zhang, L.; Lei, H.; Wang, Y.; Tang, H. Combined NMR and GC-MS analyses revealed dynamic metabolic changes associated with the carrageenan-induced rat pleurisy. J. Proteome Res. 2013, 12, 5520–5534. [Google Scholar] [CrossRef] [PubMed]
- Dong, F.; Zhang, L.; Hao, F.; Tang, H.; Wang, Y. Systemic responses of mice to dextran sulfate sodium-induced acute ulcerative colitis using 1H NMR spectroscopy. J. Proteome Res. 2013, 12, 2958–2966. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, Z.; Zhao, Y.; Cheng, S.; Ren, H. Identifying health effects of exposure to trichloroacetamide using transcriptomics and metabonomics in mice (Mus musculus). Environ. Sci. Technol. 2013, 47, 2918–2924. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Wang, H.; Xu, W.; Zeng, Y.; Hou, Y.; Zhang, Y.; Zhao, X.; Sun, C. Application of ultraperformance liquid chromatography/mass spectrometry-based metabonomic techniques to analyze the joint toxic action of long-term low-level exposure to a mixture of organophosphate pesticides on rat urine profile. Toxicol. Sci. 2013, 134, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Wang, B.; Zhang, M.; Rantalainen, M.; Wang, S.; Zhou, H.; Zhang, Y.; Shen, J.; Pang, X.; Wei, H.; et al. Symbiotic gut microbes modulate human metabolic phenotypes. Proc. Natl. Acad. Sci. USA 2008, 105, 2117–2122. [Google Scholar] [CrossRef] [Green Version]
- Dumas, M.E.; Barton, R.H.; Toye, A.; Cloarec, O.; Blancher, C.; Rothwell, A.; Fearnside, J.; Tatoud, R.; Blanc, V.; Lindon, J.C.; et al. Metabolic profiling reveals a contribution of gut microbiota to fatty liver phenotype in insulin-resistant mice. Proc. Natl. Acad. Sci. USA 2006, 103, 12511–12516. [Google Scholar] [CrossRef] [Green Version]
- Novelli, A.; Tasker, R.A. Excitatory amino acids in epilepsy: From the clinics to the laboratory. Amino Acids 2007, 32, 295–297. [Google Scholar] [CrossRef]
- Phang, J.M.; Donald, S.P.; Pandhare, J.; Liu, Y. The metabolism of proline, a stress substrate, modulates carcinogenic pathways. Amino Acids 2008, 35, 681–690. [Google Scholar] [CrossRef] [Green Version]
- Suenaga, R.; Tomonaga, S.; Yamane, H.; Kurauchi, I.; Tsuneyoshi, Y.; Sato, H.; Denbow, D.M.; Furuse, M. Intracerebroventricular injection of L-arginine induces sedative and hypnotic effects under an acute stress in neonatal chicks. Amino Acids 2008, 35, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Wu, G. Amino acids: Metabolism, functions, and nutrition. Amino Acids 2009, 37, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Chubukov, V.; Gerosa, L.; Kochanowski, K.; Sauer, U. Coordination of microbial metabolism. Nat. Rev. Microbiol. 2014, 12, 327–340. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Zhang, Y.; Zhang, R.; Wu, B.; Ding, L.; Xu, K.; Ren, H. Mice in vivo toxicity studies for monohaloacetamides emerging disinfection byproducts based on metabolomic methods. Environ. Sci. Technol. 2014, 48, 8212–8218. [Google Scholar] [CrossRef]
- Harding, H.P.; Zhang, Y.; Zeng, H.; Novoa, I.; Lu, P.D.; Calfon, M.; Sadri, N.; Yun, C.; Popko, B.; Paules, R.; et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol. Cell 2003, 11, 619–633. [Google Scholar] [CrossRef]
- Locasale, J.W. Serine, glycine and one-carbon units: Cancer metabolism in full circle. Nat. Rev. Cancer 2013, 13, 572–583. [Google Scholar] [CrossRef]
- Orlando, G.F.; Wolf, G.; Engelmann, M. Role of neuronal nitric oxide synthase in the regulation of the neuroendocrine stress response in rodents: Insights from mutant mice. Amino Acids 2008, 35, 17–27. [Google Scholar] [CrossRef]
- Willis, A.; Bender, H.U.; Steel, G.; Valle, D. PRODH variants and risk for schizophrenia. Amino Acids 2008, 35, 673–679. [Google Scholar] [CrossRef]
- Galli, F. Amino acid and protein modification by oxygen and nitrogen species. Amino Acids 2012, 42, 1–4. [Google Scholar] [CrossRef]
- Jobgen, W.S.; Fried, S.K.; Fu, W.J.; Meininger, C.J.; Wu, G. Regulatory role for the arginine-nitric oxide pathway in metabolism of energy substrates. J. Nutr. Biochem. 2006, 17, 571–588. [Google Scholar] [CrossRef]
- Marc Rhoads, J.; Wu, G. Glutamine, arginine, and leucine signaling in the intestine. Amino Acids 2009, 37, 111–122. [Google Scholar] [CrossRef] [PubMed]
- Brosnan, J.T. Amino acids, then and now—A reflection on Sir Hans Krebs’ contribution to nitrogen metabolism. Iubmb Life 2001, 52, 265–270. [Google Scholar] [CrossRef] [PubMed]
- Kaspar, H. Amino Acid Analysis in Biological Fluids by GC-MS. Ph.D. Thesis, University of Regensburg, Regensburg, Germany, 2009. [Google Scholar]
- Zhu, W.; Stevens, A.P.; Dettmer, K.; Gottfried, E.; Hoves, S.; Kreutz, M.; Holler, E.; Canelas, A.B.; Kema, I.; Oefner, P.J. Quantitative profiling of tryptophan metabolites in serum, urine, and cell culture supernatants by liquid chromatography-tandem mass spectrometry. Anal. Bioanal. Chem. 2011, 401, 3249–3261. [Google Scholar] [CrossRef] [PubMed]

| No. | Metabolites | HMDB ID | Chemical Shift (ppm) | VIP | Fold Change (MX) | Trend | Fold Change (R-MX) | Trend | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 10 mg/kg | 30 mg/kg | 60 mg/kg | 10 mg/kg | 30 mg/kg | 60 mg/kg | |||||||
| 1 | Fucose | HMDB0000174 | 1.25 (d, CH3); 3.77 (m, CH);3.81 (m, CH); 5.21 (d, CH) | 1.07 | 0.91 | 0.96 | 0.99 | - | 0.94 | 0.76 * | 0.68 * | ↓ |
| 2 | Lactate | HMDB0000190 | 1.33 (d, CH3); 4.11 (q, CH) | 2.81 | 1.22 * | 0.96 | 1.09 | ↑ | 1.30 * | 1.63 ** | 1.19 * | ↑ |
| 3 | Alanine | HMDB0000161 | 1.46 (d, CH3); 3.78 (q, CH) | 3.85 | 1.09 | 1.14 * | 1.22 * | ↑ | 1.20 * | 1.04 | 0.91 | ↑ |
| 4 | Acetate | HMDB0000042 | 1.92 (s, CH3) | 6.54 | 0.86 | 0.85 | 0.96 | - | 0.94 | 0.86 | 0.52 * | ↓ |
| 5 | Pyruvate | HMDB0000243 | 2.37 (s, CH3) | 1.26 | 0.91 | 0.99 | 0.81 * | ↓ | 0.77 * | 0.71 * | 0.69 ** | ↓ |
| 6 | Succinate | HMDB0000254 | 2.41 (s, CH) | 2.28 | 1.19 | 1.09 | 1.07 | - | 1.41 * | 1.34 * | 1.36 * | ↑ |
| 7 | Citrate | HMDB0000094 | 2.54 (d, CH2), 2.69 (d, CH′2) | 1.30 | 1.23 | 1.15 | 1.13 | - | 0.93 | 1.13 | 1.57 * | ↑ |
| 8 | MA | HMDB0000164 | 2.61 (s, CH3) | 1.22 | 0.90 | 0.87 * | 0.79 ** | ↓ | 0.8 ** | 0.81 ** | 0.77 ** | ↓ |
| 9 | DMA | HMDB0000087 | 2.72 (s, CH3) | 1.15 | 1.35 * | 1.02 | 1.07 | ↑ | 1.36 ** | 1.24 ** | 1.46 ** | ↑ |
| 10 | TMA | HMDB0000906 | 2.87 (s, CH3) | 1.74 | 0.96 | 0.92 | 0.84 * | ↓ | 1.00 | 0.73 * | 0.87 * | ↓ |
| 11 | DMG | HMDB0000092 | 2.93 (s, CH3); 3.73 (s, CH2) | 1.43 | 0.91 | 1.03 | 1.10 | - | 0.96 | 0.83 * | 0.68 * | ↓ |
| 12 | Choline | HMDB0000097 | 3.20 (s, CH3); 3.52 (m, N-CH2); 4.07 (m, O-CH2) | 2.45 | 1.12 | 1.05 | 1.12 | - | 1.11 | 1.58 ** | 1.49 ** | ↑ |
| 13 | PC | HMDB0001565 | 3.22 (s, CH3); 3.59 (m, N-CH2);4.17 (m, O-CH2) | 1.40 | 0.98 | 0.88 | 0.93 | - | 0.62 ** | 0.82 | 0.68 ** | ↓ |
| 14 | GPC | HMDB0000086 | 3.23 (s, CH3); 3.68 (m, N-CH2); 3.69 (m, O-CH2); 4.33 (m, P-O-CH2) | 2.26 | 0.85 * | 0.82 * | 0.94 | ↓ | 0.70 | 0.71 * | 0.61 ** | ↓ |
| 15 | TMAO | HMDB0000925 | 3.27 (s, CH3) | 2.99 | 1.30 | 0.93 | 0.81 | - | 1.19 | 1.13 | 1.61 * | ↑ |
| 16 | Scyllo-inositol | HMDB0006088 | 3.35 (s, CH) | 2.17 | 1.15 ** | 1.34 ** | 1.36 ** | ↑ | 1.52 ** | 1.79 ** | 2.00 ** | ↑ |
| 17 | Glycine | HMDB0000123 | 3.56 (s, CH2) | 2.46 | 1.38 * | 1.18 ** | 1.01 | ↑ | 1.39 ** | 1.10 | 1.26 * | ↑ |
| 18 | Hippurate | HMDB0000714 | 3.97 (d, CH2); 7.55 (t, CH); 7.64 (t, CH); 7.84 (d, CH) | 1.72 | 0.84 * | 0.96 | 1.01 | ↓ | 0.86 | 0.85 * | 0.78 * | ↓ |
| 19 | Formate | HMDB0000142 | 8.46 (s, CH) | 2.94 | 1.08 | 1.13 | 1.01 | - | 1.44 ** | 1.16 | 1.55 ** | ↑ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, P.; Wang, S.; He, Y.; Xu, Y.; Shi, D.; Yang, F.; Yu, W.; Zhu, W.; He, L. Identifying Metabolic Perturbations and Toxic Effects of Rac-Metalaxyl and Metalaxyl-M in Mice Using Integrative NMR and UPLC-MS/MS Based Metabolomics. Int. J. Mol. Sci. 2019, 20, 5457. https://doi.org/10.3390/ijms20215457
Zhang P, Wang S, He Y, Xu Y, Shi D, Yang F, Yu W, Zhu W, He L. Identifying Metabolic Perturbations and Toxic Effects of Rac-Metalaxyl and Metalaxyl-M in Mice Using Integrative NMR and UPLC-MS/MS Based Metabolomics. International Journal of Molecular Sciences. 2019; 20(21):5457. https://doi.org/10.3390/ijms20215457
Chicago/Turabian StyleZhang, Ping, Sheng Wang, Yuhan He, Yangyang Xu, Dongmei Shi, Furong Yang, Weizhong Yu, Wentao Zhu, and Lin He. 2019. "Identifying Metabolic Perturbations and Toxic Effects of Rac-Metalaxyl and Metalaxyl-M in Mice Using Integrative NMR and UPLC-MS/MS Based Metabolomics" International Journal of Molecular Sciences 20, no. 21: 5457. https://doi.org/10.3390/ijms20215457




