Transcriptomic and Co-Expression Network Profiling of Shoot Apical Meristem Reveal Contrasting Response to Nitrogen Rate between Indica and Japonica Rice Subspecies
Abstract
:1. Introduction
2. Results
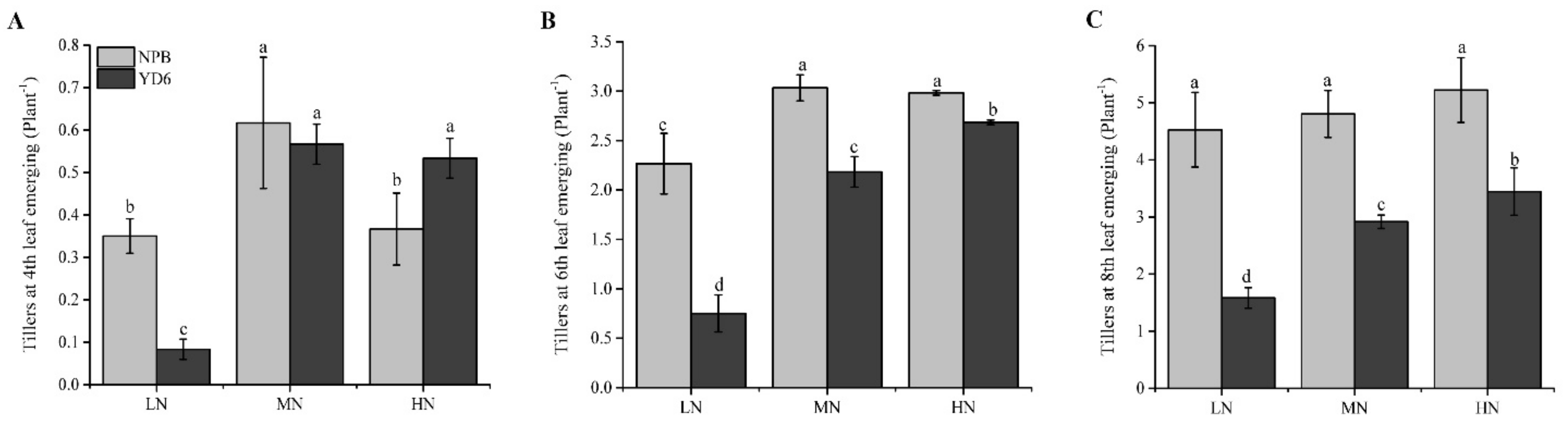
2.1. N Enrichment Promotes Tillers in a Different Pattern between NPB and YD6
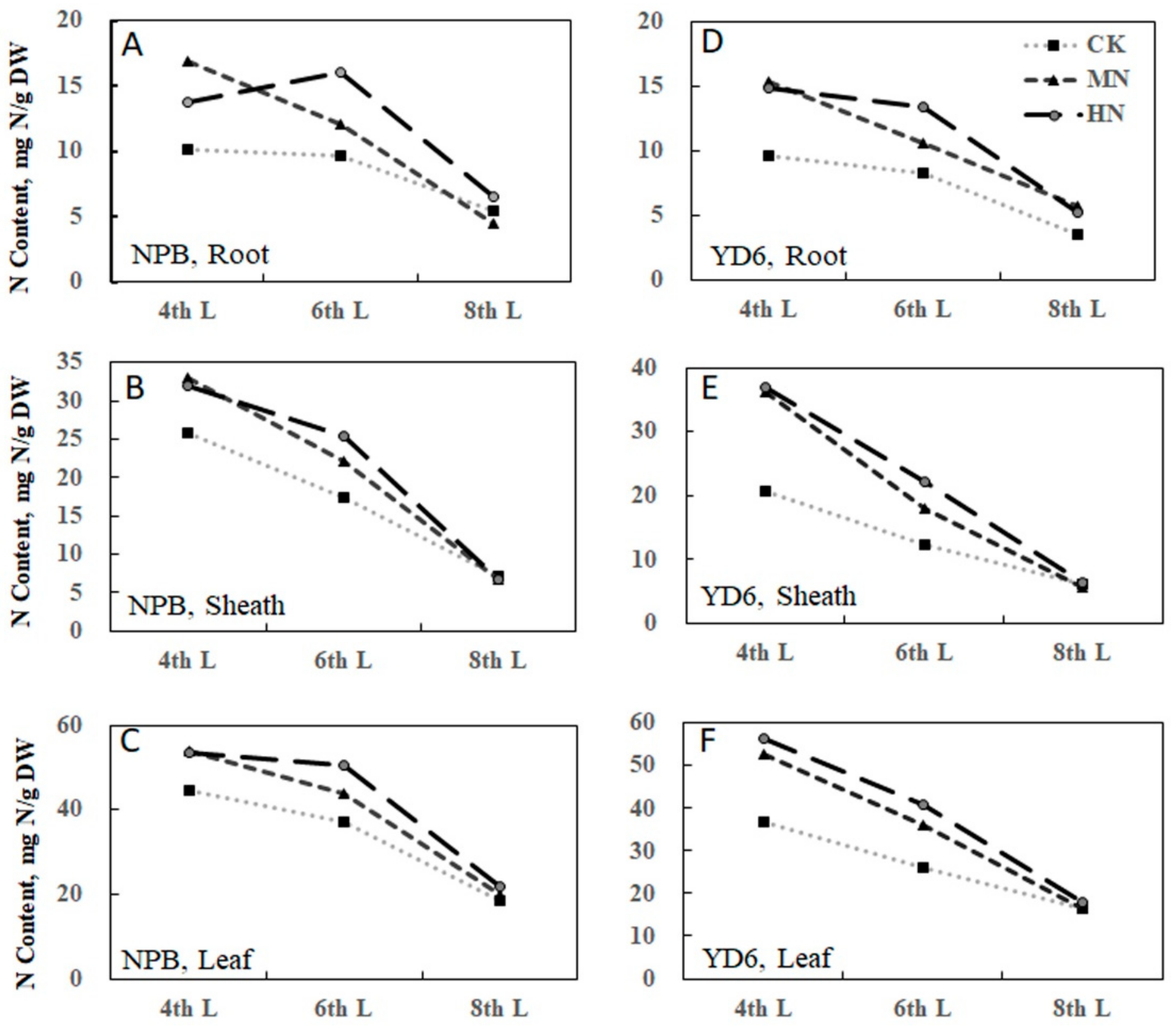
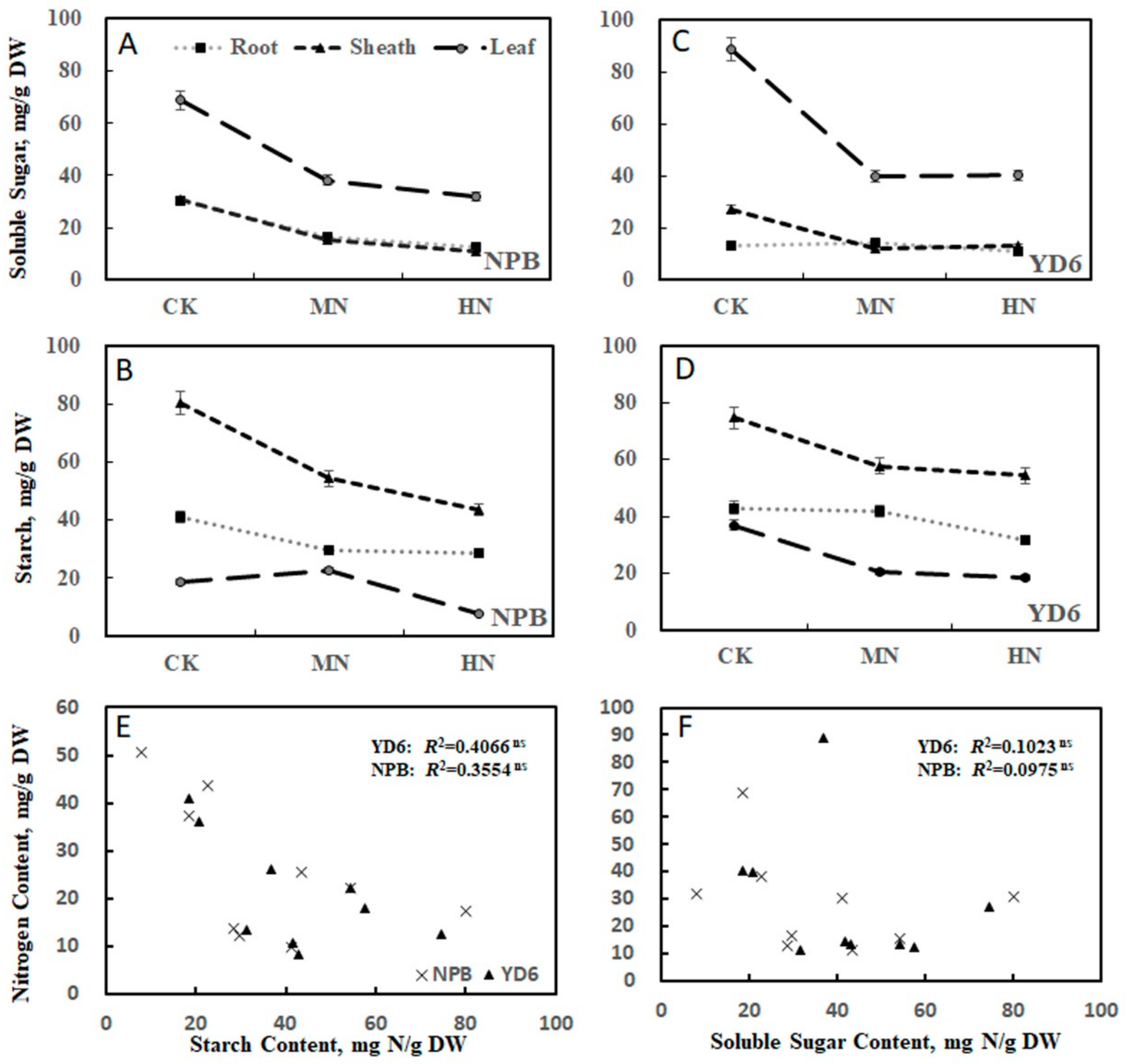
2.2. Tiller Number Is Related with N and Carbohydrate Content
2.3. RNA-Seq Data Quality and Assembly
2.4. Validation of Selected DEG Confirms RNA-Seq Data Reliability
2.5. Differentially Expressed Genes (DEGs) in Response to N Rate
2.6. Gene Ontology (GO) Analysis and KEGG Clustering
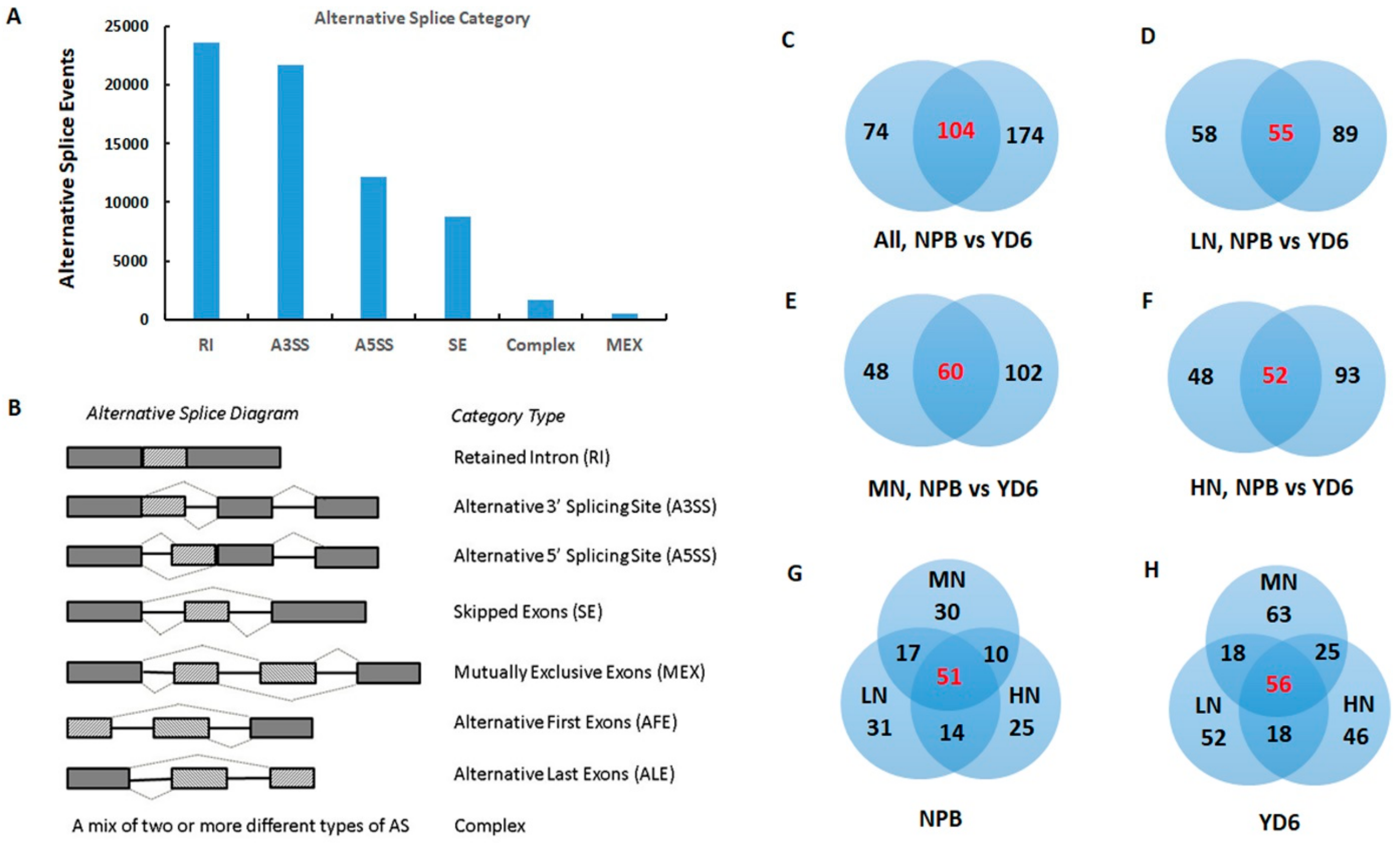
2.7. Alternative Splicing Transcripts and Novel Genes
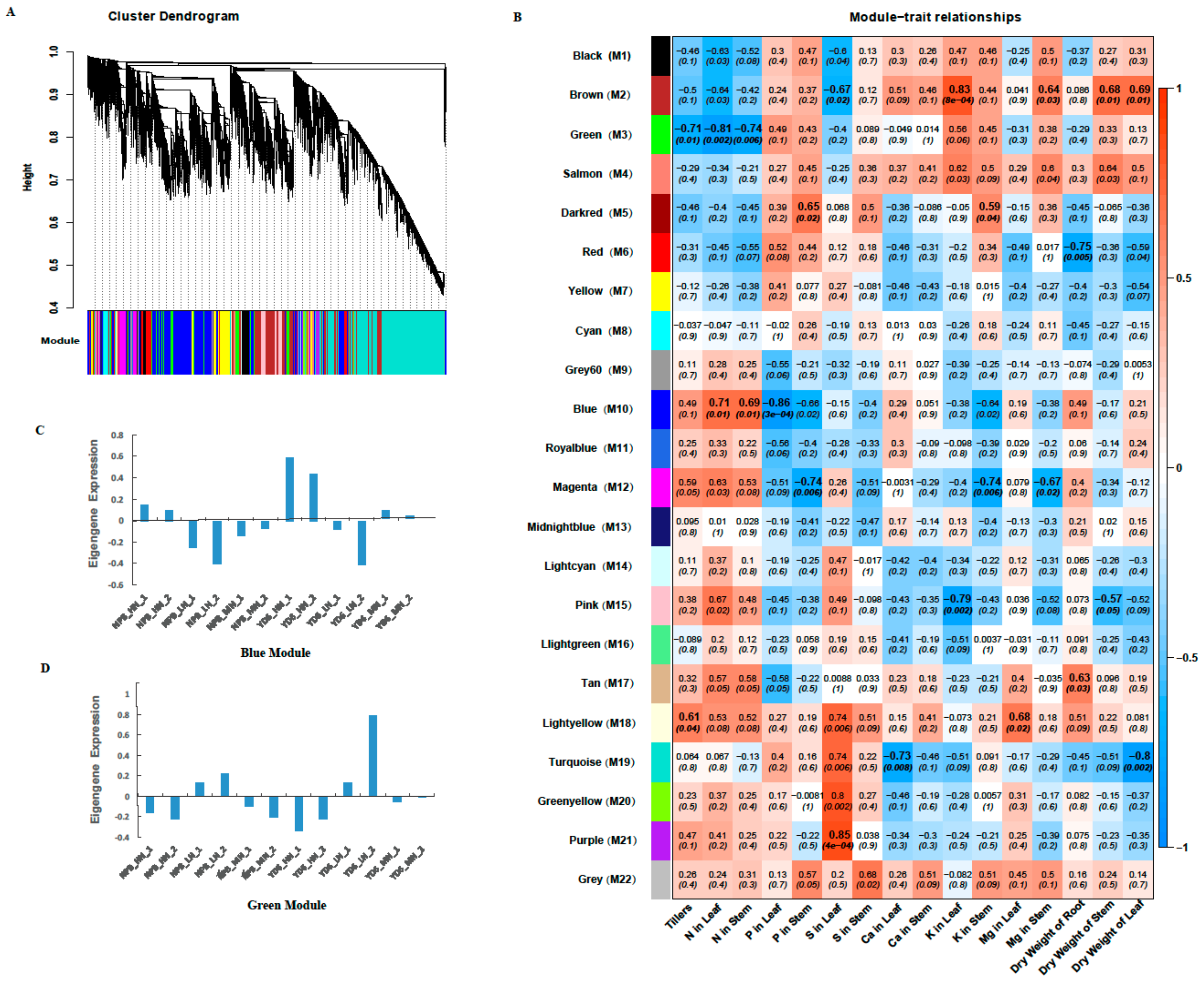
2.8. Identifition of Weighted Gene Co-Expression Network
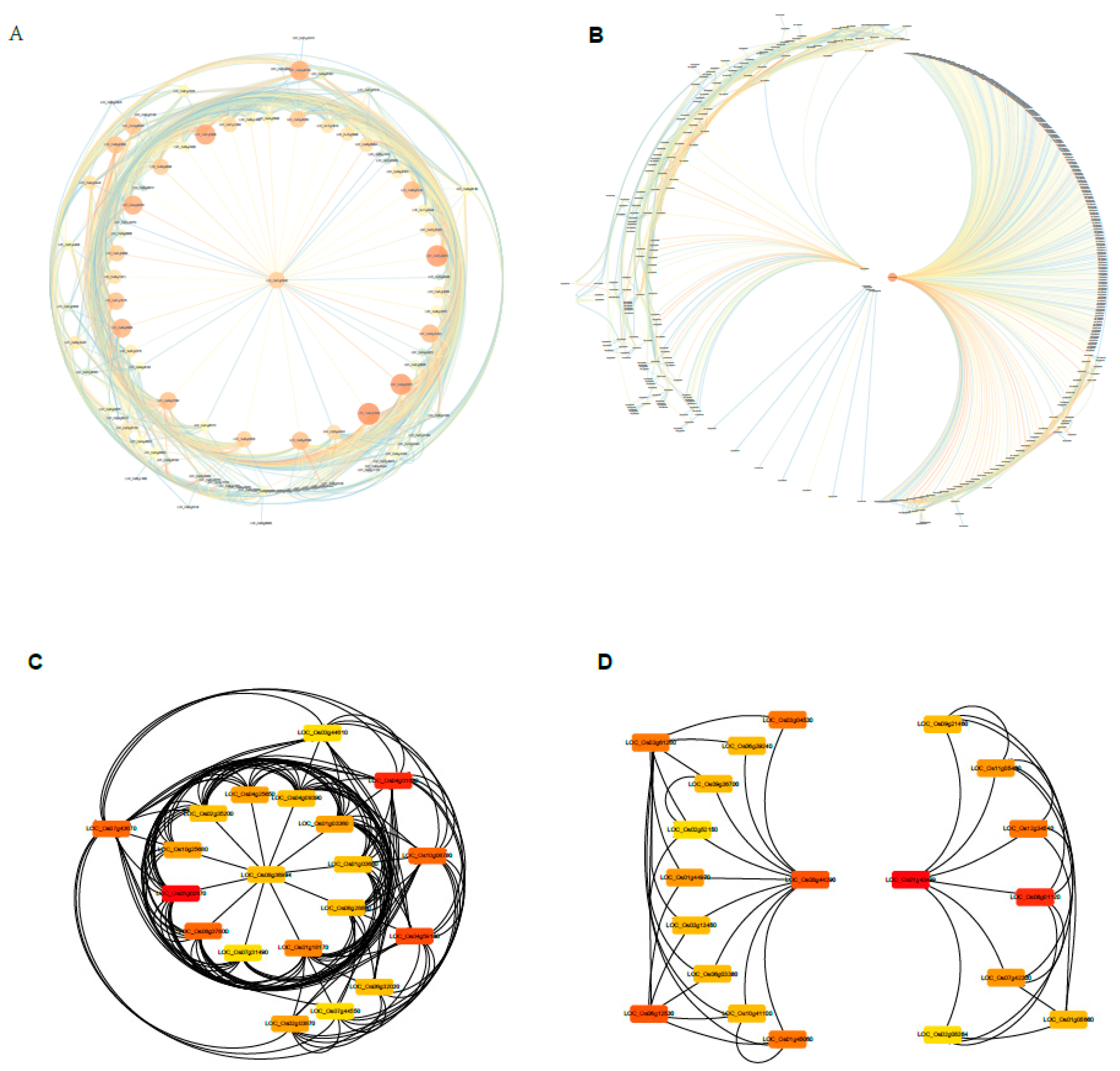
2.9. Construction of the Gene Co-Expression Networks and Identification of Candidate Hub Genes
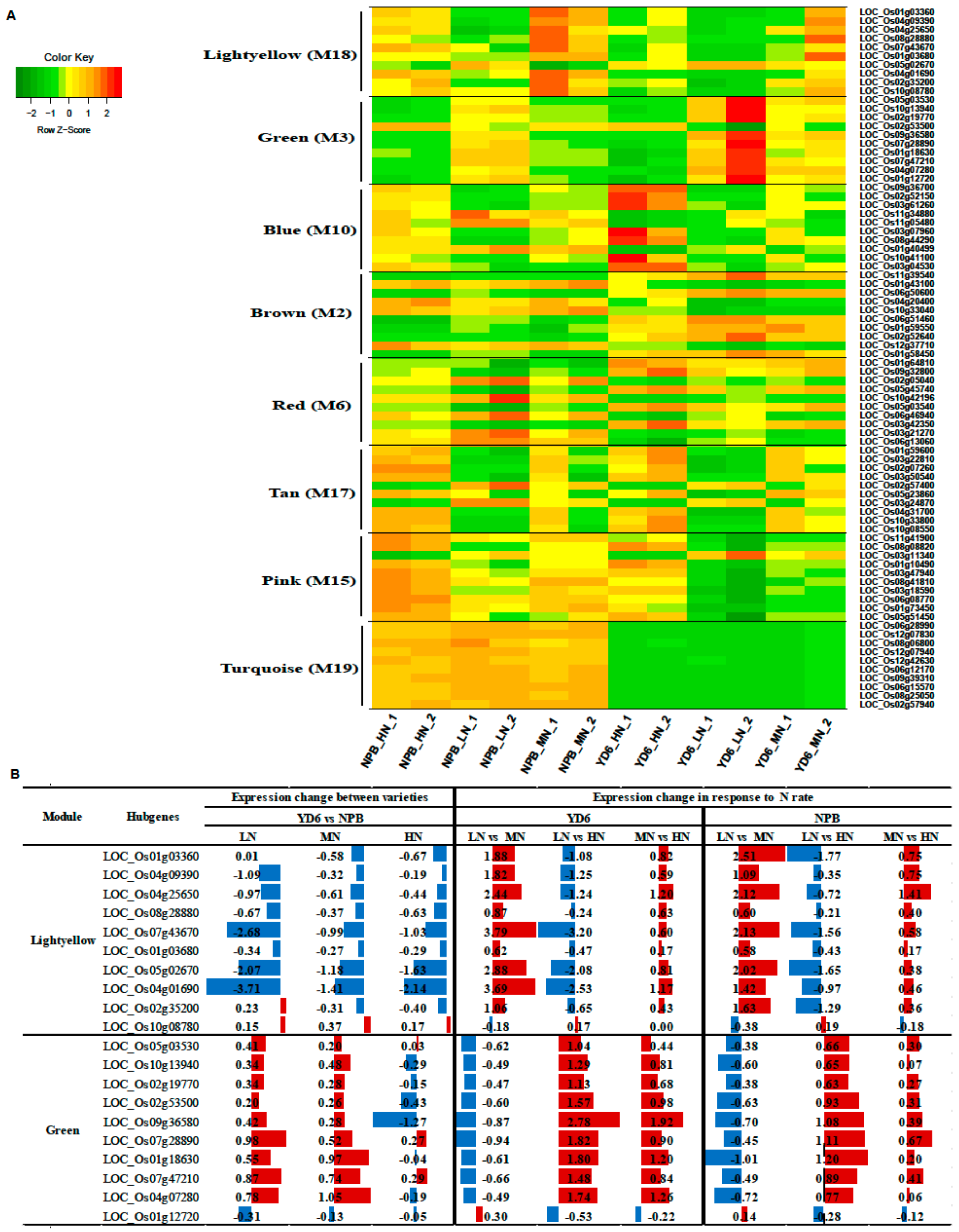
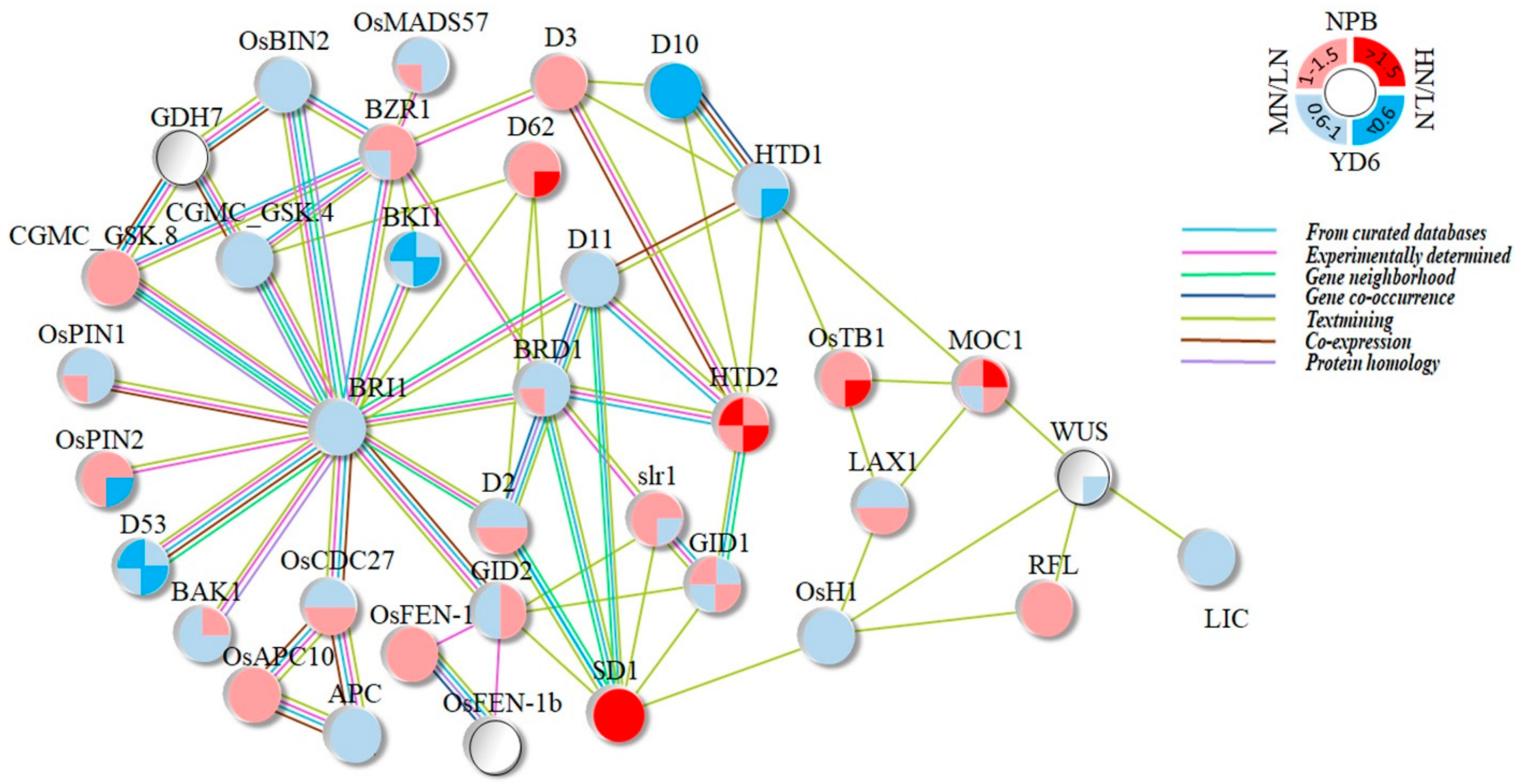
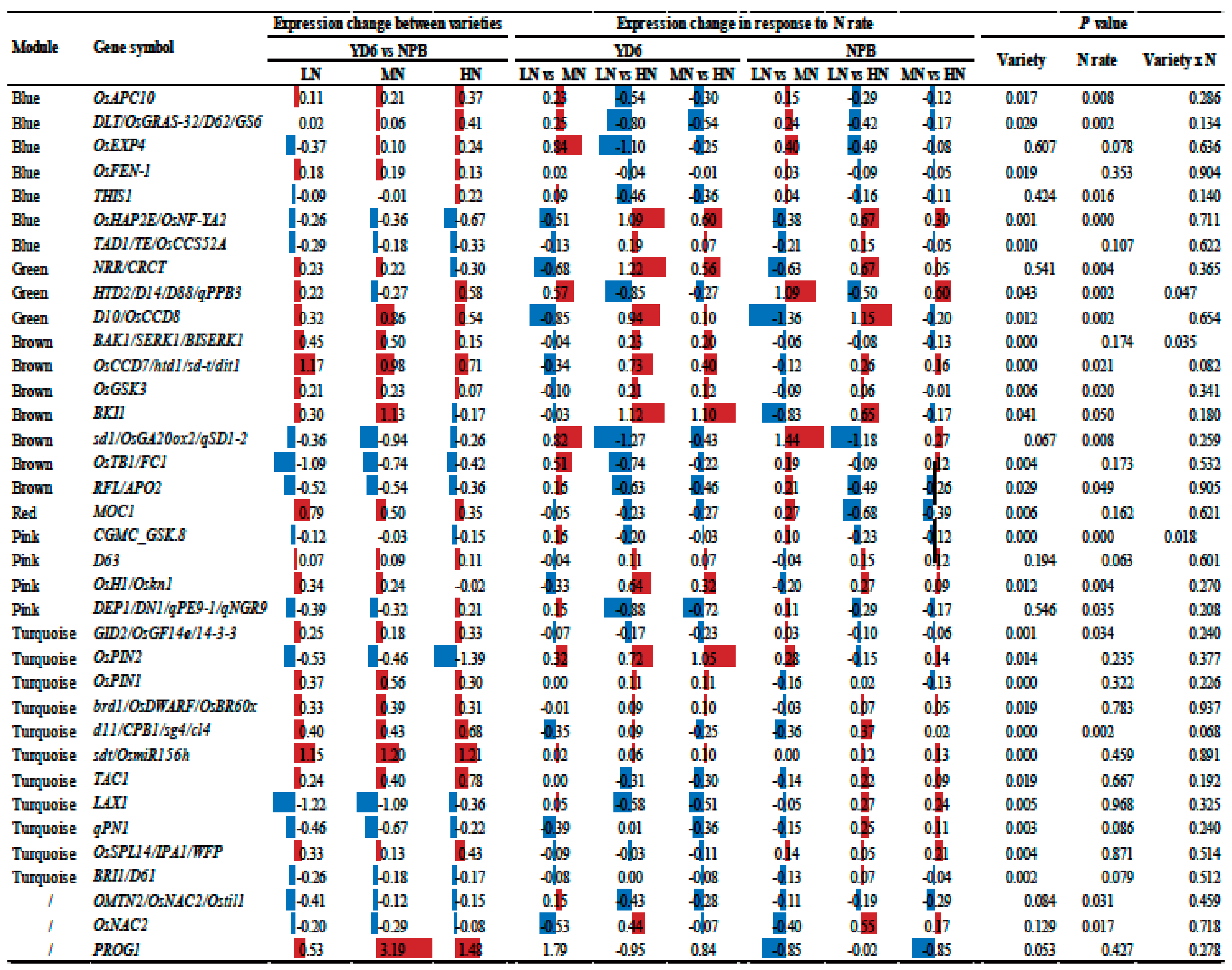
2.10. Expression Profiles of Tiller Related Genes and Their Network
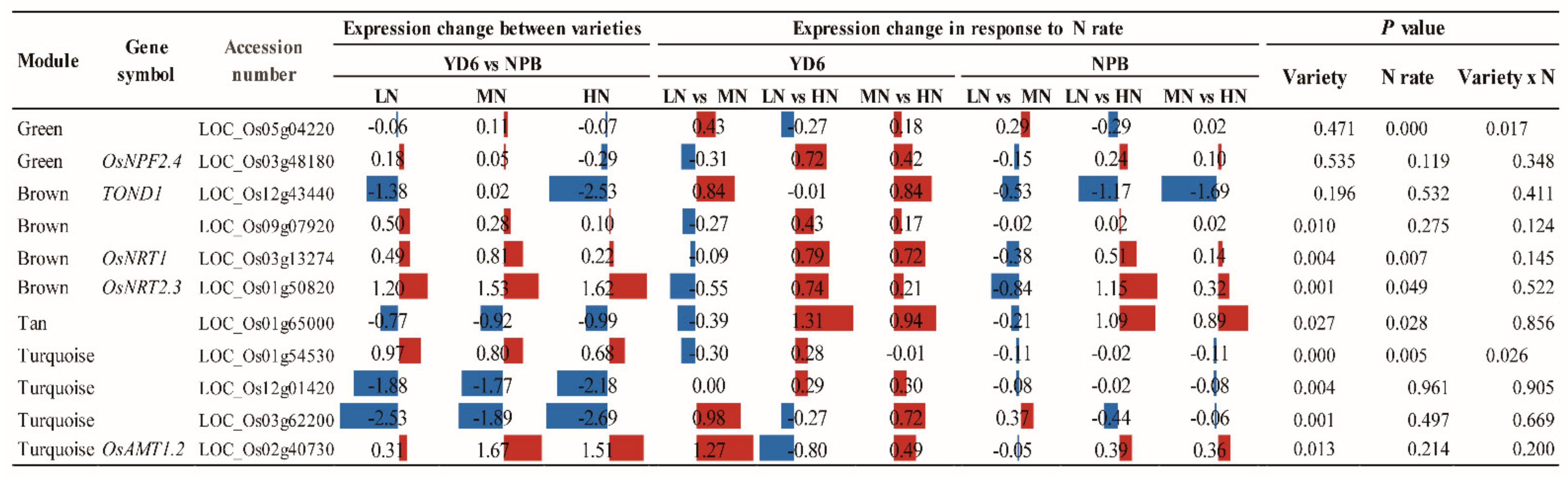
2.11. Expression Profiles of No Apical Meristem Family Genes, Carbohydrate, and N Metabolism and Transport-Related Genes
3. Discussion
3.1. Reducing N Input to Low or Moderate Rate Is Still Good to Promote Enough Tillers
3.2. Most of the Tiller Genes Are Not Drastically Respond to N Rate
4. Materials and Methods
4.1. Plant Materials
4.2. Growth Conditions, N Rate Treatment and Measurement
4.3. Samples Preparation for RNA Extraction, cDNA Library Construction and Sequencing
4.4. RNA-Seq Data Processing and Gene Expression Calculation
4.5. Novel Gene, Alternative Splicing and Enrichment Analysis
4.6. Quantitative Real Time RT-PCR Validation
4.7. Co-Expression Network Analysis for Construction of Modules
4.8. Statistical Analysis of Genes Expression Data in a Specific Pathway
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AS | Alternative splicing |
| DEGs | Differentially expressed genes |
| FPKM | Fragments per kilo-base per million reads |
| GO | Gene ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| N | Nitrogen |
| NAM | No apical meristem |
| qRT-PCR | Quantitative real-time polymerase chain reaction |
| RNA-Seq | RNA-sequencing |
| SAM | Shoot apical meristem |
| WSC | Water soluble carbohydrate |
References
- Ding, Y.; Huang, P.; Lin, Q. Relationship between emergence of tiller and nitrogen concentration of leaf blade or leaf sheath on specific node of rice. J. Nanjing Agric. Univ. 1995, 18, 14–18. [Google Scholar]
- Jin, Z.; Zhang, Y.; Pan, D.; Tong, L.; Li, D.; Li, M.; Wang, H.; Han, Y.; Zhang, Z.; Agriculture, S.O. Correlation analysis of plant carbon-nitrogen content and tiller-related gene expression with rice tiller formation. J. Northeast Agric. Univ. 2017, 48, 10–18. [Google Scholar]
- Mathieu, S.; Bernhard, M.; Bernard, N.; Gilles, P.; André, M. Long-term fate of nitrate fertilizer in agricultural soils. Proc. Natl. Acad. Sci. USA 2013, 110, 18185–18189. [Google Scholar]
- Yang, X.; Nian, J.; Xie, Q.; Feng, J.; Zhang, F.; Jing, H.; Zhang, J.; Dong, G.; Liang, Y.; Peng, J. Rice ferredoxin-dependent glutamate synthase regulates nitrogen-carbon metabolomes and is genetically differentiated between japonica and indica subspecies. Mol. Plant 2016, 9, 1520–1534. [Google Scholar] [CrossRef]
- Chu, G.; Chen, S.; Xu, C.; Wang, D.; Zhang, X. Agronomic and physiological performance of indica/japonica hybrid rice cultivar under low nitrogen conditions. Field Crop. Res. 2019, 243, 107625. [Google Scholar] [CrossRef]
- Pinson, S.R.M.; Wang, Y.; Tabien, R.E. Mapping and validation of quantitative trait loci associated with tiller production in rice. Crop Sci. 2015, 55, 1537. [Google Scholar] [CrossRef]
- Minakuchi, K.; Kameoka, H.; Yasuno, N.; Umehara, M.; Luo, L.; Kobayashi, K.; Hanada, A.; Ueno, K.; Asami, T.; Yamaguchi, S. FINE CULM1 (FC1) works downstream of strigolactones to inhibit the outgrowth of axillary buds in rice. Plant Cell Physiol. 2010, 51, 1127–1135. [Google Scholar] [CrossRef]
- Choi, M.S.; Woo, M.O.; Koh, E.B.; Lee, J.; Ham, T.H.; Seo, H.S.; Koh, H.J. Teosinte Branched 1 modulates tillering in rice plants. Plant Cell Rep. 2012, 31, 57–65. [Google Scholar] [CrossRef]
- Li, X.; Qian, Q.; Fu, Z.; Wang, Y.; Xiong, G.; Zeng, D.; Wang, X.; Liu, X.; Teng, S.; Hiroshi, F.; et al. Control of tillering in rice. Nature 2003, 422, 618–621. [Google Scholar] [CrossRef]
- Lin, Q.; Wang, D.; Dong, H.; Gu, S.; Cheng, Z.; Gong, J.; Qin, R.; Jiang, L.; Li, G.; Wang, J.L. Rice APC/CTE controls tillering by mediating the degradation of MONOCULM 1. Nat. Commun. 2012, 3, 752. [Google Scholar] [CrossRef]
- Xu, C.; Wang, Y.; Yu, Y.; Duan, J.; Liao, Z.; Xiong, G.; Meng, X.; Liu, G.; Qian, Q.; Li, J. Degradation of MONOCULM 1 by APC/CTAD1 regulates rice tillering. Nat. Commun. 2012, 3, 750. [Google Scholar] [CrossRef] [PubMed]
- Keishi, K.; Masahiko, M.; Shin, U.; Yuzuki, S.; Ikuyo, F.; Hironobu, O.; Ko, S.; Junko, K. LAX and SPA: Major regulators of shoot branching in rice. Proc. Natl. Acad. Sci. USA 2003, 100, 11765–11770. [Google Scholar]
- Liang, W.; Shang, F.; Lin, Q.; Lou, C.; Zhang, J. Tillering and panicle branching genes in rice. Gene 2014, 537, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Smith, S.M.; Li, J. Genetic regulation of shoot architecture. Annu. Rev. Plant Biol. 2018, 69, 437–468. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Pan, Y.; Luo, M. Detection and correction of assembly errors of rice Nipponbare reference sequence. Plant Biol. 2014, 16, 643–650. [Google Scholar] [CrossRef] [PubMed]
- Streets, A.M.; Huang, Y. How deep is enough in single-cell RNA-seq? Nat. Biotechnol. 2014, 32, 1005–1006. [Google Scholar] [CrossRef]
- Cole, T.; Williams, B.A.; Geo, P.; Ali, M.; Gordon, K.; Van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Lior, P. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar]
- Sammeth, M.; Foissac, S.; Guigo, R. A general definition and nomenclature for alternative splicing events. PLoS Comput. Biol. 2008, 4, e1000147. [Google Scholar] [CrossRef]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 2008, 9, 559. [Google Scholar] [CrossRef]
- Cheng, X.; Peng, J.; Ma, J.; Tang, Y.; Chen, R.; Mysore, K.S.; Wen, J. NO APICAL MERISTEM (MtNAM) regulates floral organ identity and lateral organ separation in Medicago truncatula. New Phytol. 2012, 195, 71–84. [Google Scholar] [CrossRef]
- Krahmer, J.; Ganpudi, A.; Abbas, A.; Romanowski, A.; Halliday, K.J. Phytochrome, carbon sensing, metabolism, and plant growth plasticity. Plant Physiol. 2017, 176, 1039–1048. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Yao, Q.; Gao, X.; Jiang, C.; Harberd, N.P.; Fu, X. Shoot-to-root mobile transcription factor HY5 coordinates plant carbon and nitrogen acquisition. Curr. Biol. 2016, 26, 640–646. [Google Scholar] [CrossRef]
- Brooks, J.R.; Griffin, V.K.; Kattan, M.W. A modified method for total carbohydrate analysis of glucose syrups, maltodextrins, and other starch hydrolysis products. Cereal. Chem. 1986, 63, 465–466. [Google Scholar]
- Zhou, W.; Yang, Z.; Wang, T.; Fu, Y.; Chen, Y.; Hu, B.; Yamagishi, J.; Ren, W. Environmental compensation effect and synergistic mechanism of optimized nitrogen management increasing nitrogen use efficiency in indica hybrid rice. Front. Plant Sci. 2019, 10, 245. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Zha, M.; Li, Y.; Ding, Y.; Chen, L.; Ding, C.; Wang, S. The interaction between nitrogen availability and auxin, cytokinin, and strigolactone in the control of shoot branching in rice (Oryza sativa L.). Plant Cell Rep. 2015, 34, 1647–1662. [Google Scholar] [CrossRef]
- Lin, C.M.; Koh, S.; Stacey, G.; Yu, S.; Lin, T.; Tsay, T. Cloning and functional characterization of a constitutively expressed nitrate transporter gene, OsNRT1, from rice. Plant Physiol. 2000, 122, 379–388. [Google Scholar] [CrossRef]
- Tang, Z.; Fan, X.; Li, Q.; Feng, H.; Miller, A.; Shen, Q.; Xu, G. Knockdown of a rice stelar nitrate transporter alters long-distance translocation but not root influx. Plant Physiol. 2012, 160, 2052–2063. [Google Scholar] [CrossRef]
- Fan, X.; Tang, Z.; Tan, Y.; Zhang, Y.; Luo, B.; Yang, M.; Lian, X.; Shen, Q.; Miller, A.J.; Xu, G. Overexpression of a pH-sensitive nitrate transporter in rice increases crop yields. Proc. Natl. Acad. Sci. USA 2016, 113, 7118–7123. [Google Scholar] [CrossRef]
- Ishikawa, S.; Ishimaru, Y.; Igura, M.; Kuramata, M.; Abe, T.; Senoura, T.; Hase, Y.; Arao, T.; Nishizawa, N.; Nakanishi, H. Ion-beam irradiation, gene identification, and marker-assisted breeding in the development of low-cadmium rice. Proc. Natl. Acad. Sci. USA 2018, 115, E4950–E4951. [Google Scholar] [CrossRef]
- He, F.; Karve, A.; Maslov, S.; Babst, B. Large-scale public transcriptomic data mining reveals a tight connection between the transport of nitrogen and other transport processes in Arabidopsis. Front. Plant Sci. 2016, 7, 1207. [Google Scholar] [CrossRef] [PubMed]
- Goel, P.; Sharma, N.K.; Bhuria, M.; Sharma, V.; Chauhan, R.; Pathania, S.; Swarnkar, M.K.; Chawla, V.; Acharya, V.; Shankar, R. Transcriptome and co-expression network analyses identify key genes regulating nitrogen use efficiency in Brassica juncea L. Sci. Rep. 2018, 8, 7451. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Chao, W.; Fu, Y.; Hu, G.; Si, H.; Li, Z.; Luan, W.; He, Z.; Sun, Z. Identification and characterization of HTD2: a novel gene negatively regulating tiller bud outgrowth in rice. Planta 2009, 230, 649–658. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Qian, Q.; Liu, X.; Yan, M.; Feng, Q.; Dong, G.; Liu, J.; Han, B. Dwarf 88, a novel putative esterase gene affecting architecture of rice plant. Plant Mol. Biol. 2009, 71, 265–276. [Google Scholar] [CrossRef] [PubMed]
- Arite, T.; Iwata, H.; Ohshima, K.; Maekawa, M.; Nakajima, M.M.; Sakakibara, H.; Kyozuka, J. DWARF10, an RMS1/MAX4/DAD1 ortholog, controls lateral bud outgrowth in rice. Plant J. 2010, 51, 1019–1029. [Google Scholar] [CrossRef]
- Zhang, S.; Li, G.; Fang, J.; Chen, W.; Jiang, H.; Zou, J.; Liu, X.; Zhao, X.; Li, X.; Chu, C. The interactions among DWARF10, auxin and cytokinin underlie lateral bud outgrowth in rice. J. Integr. Plant Biol. 2010, 52, 626–638. [Google Scholar] [CrossRef]
- Zhang, Y.; Yan, Y.; Wang, L.; Yang, K.; Xiao, N.; Liu, Y.; Fu, Y.; Sun, Z.; Fang, R.; Chen, X. A novel rice gene, NRR responds to macronutrient deficiency and regulates root growth. Mol. Plant 2012, 5, 63–72. [Google Scholar] [CrossRef] [Green Version]
- Morita, R.; Inoue, K.; Ikeda, K.I.; Hatanaka, T.; Misoo, S.; Fukayama, H. Starch content in leaf sheath controlled by CO2 responsive CCT protein is a potential determinant of photosynthetic capacity in rice. Plant Cell Physiol. 2016, 57, 2334–2341. [Google Scholar] [CrossRef]
- Nishida, S.; Kakei, Y.; Shimada, Y.; Fujiwara, T. Genome-wide analysis of specific alterations in transcript structure and accumulation caused by nutrient deficiencies in Arabidopsis thaliana. Plant J. 2017, 91, 741–753. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Zhang, H.; Gong, W. Genome-wide identification and comparative analysis of alternative splicing across four legume species. Planta. 2019, 249, 1133–1142. [Google Scholar] [CrossRef]
- Li, W.; Lin, W.; Ray, P.; Lan, P.; Schmidt, W. Genome-wide detection of condition-sensitive alternative splicing in Arabidopsis roots. Plant Physiol. 2013, 162, 1750–1763. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Calixto, C.; Guo, W.; James, A.; Tzioutziou, N.; Entizne, J.; Panter, P.; Knight, H.; Nimmo, H.; Zhang, R.; Brown, J. Rapid and dynamic alternative splicing impacts the Arabidopsis cold response transcriptome. Plant Cell. 2018, 30, 1424–1444. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, G.; Li, W.; Zhang, F.; Guo, W. RNA-seq analysis reveals alternative splicing under salt stress in cotton, Gossypium davidsonii. BMC Genom. 2018, 19, 73. [Google Scholar] [CrossRef] [PubMed]
- Clegg, K.M. The application of the anthrone reagent to the estimation of starch in cereals. J. Sci. Food Agric. 1956, 7, 40–44. [Google Scholar] [CrossRef]
- Trapnell, C.; Pachter LSalzberg, S.L. TopHat: Discovering splice junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef]
- Mortazavi, A.; Williams, B.A.; Mccue, K.; Schaeffer, L.; Wold, B.J. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 2008, 5, 621–628. [Google Scholar] [CrossRef]
- Sylvain, F.; Michael, S. ASTALAVISTA: Dynamic and flexible analysis of alternative splicing events in custom gene datasets. Nucleic Acids Res. 2007, 35, W297–W299. [Google Scholar]
- Michael, S. Complete alternative splicing events are bubbles in splicing graphs. J. Comput. Biol. A J. Comput. Mol. Cell Biol. 2009, 16, 1117–1140. [Google Scholar]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P. The STRING database in 2017: Quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Zhang, X.; Huang, N.; Mo, L.; Lv, M.; Gao, Y.; Wang, J.; Liu, C.; Yin, S.; Zhou, J.; Xiao, N.; et al. Global transcriptome and co-expression network analysis reveal contrasting response of japonica and indica rice cultivar to γ radiation. Int. J. Mol. Sci. 2019, 20, 4358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wisniewski, N.; Cadeiras, M.; Bondar, G.; Cheng, R.K.; Shahzad, K.; Onat, D.; Latif, F.; Korin, Y.; Reed, E.; Fakhro, R. Weighted gene coexpression network analysis (WGCNA) modeling of multiorgan dysfunction syndrome after mechanical circulatory support therapy. J. Heart Lung Transpl. 2013, 32, S223. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]


| Gene Name | Description | KME Value |
|---|---|---|
| Light yellow module (M18) with positive correlation associated with the tillers | ||
| LOC_Os04g09390 | HEV3 - Hevein family protein precursor, expressed | 0.97 |
| LOC_Os01g03680 | BBTI8 - Bowman-Birk type bran trypsin inhibitor precursor, expressed | 0.95 |
| LOC_Os08g28880 | patatin, putative, expressed | 0.94 |
| LOC_Os10g08780 | expressed protein | 0.94 |
| LOC_Os01g03360 | BBTI5 - Bowman-Birk type bran trypsin inhibitor precursor, expressed | 0.94 |
| LOC_Os02g35200 | VP15, putative, expressed | 0.88 |
| LOC_Os07g43670 | ribonuclease T2 family domain containing protein, expressed | 0.85 |
| LOC_Os04g01690 | pyridoxal-dependent decarboxylase protein, putative, expressed | 0.85 |
| LOC_Os04g25650 | cysteine-rich receptor-like protein kinase, putative, expressed | 0.85 |
| LOC_Os05g02670 | kinesin motor domain containing protein, putative, expressed | −0.80 |
| Green module (M3) with negative correlation associated with the tillers, N rate in leaf and stem | ||
| LOC_Os10g13940 | MATE efflux protein, putative, expressed | 0.98 |
| LOC_Os07g28890 | ethylene-responsive protein related, putative, expressed | 0.98 |
| LOC_Os01g12720 | protein kinase domain containing protein, expressed | 0.98 |
| LOC_Os07g47210 | GDSL-like lipase/acylhydrolase, putative, expressed | 0.97 |
| LOC_Os01g18630 | aspartic proteinase oryzasin-1 precursor, putative, expressed | 0.95 |
| LOC_Os02g19770 | eukaryotic translation initiation factor 1A, putative, expressed | 0.94 |
| LOC_Os09g36580 | thaumatin, putative, expressed | 0.93 |
| LOC_Os05g03530 | tetraspanin family protein, putative, expressed | 0.90 |
| LOC_Os04g07280 | AGAP002737-PA, putative, expressed | 0.90 |
| LOC_Os02g53500 | RFC5 - Putative clamp loader of PCNA, replication factor C subunit 5, expressed | −0.92 |
| Blue module (M10) with positive correlation associated with the N rate in leaf and stem | ||
| LOC_Os08g44290 | RNA recognition motif containing protein, putative, expressed | 0.98 |
| LOC_Os03g61260 | ribosomal L18p/L5e family protein, putative, expressed | 0.98 |
| LOC_Os03g04530 | cytochrome P450, putative, expressed | 0.95 |
| LOC_Os09g36700 | ribonuclease T2 family domain containing protein, expressed | 0.94 |
| LOC_Os02g52150 | heat shock 22 kDa protein, mitochondrial precursor, putative, expressed | 0.91 |
| LOC_Os03g07960 | expressed protein | 0.88 |
| LOC_Os10g41100 | CCT motif family protein, expressed | 0.84 |
| LOC_Os11g34880 | NB-ARC domain containing protein, expressed | −0.70 |
| LOC_Os11g05480 | transcription factor, putative, expressed | −0.76 |
| LOC_Os01g40499 | S-locus lectin protein kinase family protein, putative, expressed | −0.82 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, X.; Zhou, J.; Huang, N.; Mo, L.; Lv, M.; Gao, Y.; Chen, C.; Yin, S.; Ju, J.; Dong, G.; et al. Transcriptomic and Co-Expression Network Profiling of Shoot Apical Meristem Reveal Contrasting Response to Nitrogen Rate between Indica and Japonica Rice Subspecies. Int. J. Mol. Sci. 2019, 20, 5922. https://doi.org/10.3390/ijms20235922
Zhang X, Zhou J, Huang N, Mo L, Lv M, Gao Y, Chen C, Yin S, Ju J, Dong G, et al. Transcriptomic and Co-Expression Network Profiling of Shoot Apical Meristem Reveal Contrasting Response to Nitrogen Rate between Indica and Japonica Rice Subspecies. International Journal of Molecular Sciences. 2019; 20(23):5922. https://doi.org/10.3390/ijms20235922
Chicago/Turabian StyleZhang, Xiaoxiang, Juan Zhou, Niansheng Huang, Lanjing Mo, Minjia Lv, Yingbo Gao, Chen Chen, Shuangyi Yin, Jing Ju, Guichun Dong, and et al. 2019. "Transcriptomic and Co-Expression Network Profiling of Shoot Apical Meristem Reveal Contrasting Response to Nitrogen Rate between Indica and Japonica Rice Subspecies" International Journal of Molecular Sciences 20, no. 23: 5922. https://doi.org/10.3390/ijms20235922




