Modeling of Disordered Protein Structures Using Monte Carlo Simulations and Knowledge-Based Statistical Force Fields
Abstract
:1. Introduction
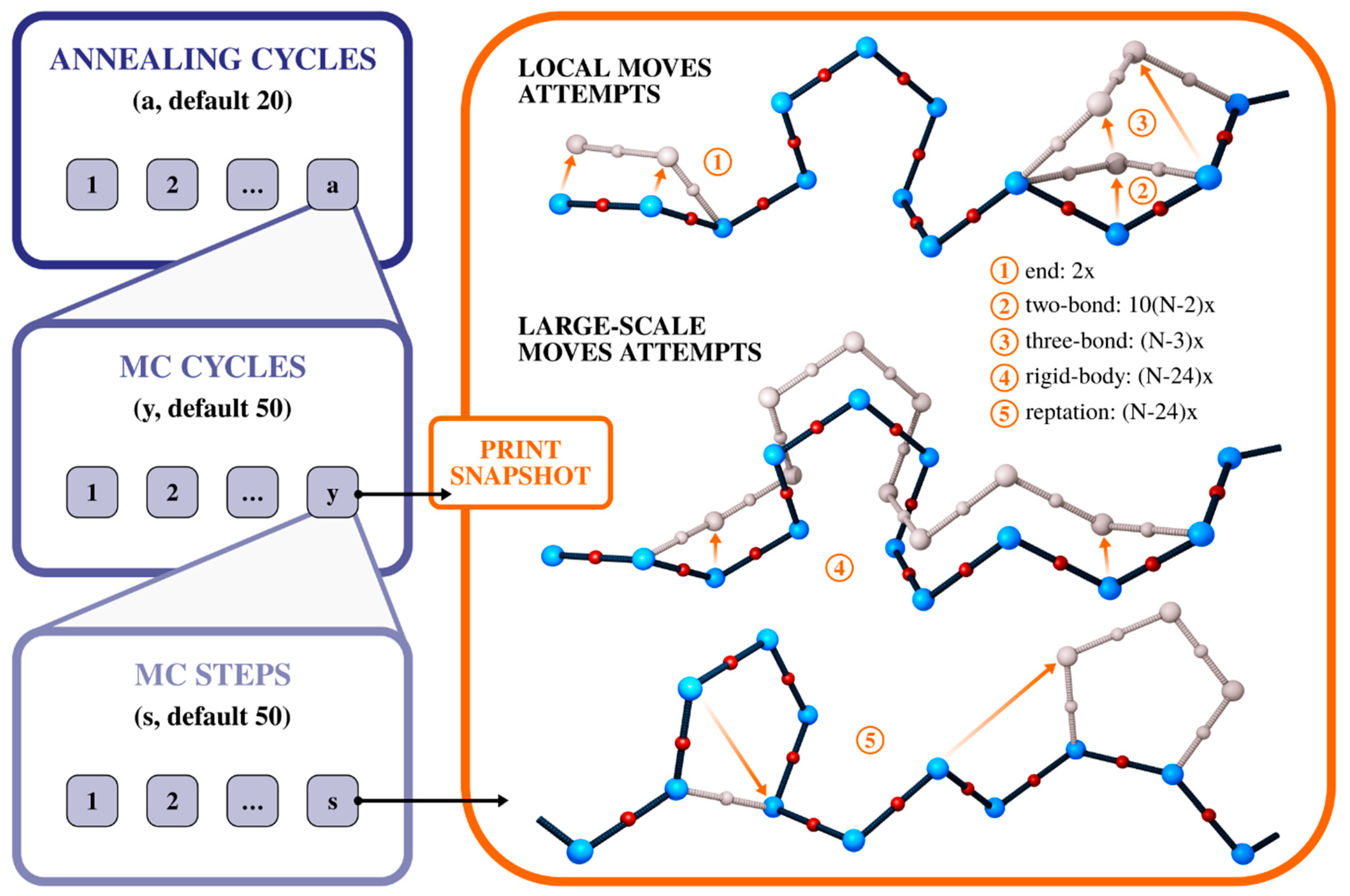
2. CABS Dynamics and Interaction Model
3. CABS Applications to Simulation of Disordered or Unfolded Proteins
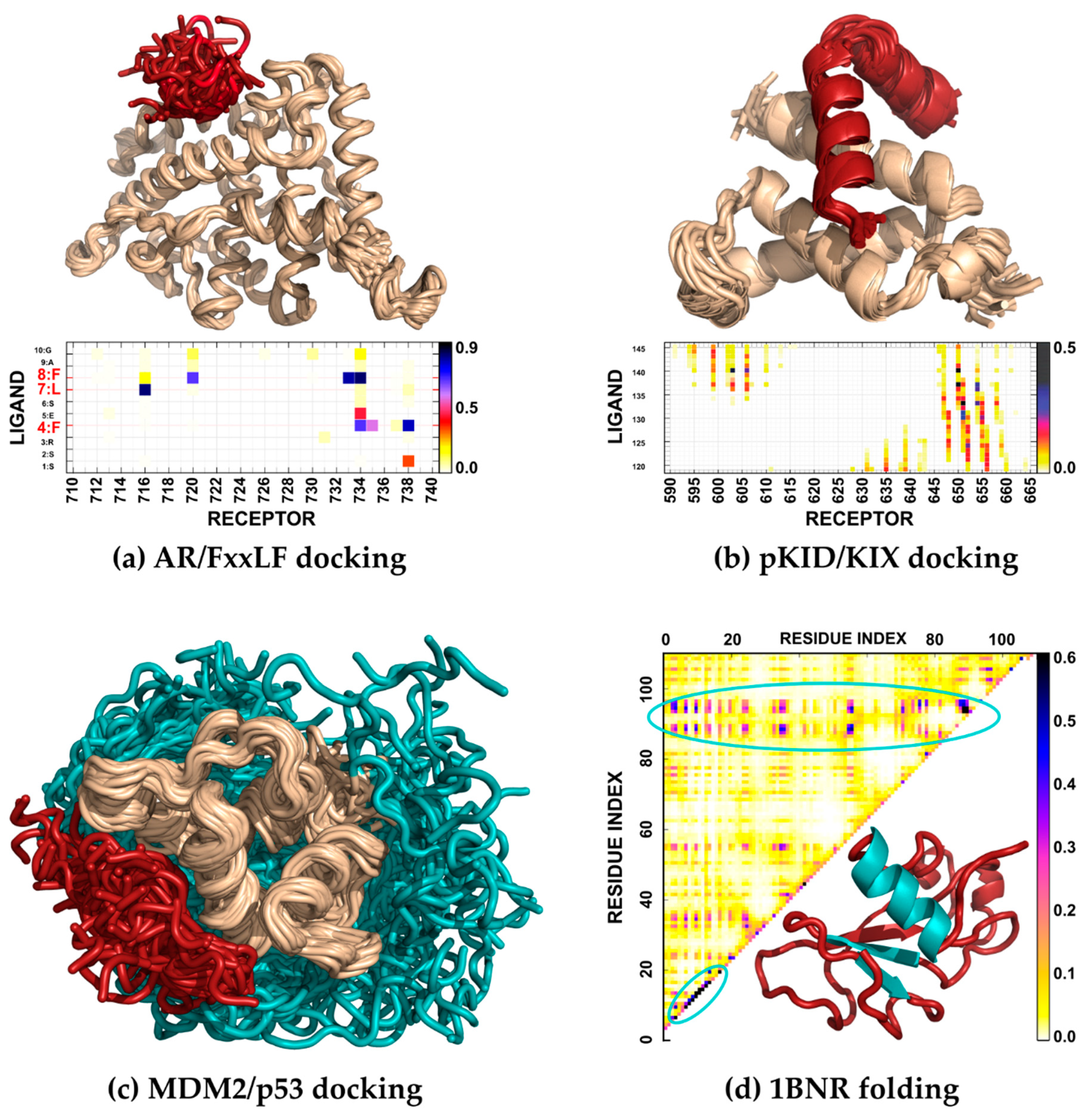
3.1. Protein–Peptide Binding
3.2. Folding and Flexibility of Globular Proteins
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| CABS | Cα, Cβ, Side chain model |
| MC | Monte Carlo |
| NMR | nuclear magnetic resonance |
| MD | molecular dynamics |
| CG | coarse-grained |
| AR | androgen receptor |
| DSSP | dictionary of protein secondary structure |
| RMSD | root-mean-square deviation of atomic positions |
| PDB | Protein Data Bank |
| CASP | Critical Assessment of protein Structure Prediction |
References
- Dishman, A.F.; Volkman, B.F. Unfolding the Mysteries of Protein Metamorphosis. ACS Chem. Biol. 2018, 13, 1438–1446. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Dancing protein clouds: The strange biology and chaotic physics of intrinsically disordered proteins. J. Biol. Chem. 2016, 291, 6681–6688. [Google Scholar] [CrossRef] [PubMed]
- Wright, P.E.; Dyson, H.J. Intrinsically disordered proteins in cellular signalling and regulation. Nat. Rev. Mol. Cell Biol. 2015, 16, 18–29. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Henzler-Wildman, K.; Kern, D. Dynamic personalities of proteins. Nature 2007, 450, 964–972. [Google Scholar] [CrossRef] [PubMed]
- Vendruscolo, M.; Dobson, C.M. Dynamic visions of enzymatic reactions. Science 2006, 313, 1586–1587. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Xi, W.; Nussinov, R.; Ma, B. Protein Ensembles: How Does Nature Harness Thermodynamic Fluctuations for Life? the Diverse Functional Roles of Conformational Ensembles in the Cell. Chem. Rev. 2016, 116, 6516–6551. [Google Scholar] [CrossRef]
- Best, R.B. Computational and theoretical advances in studies of intrinsically disordered proteins. Curr. Opin. Struct. Biol. 2017, 42, 147–154. [Google Scholar] [CrossRef]
- Kay, L.E. NMR studies of protein structure and dynamics. J. Magn. Reson. 2011, 213, 477–491. [Google Scholar] [CrossRef]
- Robustelli, P.; Piana, S.; Shaw, D.E. Developing a molecular dynamics force field for both folded and disordered protein states. Proc. Natl. Acad. Sci. USA 2018, 115, E4758–E4766. [Google Scholar] [CrossRef]
- Bowman, G.R.; Voelz, V.A.; Pande, V.S. Taming the complexity of protein folding. Curr. Opin. Struct. Biol. 2011, 21, 4–11. [Google Scholar] [CrossRef] [Green Version]
- Kmiecik, S.; Gront, D.; Kolinski, M.; Wieteska, L.; Dawid, A.E.; Kolinski, A. Coarse-Grained Protein Models and Their Applications. Chem. Rev. 2016, 116, 7898–7936. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Chan, H.S. Competition between native topology and nonnative interactions in simple and complex folding kinetics of natural and designed proteins. Proc. Natl. Acad. Sci. USA 2010, 107, 2920–2925. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shan, B.; Eliezer, D.; Raleigh, D. The unfolded state of the C-terminal domain of the ribosomal protein L9 contains both native and non-native structure. Biochemistry 2009, 48, 4707–4719. [Google Scholar] [CrossRef] [PubMed]
- Rothwarf, D.M.; Scheraga, H.A. Role of non-native aromatic and hydrophobic interactions in the folding of hen egg white lysozyme. Biochemistry 1996, 35, 13797–13807. [Google Scholar] [CrossRef] [PubMed]
- Cavalli, A.; Montalvao, R.W.; Vendruscolo, M. Using chemical shifts to determine structural changes in proteins upon complex formation. J. Phys. Chem. B 2011, 115, 9491–9494. [Google Scholar] [CrossRef] [PubMed]
- Fu, B.; Kukic, P.; Camilloni, C.; Vendruscolo, M. MD Simulations of Intrinsically Disordered Proteins with Replica-Averaged Chemical Shift Restraints. Biophys. J. 2014, 106, 481a. [Google Scholar] [CrossRef] [Green Version]
- Kar, P.; Feig, M. Recent advances in transferable coarse-grained modeling of proteins. Adv. Protein Chem. Struct. Biol. 2014, 96, 143–180. [Google Scholar] [CrossRef]
- Greener, J.G.; Filippis, I.; Sternberg, M.J.E. Predicting Protein Dynamics and Allostery Using Multi-Protein Atomic Distance Constraints. Structure 2017, 25, 546–558. [Google Scholar] [CrossRef]
- Klepeis, J.L.; Lindorff-Larsen, K.; Dror, R.O.; Shaw, D.E. Long-timescale molecular dynamics simulations of protein structure and function. Curr. Opin. Struct. Biol. 2009, 19, 120–127. [Google Scholar] [CrossRef]
- Bernardi, R.C.; Melo, M.C.R.; Schulten, K. Enhanced sampling techniques in molecular dynamics simulations of biological systems. Biochim. Biophys. Acta - Gen. Subj. 2015, 1850, 872–877. [Google Scholar] [CrossRef] [Green Version]
- Shukla, D.; Hernández, C.X.; Weber, J.K.; Pande, V.S. Markov state models provide insights into dynamic modulation of protein function. Acc. Chem. Res. 2015, 48, 414–422. [Google Scholar] [CrossRef] [PubMed]
- Kolinski, A. Toward more efficient simulations of slow processes in large biomolecular systems: Comment on “Ligand diffusion in proteins via enhanced sampling in molecular dynamics” by Jakub Rydzewski and Wieslaw Nowak. Phys. Life Rev. 2017, 22–23, 75–76. [Google Scholar] [CrossRef] [PubMed]
- Rydzewski, J.; Nowak, W. Ligand diffusion in proteins via enhanced sampling in molecular dynamics. Phys. Life Rev. 2017, 22–23, 82–84. [Google Scholar] [CrossRef] [PubMed]
- Maximova, T.; Moffatt, R.; Ma, B.; Nussinov, R.; Shehu, A. Principles and Overview of Sampling Methods for Modeling Macromolecular Structure and Dynamics. PLoS Comput. Biol. 2016, 12, e1004619. [Google Scholar] [CrossRef] [PubMed]
- Hatherley, R.; Brown, D.K.; Glenister, M.; Bishop, Ö.T. PRIMO: An interactive homology modeling pipeline. PLoS ONE 2016, 11, e0166698. [Google Scholar] [CrossRef] [PubMed]
- Das, R.; Baker, D. Macromolecular Modeling with Rosetta. Annu. Rev. Biochem. 2008, 77, 363–382. [Google Scholar] [CrossRef] [PubMed]
- Czaplewski, C.; Karczyńska, A.; Sieradzan, A.K.; Liwo, A. UNRES server for physics-based coarse-grained simulations and prediction of protein structure, dynamics and thermodynamics. Nucleic Acids Res. 2018, 46, W304–W309. [Google Scholar] [CrossRef] [PubMed]
- Kolinski, A. Protein modeling and structure prediction with a reduced representation. Acta Biochim. Pol. 2004, 51, 349–371. [Google Scholar]
- Davtyan, A.; Schafer, N.P.; Zheng, W.; Clementi, C.; Wolynes, P.G.; Papoian, G.A. AWSEM-MD: Protein structure prediction using coarse-grained physical potentials and bioinformatically based local structure biasing. J. Phys. Chem. B 2012, 116, 8494–8503. [Google Scholar] [CrossRef]
- Marrink, S.J.; Tieleman, D.P. Perspective on the Martini model. Chem. Soc. Rev. 2013, 42, 6801. [Google Scholar] [CrossRef]
- Pasi, M.; Lavery, R.; Ceres, N. PaLaCe: A coarse-grain protein model for studying mechanical properties. J. Chem. Theory Comput. 2013, 9, 785–793. [Google Scholar] [CrossRef] [PubMed]
- Basdevant, N.; Borgis, D.; Ha-Duong, T. Modeling protein-protein recognition in solution using the coarse-grained force field SCORPION. J. Chem. Theory Comput. 2013, 9, 803–813. [Google Scholar] [CrossRef] [PubMed]
- Dawid, A.E.; Gront, D.; Kolinski, A. SURPASS Low-Resolution Coarse-Grained Protein Modeling. J. Chem. Theory Comput. 2017, 13, 5766–5779. [Google Scholar] [CrossRef] [PubMed]
- Dawid, A.E.; Gront, D.; Kolinski, A. Coarse-Grained Modeling of the Interplay between Secondary Structure Propensities and Protein Fold Assembly. J. Chem. Theory Comput. 2018, 14, 2277–2287. [Google Scholar] [CrossRef] [PubMed]
- Lopes, P.E.M.; Guvench, O.; MacKerell, A.D. Current Status of Protein Force Fields for Molecular Dynamics Simulations. In Molecular Modeling of Proteins; Humana Press: New York, NY, USA, 2015; pp. 47–71. [Google Scholar]
- Cossio, P.; Granata, D.; Laio, A.; Seno, F.; Trovato, A. A simple and efficient statistical potential for scoring ensembles of protein structures. Sci. Rep. 2012, 2, 351. [Google Scholar] [CrossRef]
- Tanaka, S.; Scheraga, H.A. Medium- and Long-Range Interaction Parameters between Amino Acids for Predicting Three-Dimensional Structures of Proteins. Macromolecules 1976, 9, 945–950. [Google Scholar] [CrossRef] [PubMed]
- Tsai, J.; Bonneau, R.; Morozov, A.V.; Kuhlman, B.; Rohl, C.A.; Baker, D. An improved protein decoy set for testing energy functions for protein structure prediction. Proteins Struct. Funct. Genet. 2003, 53, 76–87. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shen, M.; Sali, A. Statistical potential for assessment and prediction of protein structures. Protein Sci. 2006, 15, 2507–2524. [Google Scholar] [CrossRef] [Green Version]
- Zhou, H.; Skolnick, J. GOAP: A Generalized Orientation-Dependent, All-Atom Statistical Potential for Protein Structure Prediction. Biophys. J. 2011, 101, 2043–2052. [Google Scholar] [CrossRef] [Green Version]
- Xu, D.; Zhang, Y. Ab initio protein structure assembly using continuous structure fragments and optimized knowledge-based force field. Proteins Struct. Funct. Bioinforma. 2012, 80, 1715–1735. [Google Scholar] [CrossRef]
- Woetzel, N.; Karakaş, M.; Staritzbichler, R.; Müller, R.; Weiner, B.E.; Meiler, J. BCL::Score—Knowledge Based Energy Potentials for Ranking Protein Models Represented by Idealized Secondary Structure Elements. PLoS ONE 2012, 7, e49242. [Google Scholar] [CrossRef] [PubMed]
- Ovchinnikov, S.; Park, H.; Kim, D.E.; Liu, Y.; Wang, R.Y.-R.; Baker, D. Structure prediction using sparse simulated NOE restraints with Rosetta in CASP11. Proteins Struct. Funct. Bioinforma. 2016, 84, 181–188. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ovchinnikov, S.; Kim, D.E.; Wang, R.Y.-R.; Liu, Y.; DiMaio, F.; Baker, D. Improved de novo structure prediction in CASP11 by incorporating coevolution information into Rosetta. Proteins Struct. Funct. Bioinforma. 2016, 84, 67–75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hirst, S.J.; Alexander, N.; Mchaourab, H.S.; Meiler, J. RosettaEPR: An integrated tool for protein structure determination from sparse EPR data. J. Struct. Biol. 2011, 173, 506–514. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Zhang, W.; He, B.; Walker, S.E.; Zhang, H.; Govindarajoo, B.; Virtanen, J.; Xue, Z.; Shen, H.B.; Zhang, Y. Template-based protein structure prediction in CASP11 and retrospect of I-TASSER in the last decade. Proteins 2016, 84, 233–246. [Google Scholar] [CrossRef]
- Russel, D.; Lasker, K.; Webb, B.; Velázquez-Muriel, J.; Tjioe, E.; Schneidman-Duhovny, D.; Peterson, B.; Sali, A. Putting the Pieces Together: Integrative Modeling Platform Software for Structure Determination of Macromolecular Assemblies. PLoS Biol. 2012, 10, e1001244. [Google Scholar] [CrossRef]
- Rodrigues, J.P.G.L.M.; Bonvin, A.M.J.J. Integrative computational modeling of protein interactions. FEBS J. 2014, 281, 1988–2003. [Google Scholar] [CrossRef] [Green Version]
- Kmiecik, S.; Kolinski, A. Folding pathway of the B1 domain of protein G explored by multiscale modeling. Biophys. J. 2008, 94, 726–736. [Google Scholar] [CrossRef]
- Kolinski, A. Multiscale approaches to protein modeling: Structure prediction, dynamics, thermodynamics and macromolecular assemblies. In Multiscale Approaches to Protein Modeling: Structure Prediction, Dynamics, Thermodynamics and Macromolecular Assemblies; Kolinski, A., Ed.; Springer: New York, NY, USA, 2011; pp. 1–355. ISBN 9781441968890. [Google Scholar]
- Kmiecik, S.; Kouza, M.; Badaczewska-Dawid, A.E.; Kloczkowski, A.; Kolinski, A. Modeling of Protein Structural Flexibility and Large-Scale Dynamics: Coarse-Grained Simulations and Elastic Network Models. Int. J. Mol. Sci. 2018, 19, 3496. [Google Scholar] [CrossRef]
- Kmiecik, S.; Kolinski, A. Characterization of protein-folding pathways by reduced-space modeling. Proc. Natl. Acad. Sci. USA 2007, 104, 12330–12335. [Google Scholar] [CrossRef]
- Kmiecik, S.; Gront, D.; Kouza, M.; Kolinski, A. From coarse-grained to atomic-level characterization of protein dynamics: Transition state for the folding of B domain of protein A. J. Phys. Chem. B 2012, 116, 7026–7032. [Google Scholar] [CrossRef] [PubMed]
- Kmiecik, S.; Kurcinski, M.; Rutkowska, A.; Gront, D.; Kolinski, A. Denatured proteins and early folding intermediates simulated in a reduced conformational space. Acta Biochim. Pol. 2006, 53, 131–143. [Google Scholar] [CrossRef] [PubMed]
- Kmiecik, S.; Kolinski, A. Simulation of chaperonin effect on protein folding: A shift from nucleation - Condensation to framework mechanism. J. Am. Chem. Soc. 2011, 133, 10283–10289. [Google Scholar] [CrossRef] [PubMed]
- Jamroz, M.; Kolinski, A.; Kmiecik, S. Protocols for efficient simulations of long-time protein dynamics using coarse-grained CABS model. Methods Mol. Biol. 2014, 1137, 235–250. [Google Scholar] [CrossRef] [PubMed]
- Wabik, J.; Kmiecik, S.; Gront, D.; Kouza, M.; Koliński, A. Combining coarse-grained protein models with replica-exchange all-atom molecular dynamics. Int. J. Mol. Sci. 2013, 14, 9893–9905. [Google Scholar] [CrossRef] [PubMed]
- Blaszczyk, M.; Jamroz, M.; Kmiecik, S.; Kolinski, A. CABS-fold: Server for the de novo and consensus-based prediction of protein structure. Nucleic Acids Res. 2013, 41, W406–W411. [Google Scholar] [CrossRef] [PubMed]
- Kmiecik, S.; Jamroz, M.; Kolinski, M. Structure prediction of the second extracellular loop in G-protein-coupled receptors. Biophys. J. 2014, 106, 2408–2416. [Google Scholar] [CrossRef]
- Koliński, A.; Bujnicki, J.M. Generalized protein structure prediction based on combination of fold-recognition with de novo folding and evaluation of models. Proteins Struct. Funct. Genet. 2005, 61, 84–90. [Google Scholar] [CrossRef] [Green Version]
- Jamroz, M.; Kolinski, A. Modeling of loops in proteins: A multi-method approach. BMC Struct. Biol. 2010, 10. [Google Scholar] [CrossRef]
- Kurcinski, M.; Jamroz, M.; Blaszczyk, M.; Kolinski, A.; Kmiecik, S. CABS-dock web server for the flexible docking of peptides to proteins without prior knowledge of the binding site. Nucleic Acids Res. 2015, 43, W419–W424. [Google Scholar] [CrossRef]
- Kurcinski, M.; Kolinski, A.; Kmiecik, S. Mechanism of folding and binding of an intrinsically disordered protein as revealed by ab initio simulations. J. Chem. Theory Comput. 2014, 10, 2224–2231. [Google Scholar] [CrossRef] [PubMed]
- Ciemny, M.P.; Debinski, A.; Paczkowska, M.; Kolinski, A.; Kurcinski, M.; Kmiecik, S. Protein-peptide molecular docking with large-scale conformational changes: The p53-MDM2 interaction. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Blaszczyk, M.; Kurcinski, M.; Kouza, M.; Wieteska, L.; Debinski, A.; Kolinski, A.; Kmiecik, S. Modeling of protein-peptide interactions using the CABS-dock web server for binding site search and flexible docking. Methods 2016, 93, 72–83. [Google Scholar] [CrossRef] [PubMed]
- Ciemny, M.; Kurcinski, M.; Kozak, K.; Kolinski, A.; Kmiecik, S. Highly flexible protein-peptide docking using cabs-dock. Methods Mol. Biol. 2017, 1561, 69–94. [Google Scholar] [CrossRef]
- Blaszczyk, M.; Ciemny, M.P.; Kolinski, A.; Kurcinski, M.; Kmiecik, S. Protein–peptide docking using CABS-dock and contact information. Brief. Bioinform. 2018, bby080. [Google Scholar] [CrossRef]
- Ciemny, M.P.; Kurcinski, M.; Blaszczyk, M.; Kolinski, A.; Kmiecik, S. Modeling EphB4-EphrinB2 protein-protein interaction using flexible docking of a short linear motif. Biomed. Eng. Online 2017, 16, 71. [Google Scholar] [CrossRef]
- Jamroz, M.; Orozco, M.; Kolinski, A.; Kmiecik, S. Consistent view of protein fluctuations from all-atom molecular dynamics and coarse-grained dynamics with knowledge-based force-field. J. Chem. Theory Comput. 2013, 9, 119–125. [Google Scholar] [CrossRef]
- Jamroz, M.; Kolinski, A.; Kmiecik, S. CABS-flex: Server for fast simulation of protein structure fluctuations. Nucleic Acids Res. 2013, 41, W427–W431. [Google Scholar] [CrossRef] [PubMed]
- Jamroz, M.; Kolinski, A.; Kmiecik, S. CABS-flex predictions of protein flexibility compared with NMR ensembles. Bioinformatics 2014, 30, 2150–2154. [Google Scholar] [CrossRef] [PubMed]
- Kurcinski, M.; Oleniecki, T.; Ciemny, P.M.; Kuriata, A.; Kolinski, A.; Kmiecik, S. CABS-flex standalone: A simulation environment for fast modeling of protein flexibility. Bioinformatics 2018, bty685. [Google Scholar] [CrossRef] [PubMed]
- Kuriata, A.; Gierut, A.M.; Oleniecki, T.; Ciemny, M.P.; Kolinski, A.; Kurcinski, M.; Kmiecik, S. CABS-flex 2.0: A web server for fast simulations of flexibility of protein structures. Nucleic Acids Res. 2018, 46, W338–W343. [Google Scholar] [CrossRef] [PubMed]
- Eswar, N.; John, B.; Mirkovic, N.; Fiser, A.; Ilyin, V.A.; Pieper, U.; Stuart, A.C.; Marti-Renom, M.A.; Madhusudhan, M.S.; Yerkovich, B.; Sali, A. Tools for comparative protein structure modeling and analysis. Nucleic Acids Res. 2003, 31, 3375–3380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gront, D.; Kmiecik, S.; Kolinski, A. Backbone building from quadrilaterals: A fast and accurate algorithm for protein backbone reconstruction from alpha carbon coordinates. J. Comput. Chem. 2007, 28, 1593–1597. [Google Scholar] [CrossRef] [PubMed]
- Canutescu, A.A.; Shelenkov, A.A.; Dunbrack, R.L. A graph-theory algorithm for rapid protein side-chain prediction. Protein Sci. 2003, 12, 2001–2014. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gront, D.; Kmiecik, S.; Blaszczyk, M.; Ekonomiuk, D.; Koliński, A. Optimization of protein models. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2012, 2, 479–493. [Google Scholar] [CrossRef]
- Kmiecik, S.; Kolinski, A. One-dimensional structural properties of proteins in the coarse-grained cabs model. Methods Mol. Biol. 2017, 1484, 83–113. [Google Scholar] [CrossRef] [PubMed]
- Pulawski, W.; Jamroz, M.; Kolinski, M.; Kolinski, A.; Kmiecik, S. Coarse-grained simulations of membrane insertion and folding of small helical proteins using the CABS model. J. Chem. Inf. Model. 2016, 56, 2207–2215. [Google Scholar] [CrossRef]
- Adhikari, A.N.; Freed, K.F.; Sosnick, T.R. De novo prediction of protein folding pathways and structure using the principle of sequential stabilization. Proc. Natl. Acad. Sci. USA 2012, 109, 17442–17447. [Google Scholar] [CrossRef] [Green Version]
- Adhikari, A.N.; Freed, K.F.; Sosnick, T.R. Simplified protein models: Predicting folding pathways and structure using amino acid sequences. Phys. Rev. Lett. 2013, 111, 028103. [Google Scholar] [CrossRef]
- Konrat, R. NMR contributions to structural dynamics studies of intrinsically disordered proteins. J. Magn. Reson. 2014, 241, 74–85. [Google Scholar] [CrossRef] [Green Version]
- Kmiecik, S.; Wabik, J.; Kolinski, M.; Kouza, M.; Kolinski, A. Coarse-Grained Modeling of Protein Dynamics. In Computational Methods to Study the Structure and Dynamics of Biomolecules; Springer: Berlin/Heidelberg, Germany, 2014; Volume 1, pp. 55–79. ISBN 978-3-642-28553-0. [Google Scholar]
- Kurcinski, M.; Ciemny, M.P.; Oleniecki, T.; Kuriata, A.; Badaczewska-Dawid, A.E.; Kolinski, A.; Kmiecik, S. CABS-dock standalone: A toolbox for flexible protein-peptide docking. Bioinformatics 2019. submitted. [Google Scholar]
- Ciemny, M.; Kurcinski, M.; Kamel, K.; Kolinski, A.; Alam, N.; Schueler-Furman, O.; Kmiecik, S. Protein–peptide docking: Opportunities and challenges. Drug Discov. Today 2018, 23, 1530–1537. [Google Scholar] [CrossRef] [PubMed]
- Zambrano, R.; Jamroz, M.; Szczasiuk, A.; Pujols, J.; Kmiecik, S.; Ventura, S. AGGRESCAN3D (A3D): Server for prediction of aggregation properties of protein structures. Nucleic Acids Res. 2015, 43, W306–W313. [Google Scholar] [CrossRef]
- London, N.; Movshovitz-Attias, D.; Schueler-Furman, O. The Structural Basis of Peptide-Protein Binding Strategies. Structure 2010, 18, 188–199. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hur, E.; Pfaff, S.J.; Sturgis Payne, E.; Grøn, H.; Buehrer, B.M.; Fletterick, R.J. Recognition and accommodation at the androgen receptor coactivator binding interface. PLoS Biol. 2004, 2, E274. [Google Scholar] [CrossRef] [PubMed]
- Kabsch, W.; Sander, C. Dictionary of protein secondary structure: Pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 1983, 22, 2577–2637. [Google Scholar] [CrossRef] [PubMed]
- Kussie, P.H.; Gorina, S.; Marechal, V.; Elenbaas, B.; Moreau, J.; Levine, A.J.; Pavletich, N.P. Structure of the MDM2 oncoprotein bound to the p53 tumor suppressor transactivation domain. Science 1996, 274, 948–953. [Google Scholar] [CrossRef] [PubMed]
- Ozenne, V.; Bauer, F.; Salmon, L.; Huang, J.R.; Jensen, M.R.; Segard, S.; Bernadó, P.; Charavay, C.; Blackledge, M. Flexible-meccano: A tool for the generation of explicit ensemble descriptions of intrinsically disordered proteins and their associated experimental observables. Bioinformatics 2012, 28, 1463–1470. [Google Scholar] [CrossRef] [PubMed]
- Feldman, H.J.; Hogue, C.W.V. Probabilistic sampling of protein conformations: New hope for brute force? Proteins Struct. Funct. Genet. 2002, 46, 8–23. [Google Scholar] [CrossRef] [PubMed]
- Vitalis, A.; Pappu, R.V. ABSINTH: A new continuum solvation model for simulations of polypeptides in aqueous solutions. J. Comput. Chem. 2009, 30, 673–699. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baul, U.; Chakraborty, D.; Mugnai, M.L.; Straub, J.E.; Thirumalai, D. Sequence effects on size, shape, and structural heterogeneity in Intrinsically Disordered Proteins. bioRxiv 2018, 427476. [Google Scholar] [CrossRef]
- Estaña, A.; Sibille, N.; Delaforge, E.; Vaisset, M.; Cortés, J.; Bernadó, P. Realistic Ensemble Models of Intrinsically Disordered Proteins Using a Structure-Encoding Coil Database. Structure 2018. [Google Scholar] [CrossRef] [PubMed]




© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ciemny, M.P.; Badaczewska-Dawid, A.E.; Pikuzinska, M.; Kolinski, A.; Kmiecik, S. Modeling of Disordered Protein Structures Using Monte Carlo Simulations and Knowledge-Based Statistical Force Fields. Int. J. Mol. Sci. 2019, 20, 606. https://doi.org/10.3390/ijms20030606
Ciemny MP, Badaczewska-Dawid AE, Pikuzinska M, Kolinski A, Kmiecik S. Modeling of Disordered Protein Structures Using Monte Carlo Simulations and Knowledge-Based Statistical Force Fields. International Journal of Molecular Sciences. 2019; 20(3):606. https://doi.org/10.3390/ijms20030606
Chicago/Turabian StyleCiemny, Maciej Pawel, Aleksandra Elzbieta Badaczewska-Dawid, Monika Pikuzinska, Andrzej Kolinski, and Sebastian Kmiecik. 2019. "Modeling of Disordered Protein Structures Using Monte Carlo Simulations and Knowledge-Based Statistical Force Fields" International Journal of Molecular Sciences 20, no. 3: 606. https://doi.org/10.3390/ijms20030606
APA StyleCiemny, M. P., Badaczewska-Dawid, A. E., Pikuzinska, M., Kolinski, A., & Kmiecik, S. (2019). Modeling of Disordered Protein Structures Using Monte Carlo Simulations and Knowledge-Based Statistical Force Fields. International Journal of Molecular Sciences, 20(3), 606. https://doi.org/10.3390/ijms20030606





