Effect of A22 on the Conformation of Bacterial Actin MreB
Abstract
:1. Introduction
2. Results and Discussion
2.1. A22 Impedes ATP-Induced Backbone Conformational Change in MreB
2.2. MreB Adopts One Main Low-Energy Structure in the Apo, ATP+, and ATP-A22+ States
2.3. The ATP-A22+ MreB Structure Differs from the ATP+ Form
2.4. A22 Affects the Conformational Change of the Ile55-Ile219 and Ser232-Val316 MreB Segments
2.5. Water Dynamics in the Active Site of MreB Protofilament
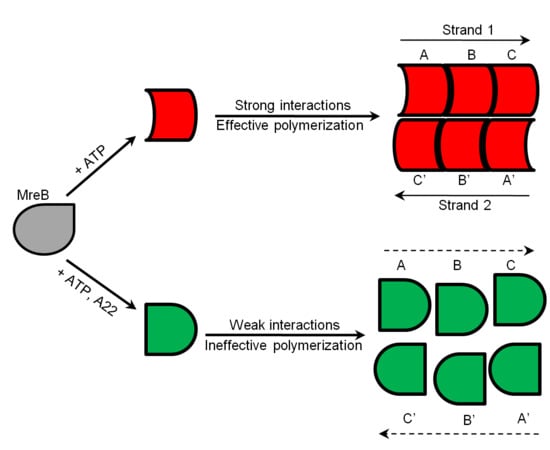
2.6. Proposed Effect of A22
3. Methods
3.1. Preparation of Structures
3.2. Molecular Dynamics Simulations
4. Conclusions
5. Recommendation
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| PDB | Protein Data Bank |
| CcMreB | Caulobacter cresentus MreB |
| NT | Nucleotide |
| MD | Molecular dynamics |
| AMBER | Assisted Model Building and Energy Refinement |
| GAFF | General Amber Force Field |
| RMSD | Root Mean Square Deviation |
| PCA | Principal Component Analysis |
| FEL | Free Energy Landscape |
| RMSF | Root Mean Square Fluctuation |
| AMPPNP | Adenylyl imidodiphosphate |
References
- Jones, L.J.F.; Carballido-Lopez, R.; Errington, J. Control of cell shape in bacteria: Helical, actin-like filaments in Bacillus subtilis. Cell 2001, 104, 913–922. [Google Scholar] [CrossRef]
- van den Ent, F.; Amos, L.A.; Lowe, J. Prokaryotic origin of the actin cytoskeleton. Nature 2001, 413, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Doi, M.; Wachi, M.; Ishino, F.; Tomioka, S.; Ito, M.; Sakagami, Y.; Suzuki, A.; Matsuhashi, M. Determinations of the DNA-sequence of the MreB-gene and of the Gene-products of the Mre-region that function in formation of the rod shape of Escherichia coli cells. J. Bacteriol. 1988, 170, 4619–4624. [Google Scholar] [CrossRef]
- Wachi, M.; Matsuhashi, M. Negative control of cell-division by MreB, a gene that functions in determining the rod shape of Escherichia coli cells. J. Bacteriol. 1989, 171, 3123–3127. [Google Scholar] [CrossRef] [PubMed]
- Soufo, H.J.D.; Graumann, P.L. Actin-like proteins MreB and Mbl from Bacillus subtilis are required for bipolar positioning of replication origins. Curr. Biol. 2003, 13, 1916–1920. [Google Scholar] [CrossRef] [PubMed]
- Kruse, T.; Gerdes, K. Bacterial DNA segregation by the actin-like MreB protein. Trends Cell Biol. 2005, 15, 343–345. [Google Scholar] [CrossRef]
- Kruse, T.; Moller-Jensen, J.; Lobner-Olesen, A.; Gerdes, K. Dysfunctional MreB inhibits chromosome segregation in Escherichia coli. EMBO J. 2003, 22, 5283–5292. [Google Scholar] [CrossRef] [Green Version]
- Gital, Z.; Dye, N.A.; Reisenauer, A.; Wachi, M.; Shapiro, L. MreB actin-mediated segregation of a specific region of a bacterial chromosome. Cell 2005, 120, 329–341. [Google Scholar]
- Soufo, H.J.D.; Graumann, P.L. Bacillus subtilis actin-like protein MreB influences the positioning of the replication machinery and requires membrane proteins MreC/D and other actin-like proteins for proper localization. BMC Cell Biol. 2005, 6, 1–11. [Google Scholar]
- Gitai, Z.; Dye, N.; Shapiro, L. An actin-like gene can determine cell polarity in bacteria. Proc. Natl. Acad. Sci. USA 2004, 101, 8643–8648. [Google Scholar] [CrossRef] [Green Version]
- van den Ent, F.; Izore, T.; Bharat, T.A.M.; Johnson, C.M.; Loewe, J. Bacterial actin MreB forms antiparallel double filaments. Elife 2014, 3, 1–22. [Google Scholar] [CrossRef]
- Kruse, T.; Blagoev, B.; Lobner-Olesen, A.; Wachi, M.; Sasaki, K.; Iwai, N.; Mann, M.; Gerdes, K. Actin homolog MreB and RNA polymerase interact and are both required for chromosome segregation in Escherichia coli. Genes Dev. 2006, 20, 113–124. [Google Scholar] [CrossRef] [Green Version]
- Iwai, N.; Nagai, K.; Wachi, M. Novel S-benzylisothiourea compound that induces spherical cells in Escherichia coli probably by acting on a rod-shape-determining protein(s) other than penicillin-binding protein 2. Biosci. Biotechnol. Biochem. 2002, 66, 2658–2662. [Google Scholar] [CrossRef]
- Awuni, Y.; Jiang, S.M.; Robinson, R.C.; Mu, Y.G. Exploring the A22-bacterial actin MreB interaction through molecular dynamics simulations. J. Phys. Chem. B 2016, 120, 9867–9874. [Google Scholar] [CrossRef]
- Esue, O.; Cordero, M.; Wirtz, D.; Tseng, Y. The assembly of MreB, a prokaryotic homolog of actin. J. Biol. Chem. 2005, 280, 2628–2635. [Google Scholar] [CrossRef]
- Esue, O.; Wirtz, D.; Tseng, Y. GTPase activity, structure, and mechanical properties of filaments assembled from bacterial cytoskeleton protein MreB. J. Bacteriol. 2006, 188, 968–976. [Google Scholar] [CrossRef]
- Bean, G.J.; Amann, K.J. Polymerization properties of the Thermotoga maritima actin MreB: Roles of temperature, nucleotides, and ions. Biochemistry 2008, 47, 826–835. [Google Scholar] [CrossRef]
- Mayer, J.A.; Amann, K.J. Assembly properties of the Bacillus subtilis actin, MreB. Cell Motil. Cytoskeleton 2009, 66, 109–118. [Google Scholar] [CrossRef]
- Maisuradze, G.G.; Leitner, D.M. Free energy landscape of a biomolecule in dihedral principal component space: Sampling convergence and correspondence between structures and minima. Proteins Struct. Funct. Bioinf. 2007, 67, 569–578. [Google Scholar] [CrossRef]
- Parke, C.L.; Wojcik, E.J.; Kim, S.; Worthylake, D.K. ATP hydrolysis in Eg5 kinesin involves a catalytic two-water mechanism. J. Biol. Chem. 2010, 285, 5859–5867. [Google Scholar] [CrossRef]
- Grigorenko, B.L.; Nemukhin, A.V.; Shadrina, M.S.; Topol, I.A.; Burt, S.K. Mechanisms of guanosine triphosphate hydrolysis by Ras and Ras-GAP proteins as rationalized by ab initio QM/MM simulations. Proteins Struct. Funct. Bioinf. 2007, 66, 456–466. [Google Scholar] [CrossRef] [PubMed]
- Grigorenko, B.L.; Nemukhin, A.V.; Topol, I.A.; Cachau, R.E.; Burt, S.K. QM/MM modeling the Ras-GAP catalyzed hydrolysis of guanosine triphosphate. Proteins Struct. Funct. Bioinf. 2005, 60, 495–503. [Google Scholar] [CrossRef]
- Popp, D.; Narita, A.; Maeda, K.; Fujisawa, T.; Ghoshdastider, U.; Iwasa, M.; Maeda, Y.; Robinson, R.C. Filament structure, organization, and dynamics in MreB sheets. J. Biol. Chem. 2010, 285, 15858–15865. [Google Scholar] [CrossRef]
- Kim, S.Y.; Gitai, Z.; Kinkhabwala, A.; Shapiro, L.; Moerner, W.E. Single molecules of the bacterial actin MreB undergo directed treadmilling motion in Caulobacter crescentus. Proc. Natl. Acad. Sci. USA 2006, 103, 10929. [Google Scholar] [CrossRef]
- Aylett, C.H.S.; Löwe, J.; Amos, L.A. Chapter One—New insights into the mechanisms of cytomotive actin and tubulin filaments. Int. Rev. Cell Mol. Biol. 2011, 292, 1–71. [Google Scholar]
- Arnold, K.; Bordoli, L.; Kopp, J.; Schwede, T. The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics 2006, 22, 195–201. [Google Scholar] [CrossRef]
- Cornell, W.D.; Cieplak, P.; Bayly, C.I.; Gould, I.R.; Merz, K.M., Jr.; Ferguson, D.M.; Spellmeyer, D.C.; Fox, T.; Caldwell, J.W.; Kollman, P.A. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 1995, 5179–5197. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G.A.; et al. Gaussian 09; Gaussian, Inc.: Wallingford, CT, USA, 2009. [Google Scholar]
- Dupradeau, F.-Y.; Pigache, A.; Zaffran, T.; Savineau, C.; Lelong, R.; Grivel, N.; Lelong, D.; Rosanski, W.; Cieplak, P. The R.E.D. tools: Advances in RESP and ESP charge derivation and force field library building. Phys. Chem. Chem. Phys. 2010, 12, 7821–7839. [Google Scholar] [CrossRef]
- Wang, J.; Wang, W.; Kollman, P.A.; Case, D.A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 2006, 25, 247–260. [Google Scholar] [CrossRef]
- Case, D.A.; Cheatham, T.E.; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.M.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber biomolecular simulation programs. J. Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef] [Green Version]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald - An N.log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 2007, 126, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Shan, Y.; Arkhipov, A.; Kim, E.T.; Pan, A.C.; Shaw, D.E. Transitions to catalytically inactive conformations in EGFR kinase. Proc. Natl. Acad. Sci. USA 2013, 110, 7270. [Google Scholar] [CrossRef] [PubMed]









| System | State of MreB Protofilament | ATP | A22 | Mg2+ | Time (ns) | Repeats |
|---|---|---|---|---|---|---|
| 1 | Apo (no ATP and A22) | - | - | + | 100 | 3 |
| 2 | ATP-bound (ATP+) | + | - | + | 100 | 3 |
| 3 | ATP-A22-bound (ATP-A22+) | + | + | + | 100 | 3 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Awuni, E.; Mu, Y. Effect of A22 on the Conformation of Bacterial Actin MreB. Int. J. Mol. Sci. 2019, 20, 1304. https://doi.org/10.3390/ijms20061304
Awuni E, Mu Y. Effect of A22 on the Conformation of Bacterial Actin MreB. International Journal of Molecular Sciences. 2019; 20(6):1304. https://doi.org/10.3390/ijms20061304
Chicago/Turabian StyleAwuni, Elvis, and Yuguang Mu. 2019. "Effect of A22 on the Conformation of Bacterial Actin MreB" International Journal of Molecular Sciences 20, no. 6: 1304. https://doi.org/10.3390/ijms20061304