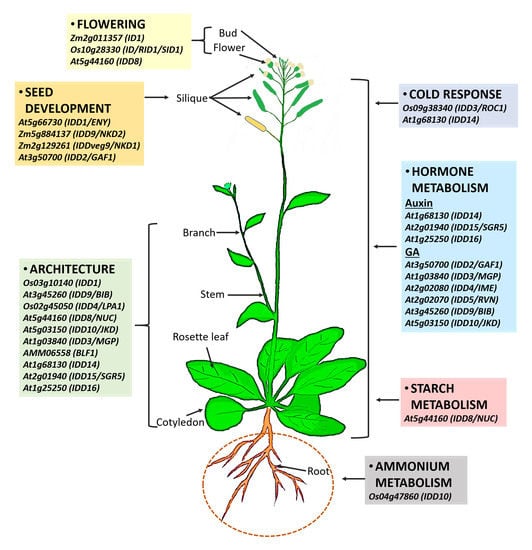
Role of the INDETERMINATE DOMAIN Genes in Plants
Abstract
:1. Introduction
2. Structure and Phylogenetic Analysis of IDD Proteins
3. Biological Functions of IDDs
3.1. Modulation of Sugar Metabolism and Floral Transition
3.2. Starch Accumulation and Cold Response
3.3. Regulation of Seed Development
3.4. Modulation of Plant Architecture, Shoot Gravitropism, and Secondary Cell Wall Formation
3.5. Regulation of Hormonal Signaling
3.6. Ammonium Metabolism
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| IDD | INDETERMINATE DOMAIN |
| LPA | Loose Plant Architecture |
| TPS | Trehalose-6-phosphate synthase |
| FT | Flowering Locus T |
| RID | Rice Indeterminate |
| Ehd | Early heading date |
| SUS | Sucrose Synthase |
| BLF | Broadleaf |
| AMT | Ammonium Transporter |
| NKD | Naked Endosperm |
| ENY | Enhydrous |
| HD | Heading Date |
| Rft1 | Rice Flowering Locus T1 |
References
- Pabo, C.O.; Sauer, R.T. Transcription factors: Structural families and principles of DNA recognition. Annu. Rev. Biochem. 1992, 61, 1053–1095. [Google Scholar] [CrossRef] [PubMed]
- Englbrecht, C.C.; Schoof, H.; Bohm, S. Conservation, diversification and expansion of C2H2 zinc finger proteins in the Arabidopsis thaliana genome. BMC Genom. 2004, 5, 39. [Google Scholar] [CrossRef]
- Liu, Q.G.; Wang, Z.C.; Xu, X.M.; Zhang, H.Z.; Li, C.H. Genome-Wide Analysis of C2H2 Zinc-Finger Family Transcription Factors and Their Responses to Abiotic Stresses in Poplar (Populus trichocarpa). PLoS ONE 2015, 10, e0134753. [Google Scholar] [CrossRef]
- Colasanti, J.; Tremblay, R.; Wong, A.Y.; Coneva, V.; Kozaki, A.; Mable, B.K. The maize INDETERMINATE1 flowering time regulator defines a highly conserved zinc finger protein family in higher plants. BMC Genom. 2006, 7, 158. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, C.; Frommer, W.B. A novel zinc finger protein encoded by a couch potato homologue from Solanum tuberosum enables a sucrose transport-deficient yeast strain to grow on sucrose. Mol. Gen. Genet. 1995, 247, 759–763. [Google Scholar] [CrossRef]
- Cui, D.; Zhao, J.; Jing, Y.; Fan, M.; Liu, J.; Wang, Z.; Xin, W.; Hu, Y. The arabidopsis IDD14, IDD15, and IDD16 cooperatively regulate lateral organ morphogenesis and gravitropism by promoting auxin biosynthesis and transport. PLoS Genet. 2013, 9, e1003759. [Google Scholar] [CrossRef]
- Feurtado, J.A.; Huang, D.; Wicki-Stordeur, L.; Hemstock, L.E.; Potentier, M.S.; Tsang, E.W.; Cutler, A.J. The Arabidopsis C2H2 zinc finger INDETERMINATE DOMAIN1/ENHYDROUS promotes the transition to germination by regulating light and hormonal signaling during seed maturation. Plant Cell 2011, 23, 1772–1794. [Google Scholar] [CrossRef]
- Fukazawa, J.; Teramura, H.; Murakoshi, S.; Nasuno, K.; Nishida, N.; Ito, T.; Yoshida, M.; Kamiya, Y.; Yamaguchi, S.; Takahashi, Y. DELLAs function as coactivators of GAI-ASSOCIATED FACTOR1 in regulation of gibberellin homeostasis and signaling in Arabidopsis. Plant Cell 2014, 26, 2920–2938. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, H.; Hirano, K.; Sato, T.; Mitsuda, N.; Nomoto, M.; Maeo, K.; Koketsu, E.; Mitani, R.; Kawamura, M.; Ishiguro, S.; et al. DELLA protein functions as a transcriptional activator through the DNA binding of the indeterminate domain family proteins. Proc. Natl. Acad. Sci. USA 2014, 111, 7861–7866. [Google Scholar] [CrossRef] [PubMed]
- Xuan, Y.H.; Priatama, R.A.; Huang, J.; Je, B.I.; Liu, J.M.; Park, S.J.; Piao, H.L.; Son, D.Y.; Lee, J.J.; Park, S.H.; et al. Indeterminate domain 10 regulates ammonium-mediated gene expression in rice roots. New Phytol. 2013, 197, 791–804. [Google Scholar] [CrossRef]
- Hirano, Y.; Nakagawa, M.; Suyama, T.; Murase, K.; Shirakawa, M.; Takayama, S.; Sun, T.P.; Hakoshima, T. Structure of the SHR-SCR heterodimer bound to the BIRD/IDD transcriptional factor JKD. Nat. Plants 2017, 3, 17010. [Google Scholar] [CrossRef]
- Clemens, K.R.; Wolf, V.; Mcbryant, S.J.; Zhang, P.H.; Liao, X.B.; Wright, P.E.; Gottesfeld, J.M. Molecular-Basis for Specific Recognition of Both Rna and DNA by a Zinc Finger Protein. Science 1993, 260, 530–533. [Google Scholar] [CrossRef]
- Searles, M.A.; Lu, D.; Klug, A. The role of the central zinc fingers of transcription factor IIIA in binding to 5 S RNA. J. Mol. Biol. 2000, 301, 47–60. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic Local Alignment Search Tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The Rapid Generation of Mutation Data Matrices from Protein Sequences. Comput. Appl. Biosci. 1992, 8, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Singleton, W.R. Inheritance of indeterminate growth in maize. J. Hered. 1946, 37, 61–64. [Google Scholar] [CrossRef]
- Wong, A.Y.; Colasanti, J. Maize floral regulator protein INDETERMINATE1 is localized to developing leaves and is not altered by light or the sink/source transition. J. Exp. Bot. 2007, 58, 403–414. [Google Scholar] [CrossRef] [PubMed]
- Kozaki, A.; Hake, S.; Colasanti, J. The maize ID1 flowering time regulator is a zinc finger protein with novel DNA binding properties. Nucleic Acids Res. 2004, 32, 1710–1720. [Google Scholar] [CrossRef]
- Muszynski, M.G.; Dam, T.; Li, B.; Shirbroun, D.M.; Hou, Z.; Bruggemann, E.; Archibald, R.; Ananiev, E.V.; Danilevskaya, O.N. Delayed flowering1 encodes a basic leucine zipper protein that mediates floral inductive signals at the shoot apex in maize. Plant Physiol. 2006, 142, 1523–1536. [Google Scholar] [CrossRef]
- Meng, X.; Muszynski, M.G.; Danilevskaya, O.N. The FT-like ZCN8 Gene Functions as a Floral Activator and Is Involved in Photoperiod Sensitivity in Maize. Plant Cell 2011, 23, 942–960. [Google Scholar] [CrossRef]
- Matsubara, K.; Yamanouchi, U.; Wang, Z.X.; Minobe, Y.; Izawa, T.; Yano, M. Ehd2, a rice ortholog of the maize INDETERMINATE1 gene, promotes flowering by up-regulating Ehd1. Plant Physiol. 2008, 148, 1425–1435. [Google Scholar] [CrossRef] [PubMed]
- Park, S.J.; Kim, S.L.; Lee, S.; Je, B.I.; Piao, H.L.; Park, S.H.; Kim, C.M.; Ryu, C.H.; Park, S.H.; Xuan, Y.H.; et al. Rice Indeterminate 1 (OsId1) is necessary for the expression of Ehd1 (Early heading date 1) regardless of photoperiod. Plant J. 2008, 56, 1018–1029. [Google Scholar] [CrossRef]
- Wu, C.; You, C.; Li, C.; Long, T.; Chen, G.; Byrne, M.E.; Zhang, Q. RID1, encoding a Cys2/His2-type zinc finger transcription factor, acts as a master switch from vegetative to floral development in rice. Proc. Natl. Acad. Sci. USA 2008, 105, 12915–12920. [Google Scholar] [CrossRef] [Green Version]
- Bernier, G.; Havelange, A.; Houssa, C.; Petitjean, A.; Lejeune, P. Physiological Signals That Induce Flowering. Plant Cell 1993, 5, 1147–1155. [Google Scholar] [CrossRef] [PubMed]
- Seo, P.J.; Ryu, J.; Kang, S.K.; Park, C.M. Modulation of sugar metabolism by an INDETERMINATE DOMAIN transcription factor contributes to photoperiodic flowering in Arabidopsis. Plant J. 2011, 65, 418–429. [Google Scholar] [CrossRef] [PubMed]
- King, R.W.; Hisamatsu, T.; Goldschmidt, E.E.; Blundell, C. The nature of floral signals in Arabidopsis. I. Photosynthesis and a far-red photoresponse independently regulate flowering by increasing expression of FLOWERING LOCUS T (FT). J. Exp. Bot. 2008, 59, 3811–3820. [Google Scholar] [CrossRef] [Green Version]
- Jeong, E.Y.; Seo, P.J.; Woo, J.C.; Park, C.M. AKIN10 delays flowering by inactivating IDD8 transcription factor through protein phosphorylation in Arabidopsis. BMC Plant Biol. 2015, 15, 110. [Google Scholar] [CrossRef]
- Coneva, V.; Guevara, D.; Rothstein, S.J.; Colasanti, J. Transcript and metabolite signature of maize source leaves suggests a link between transitory starch to sucrose balance and the autonomous floral transition. J. Exp. Bot. 2012, 63, 5079–5092. [Google Scholar] [CrossRef] [Green Version]
- Figueroa, C.M.; Lunn, J.E. A Tale of Two Sugars: Trehalose 6-Phosphate and Sucrose. Plant Physiol. 2016, 172, 7–27. [Google Scholar] [CrossRef]
- Gomez-Ariza, J.; Galbiati, F.; Goretti, D.; Brambilla, V.; Shrestha, R.; Pappolla, A.; Courtois, B.; Fornara, F. Loss of floral repressor function adapts rice to higher latitudes in Europe. J. Exp. Bot. 2015, 66, 2027–2039. [Google Scholar] [CrossRef] [Green Version]
- Wahl, V.; Ponnu, J.; Schlereth, A.; Arrivault, S.; Langenecker, T.; Franke, A.; Feil, R.; Lunn, J.E.; Stitt, M.; Schmid, M. Regulation of flowering by trehalose-6-phosphate signaling in Arabidopsis thaliana. Science 2013, 339, 704–707. [Google Scholar] [CrossRef]
- Deng, L.; Li, L.; Zhang, S.; Shen, J.; Li, S.; Hu, S.; Peng, Q.; Xiao, J.; Wu, C. Suppressor of rid1 (SID1) shares common targets with RID1 on florigen genes to initiate floral transition in rice. PLoS Genet. 2017, 13, e1006642. [Google Scholar] [CrossRef]
- Thomashow, M.F. Role of cold-responsive genes in plant freezing tolerance. Plant Physiol. 1998, 118, 1–8. [Google Scholar] [CrossRef]
- Li, L.; Foster, C.M.; Gan, Q.; Nettleton, D.; James, M.G.; Myers, A.M.; Wurtele, E.S. Identification of the novel protein QQS as a component of the starch metabolic network in Arabidopsis leaves. Plant J. 2009, 58, 485–498. [Google Scholar] [CrossRef] [Green Version]
- Seo, P.J.; Kim, M.J.; Ryu, J.Y.; Jeong, E.Y.; Park, C.M. Two splice variants of the IDD14 transcription factor competitively form nonfunctional heterodimers which may regulate starch metabolism. Nat. Commun. 2011, 2, 303. [Google Scholar] [CrossRef] [Green Version]
- Dou, M.; Cheng, S.; Zhao, B.; Xuan, Y.; Shao, M. The Indeterminate Domain Protein ROC1 Regulates Chilling Tolerance via Activation of DREB1B/CBF1 in Rice. Int. J. Mol. Sci. 2016, 17, 233. [Google Scholar] [CrossRef]
- Jaglo-Ottosen, K.R.; Gilmour, S.J.; Zarka, D.G.; Schabenberger, O.; Thomashow, M.F. Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 1998, 280, 104–106. [Google Scholar] [CrossRef]
- Agarwal, M.; Hao, Y.; Kapoor, A.; Dong, C.H.; Fujii, H.; Zheng, X.; Zhu, J.K. A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J. Biol. Chem. 2006, 281, 37636–37645. [Google Scholar] [CrossRef]
- Chinnusamy, V.; Zhu, J.K.; Sunkar, R. Gene regulation during cold stress acclimation in plants. Methods Mol. Biol. 2010, 639, 39–55. [Google Scholar]
- Dong, C.H.; Agarwal, M.; Zhang, Y.; Xie, Q.; Zhu, J.K. The negative regulator of plant cold responses, HOS1, is a RING E3 ligase that mediates the ubiquitination and degradation of ICE1. Proc. Natl. Acad. Sci. USA 2006, 103, 8281–8286. [Google Scholar] [CrossRef] [Green Version]
- Shibasaki, K.; Uemura, M.; Tsurumi, S.; Rahman, A. Auxin response in Arabidopsis under cold stress: Underlying molecular mechanisms. Plant Cell 2009, 21, 3823–3838. [Google Scholar] [CrossRef]
- Yi, G.; Neelakandan, A.K.; Gontarek, B.C.; Vollbrecht, E.; Becraft, P.W. The naked endosperm genes encode duplicate INDETERMINATE domain transcription factors required for maize endosperm cell patterning and differentiation. Plant Physiol. 2015, 167, 443–456. [Google Scholar] [CrossRef]
- Gontarek, B.C.; Neelakandan, A.K.; Wu, H.; Becraft, P.W. NKD Transcription Factors Are Central Regulators of Maize Endosperm Development. Plant Cell 2016, 28, 2916–2936. [Google Scholar] [CrossRef]
- Huang, P.; Yoshida, H.; Yano, K.; Kinoshita, S.; Kawai, K.; Koketsu, E.; Hattori, M.; Takehara, S.; Huang, J.; Hirano, K.; et al. OsIDD2, a zinc finger and INDETERMINATE DOMAIN protein, regulates secondary cell wall formation. J. Integr. Plant Biol. 2018, 60, 130–143. [Google Scholar] [CrossRef]
- Hirano, K.; Aya, K.; Morinaka, Y.; Nagamatsu, S.; Sato, Y.; Antonio, B.A.; Namiki, N.; Nagamura, Y.; Matsuoka, M. Survey of genes involved in rice secondary cell wall formation through a co-expression network. Plant Cell Physiol. 2013, 54, 1803–1821. [Google Scholar] [CrossRef]
- Liu, J.M.; Park, S.J.; Huang, J.; Lee, E.J.; Xuan, Y.H.; Je, B.I.; Kumar, V.; Priatama, R.A.; Raj, K.V.; Kim, S.H.; et al. Loose Plant Architecture1 (LPA1) determines lamina joint bending by suppressing auxin signalling that interacts with C-22-hydroxylated and 6-deoxo brassinosteroids in rice. J. Exp. Bot. 2016, 67, 1883–1895. [Google Scholar] [CrossRef] [Green Version]
- Wu, X.; Tang, D.; Li, M.; Wang, K.; Cheng, Z. Loose Plant Architecture1, an INDETERMINATE DOMAIN protein involved in shoot gravitropism, regulates plant architecture in rice. Plant Physiol. 2013, 161, 317–329. [Google Scholar] [CrossRef]
- Jost, M.; Hensel, G.; Kappel, C.; Druka, A.; Sicard, A.; Hohmann, U.; Beier, S.; Himmelbach, A.; Waugh, R.; Kumlehn, J.; et al. The INDETERMINATE DOMAIN Protein BROAD LEAF1 Limits Barley Leaf Width by Restricting Lateral Proliferation. Curr. Biol. 2016, 26, 903–909. [Google Scholar] [CrossRef]
- Richards, R.A. Selectable traits to increase crop photosynthesis and yield of grain crops. J. Exp. Bot. 2000, 51, 447–458. [Google Scholar] [CrossRef] [Green Version]
- Peng, S.; Khush, G.S.; Virk, P.; Tang, Q.; Zou, Y. Progress in ideotype breeding to increase rice yield potential. Field Crop. Res. 2008, 108, 32–38. [Google Scholar] [CrossRef]
- Kim, J.Y.; Ryu, J.Y.; Baek, K.; Park, C.M. High temperature attenuates the gravitropism of inflorescence stems by inducing SHOOT GRAVITROPISM 5 alternative splicing in Arabidopsis. New Phytol. 2016, 209, 265–279. [Google Scholar] [CrossRef]
- Ryu, J.Y.; Kim, J.Y.; Park, C.M. Adaptive thermal control of stem gravitropism through alternative RNA splicing in Arabidopsis. Plant Signal Behav. 2015, 10, e1093715. [Google Scholar] [CrossRef] [Green Version]
- Hassan, H.; Scheres, B.; Blilou, I. JACKDAW controls epidermal patterning in the Arabidopsis root meristem through a non-cell-autonomous mechanism. Development 2010, 137, 1523–1529. [Google Scholar] [CrossRef] [Green Version]
- Ogasawara, H.; Kaimi, R.; Colasanti, J.; Kozaki, A. Activity of transcription factor JACKDAW is essential for SHR/SCR-dependent activation of SCARECROW and MAGPIE and is modulated by reciprocal interactions with MAGPIE, SCARECROW and SHORT ROOT. Plant Mol. Biol. 2011, 77, 489–499. [Google Scholar] [CrossRef] [Green Version]
- Welch, D.; Hassan, H.; Blilou, I.; Immink, R.; Heidstra, R.; Scheres, B. Arabidopsis JACKDAW and MAGPIE zinc finger proteins delimit asymmetric cell division and stabilize tissue boundaries by restricting SHORT-ROOT action. Genes Dev. 2007, 21, 2196–2204. [Google Scholar] [CrossRef]
- Yoon, E.K.; Dhar, S.; Lee, M.H.; Song, J.H.; Lee, S.A.; Kim, G.; Jang, S.; Choi, J.W.; Choe, J.E.; Kim, J.H.; et al. Conservation and Diversification of the SHR-SCR-SCL23 Regulatory Network in the Development of the Functional Endodermis in Arabidopsis Shoots. Mol. Plant 2016, 9, 1197–1209. [Google Scholar] [CrossRef]
- Salazar-Henao, J.E.; Velez-Bermudez, I.C.; Schmidt, W. The regulation and plasticity of root hair patterning and morphogenesis. Development 2016, 143, 1848–1858. [Google Scholar] [CrossRef] [Green Version]
- Durbak, A.; Yao, H.; McSteen, P. Hormone signaling in plant development. Curr. Opin. Plant Biol. 2012, 15, 92–96. [Google Scholar] [CrossRef]
- Zentella, R.; Zhang, Z.L.; Park, M.; Thomas, S.G.; Endo, A.; Murase, K.; Fleet, C.M.; Jikumaru, Y.; Nambara, E.; Kamiya, Y.; et al. Global analysis of della direct targets in early gibberellin signaling in Arabidopsis. Plant Cell 2007, 19, 3037–3057. [Google Scholar] [CrossRef]
- Zhang, Z.L.; Ogawa, M.; Fleet, C.M.; Zentella, R.; Hu, J.; Heo, J.O.; Lim, J.; Kamiya, Y.; Yamaguchi, S.; Sun, T.P. Scarecrow-like 3 promotes gibberellin signaling by antagonizing master growth repressor DELLA in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 2160–2165. [Google Scholar] [CrossRef]
- Vanneste, S.; Friml, J. Auxin: A trigger for change in plant development. Cell 2009, 136, 1005–1016. [Google Scholar] [CrossRef] [PubMed]
- Woodward, A.W.; Bartel, B. Auxin: Regulation, action, and interaction. Ann. Bot. 2005, 95, 707–735. [Google Scholar] [CrossRef]
- Zhao, Y. Auxin biosynthesis and its role in plant development. Annu. Rev. Plant Biol. 2010, 61, 49–64. [Google Scholar] [CrossRef] [Green Version]
- Cheng, Y.; Dai, X.; Zhao, Y. Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev. 2006, 20, 1790–1799. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, J.I.; Murphy, A.S.; Baek, D.; Lee, S.W.; Yun, D.J.; Bressan, R.A.; Narasimhan, M.L. YUCCA6 over-expression demonstrates auxin function in delaying leaf senescence in Arabidopsis thaliana. J. Exp. Bot. 2011, 62, 3981–3992. [Google Scholar] [CrossRef] [Green Version]
- Kim, J.I.; Sharkhuu, A.; Jin, J.B.; Li, P.; Jeong, J.C.; Baek, D.; Lee, S.Y.; Blakeslee, J.J.; Murphy, A.S.; Bohnert, H.J.; et al. yucca6, a dominant mutation in Arabidopsis, affects auxin accumulation and auxin-related phenotypes. Plant Physiol. 2007, 145, 722–735. [Google Scholar] [CrossRef]
- Zhao, Y.; Christensen, S.K.; Fankhauser, C.; Cashman, J.R.; Cohen, J.D.; Weigel, D.; Chory, J. A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 2001, 291, 306–309. [Google Scholar] [CrossRef]
- Shi, H.; Zhang, S.; Lin, D.; Wei, Y.; Yan, Y.; Liu, G.; Reiter, R.J.; Chan, Z. Zinc finger of Arabidopsis thaliana 6 is involved in melatonin-mediated auxin signaling through interacting INDETERMINATE DOMAIN15 and INDOLE-3-ACETIC ACID 17. J. Pineal Res. 2018, 65, e12494. [Google Scholar] [CrossRef]
- Fukumorita, T.; Chino, M. Sugar, Amino Acid and Inorganic Contents in Rice Phloem Sap. Plant Cell Physiol. 1982, 23, 273–283. [Google Scholar]
- Gutierrez, R.A.; Stokes, T.L.; Thum, K.; Xu, X.; Obertello, M.; Katari, M.S.; Tanurdzic, M.; Dean, A.; Nero, D.C.; McClung, C.R.; et al. Systems approach identifies an organic nitrogen-responsive gene network that is regulated by the master clock control gene CCA1. Proc. Natl. Acad. Sci. USA 2008, 105, 4939–4944. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Patterson, K.; Cakmak, T.; Cooper, A.; Lager, I.; Rasmusson, A.G.; Escobar, M.A. Distinct signalling pathways and transcriptome response signatures differentiate ammonium- and nitrate-supplied plants. Plant Cell Environ. 2010, 33, 1486–1501. [Google Scholar] [CrossRef] [PubMed]
- Patterson, K.; Walters, L.A.; Cooper, A.M.; Olvera, J.G.; Rosas, M.A.; Rasmusson, A.G.; Escobar, M.A. Nitrate-Regulated Glutaredoxins Control Arabidopsis Primary Root Growth. Plant Physiol. 2016, 170, 989–999. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Guegler, K.; LaBrie, S.T.; Crawford, N.M. Genomic analysis of a nutrient response in Arabidopsis reveals diverse expression patterns and novel metabolic and potential regulatory genes induced by nitrate. Plant Cell 2000, 12, 1491–1509. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Tischner, R.; Gutierrez, R.A.; Hoffman, M.; Xing, X.; Chen, M.; Coruzzi, G.; Crawford, N.M. Genomic analysis of the nitrate response using a nitrate reductase-null mutant of Arabidopsis. Plant Physiol. 2004, 136, 2512–2522. [Google Scholar] [CrossRef] [PubMed]
- Xuan, Y.H.; Priatama, R.A.; Kumar, V.; Han, C.D. Regulatory role of indeterminate domain 10 (IDD10) in ammonium-dependent gene expression in rice roots. Plant Signal Behav. 2013, 8, e24139. [Google Scholar] [CrossRef]
- Xuan, Y.H.; Kumar, V.; Zhu, X.F.; Je, B.I.; Kim, C.M.; Huang, J.; Cho, J.H.; Yi, G.; Han, C.D. IDD10 is Involved in the Interaction between NH4+ and Auxin Signaling in Rice Roots. J. Plant Biol. 2018, 61, 72–79. [Google Scholar] [CrossRef]
- Coelho, C.P.; Huang, P.; Lee, D.Y.; Brutnell, T.P. Making Roots, Shoots, and Seeds: IDD Gene Family Diversification in Plants. Trends Plant Sci. 2018, 23, 66–78. [Google Scholar] [CrossRef]
- Liu, T.; Reinhart, B.J.; Magnani, E.; Huang, T.; Kerstetter, R.; Barton, M.K. Of blades and branches: Understanding and expanding the Arabidopsis ad/abaxial regulatory network through target gene identification. Cold Spring Harb. Symp. Quant. Biol. 2012, 77, 31–45. [Google Scholar] [CrossRef]
- Moreno-Risueno, M.A.; Sozzani, R.; Yardimci, G.G.; Petricka, J.J.; Vernoux, T.; Blilou, I.; Alonso, J.; Winter, C.M.; Ohler, U.; Scheres, B.; et al. Transcriptional control of tissue formation throughout root development. Science 2015, 350, 426–430. [Google Scholar] [CrossRef]
- Morita, M.T.; Sakaguchi, K.; Kiyose, S.; Taira, K.; Kato, T.; Nakamura, M.; Tasaka, M. A C2H2-type zinc finger protein, SGR5, is involved in early events of gravitropism in Arabidopsis inflorescence stems. Plant J. 2006, 47, 619–628. [Google Scholar] [CrossRef] [Green Version]
- Reinhart, B.J.; Liu, T.; Newell, N.R.; Magnani, E.; Huang, T.; Kerstetter, R.; Michaels, S.; Barton, M.K. Establishing a framework for the Ad/abaxial regulatory network of Arabidopsis: Ascertaining targets of class III homeodomain leucine zipper and KANADI regulation. Plant Cell 2013, 25, 3228–3249. [Google Scholar] [CrossRef] [PubMed]
- Tanimoto, M.; Tremblay, R.; Colasanti, J. Altered gravitropic response, amyloplast sedimentation and circumnutation in the Arabidopsis shoot gravitropism 5 mutant are associated with reduced starch levels. Plant Mol. Biol. 2008, 67, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Levesque, M.P.; Vernoux, T.; Busch, W.; Cui, H.C.; Wang, J.Y.; Blilou, I.; Hassan, H.; Nakajima, K.; Matsumoto, N.; Lohmann, J.U.; et al. Whole-genome analysis of the SHORT-ROOT developmental pathway in Arabidopsis (vol 4, pg 739, 2006). PLoS Biol. 2006, 4, 1284. [Google Scholar]
- Ingkasuwan, P.; Netrphan, S.; Prasitwattanaseree, S.; Tanticharoen, M.; Bhumiratana, S.; Meechai, A.; Chaijaruwanich, J.; Takahashi, H.; Cheevadhanarak, S. Inferring transcriptional gene regulation network of starch metabolism in Arabidopsis thaliana leaves using graphical Gaussian model. BMC Syst. Biol. 2012, 6, 100. [Google Scholar] [CrossRef] [PubMed]
- Bustillo-Avendano, E.; Ibanez, S.; Sanz, O.; Barros, J.A.S.; Gude, I.; Perianez-Rodriguez, J.; Micol, J.L.; Del Pozo, J.C.; Moreno-Risueno, M.A.; Perez-Perez, J.M. Regulation of Hormonal Control, Cell Reprogramming, and Patterning during De Novo Root Organogenesis. Plant Physiol. 2018, 176, 1709–1727. [Google Scholar] [CrossRef] [PubMed]
- Long, Y.; Goedhart, J.; Schneijderberg, M.; Terpstra, I.; Shimotohno, A.; Bouchet, B.P.; Akhmanova, A.; Gadella, T.W., Jr.; Heidstra, R.; Scheres, B.; et al. SCARECROW-LIKE23 and SCARECROW jointly specify endodermal cell fate but distinctly control SHORT-ROOT movement. Plant J. 2015, 84, 773–784. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Long, Y.C.; Smet, W.; Cruz-Ramirez, A.; Castelijns, B.; de Jonge, W.; Mahonen, A.P.; Bouchet, B.P.; Perez, G.S.; Akhmanova, A.; Scheres, B.; et al. Arabidopsis BIRD Zinc Finger Proteins Jointly Stabilize Tissue Boundaries by Confining the Cell Fate Regulator SHORT-ROOT and Contributing to Fate Specification. Plant Cell 2015, 27, 1185–1199. [Google Scholar] [CrossRef] [Green Version]
- Long, Y.; Stahl, Y.; Weidtkamp-Peters, S.; Postma, M.; Zhou, W.; Goedhart, J.; Sanchez-Perez, M.I.; Gadella, T.W.J.; Simon, R.; Scheres, B.; et al. In vivo FRET-FLIM reveals cell-type-specific protein interactions in Arabidopsis roots. Nature 2017, 548, 97–102. [Google Scholar] [CrossRef]
- Colasanti, J.; Yuan, Z.; Sundaresan, V. The indeterminate gene encodes a zinc finger protein and regulates a leaf-generated signal required for the transition to flowering in maize. Cell 1998, 93, 593–603. [Google Scholar] [CrossRef]
- Galinat, W.C.; Naylor, A.W. Relation of Photoperiod to Inflorescence proliferation in Zea Mays L. Am. J. Bot. 1951, 38, 38–47. [Google Scholar] [CrossRef]
- Becraft, P.W.; Asuncion-Crabb, Y. Positional cues specify and maintain aleurone cell fate in maize endosperm development. Development 2000, 127, 4039–4048. [Google Scholar] [PubMed]







| Gene Name | Phenotype | Key Role | References |
|---|---|---|---|
| Arabidopsis | |||
| At5g66730 (AtIDD1)/ENHYDROUS | Overexpression of AtIDD1 increases starch retention, endosperm-specific fatty acids, and defective mucilage extrusion in mature seed. | Seed development | [7] |
| At3g50700 (AtIDD2)/CARRION CROW/GAF1 | idd2/idd1 double mutant exhibit decreased GA responsiveness, whereas IDD2 overexpressor enhance GA responsiveness. | GA signaling | [8] |
| At1g03840 (AtIDD3)/MAGPIE | SHORT-ROOT (SHR) transcription factor directly targets AtIDD3/MGP, which plays key roles in specifying the radial root patterning and root stem cell niche. | Root development, DELLA interacting protein | [9,83] |
| At2g02080 (AtIDD4)/IMPERIAL EAGLE | IDD4 and GR-REV act together to coordinate and promote outgrowth of a flattened leaf blade. IDD4 is an SHR target. | DELLA interacting protein, leaf polarity | [79,81] |
| At2g02070 (AtIDD5)/RAVEN | Positive regulator of starch synthase 4. atidd5 mutants have deformed chloroplasts and starch granules. | DELLA interacting protein | [84] |
| At1g14580 (AtIDD6)/BLUEJAY | Stem cell regulator and participant in root initiation, and is required to pattern new organs. | Root development | [79,85] |
| At1g55110 (AtIDD7)/REDSTART | N/R | ||
| At5g44160 (AtIDD8)/NUTCRACKER | idd8 mutants exhibit delayed flowering under LD condition. | Control floral transition via modulation of sugar metabolism, root development | [25,27,79,86] |
| At3g45260 (AtIDD9)/BALDIBIS (Cabib and Leloir) | Jointly stabilizes tissue boundaries by confining the cell fate regulator SHORT-ROOT and contributing to fate specification. | Restrict SHR movement in root tissues | [87] |
| At5g03150 (AtIDD10)/JACKDAW | JKD physically interacts with cell fate determinants SHR and SCR in a cell-type-specific manner. | Root development, DELLA interacting protein | [9,53,54,55,88] |
| At3g13810 (AtIDD11)/WARBLER | N/R | Leaf polarity (regulated by REV and KAN1) | [81] |
| At4g02670 (AtIDD12)/WOODPECKER | N/R | ||
| At5g60470 (AtIDD13)/EGRET | N/R | ||
| At1g68130 (AtIDD14) | idd14-1 mutant shows diverse leaf phenotypes. 35S: IDD14α has retarded growth and downward leaf curling (same 35S: QQS), whereas 35S: IDD14β and idd14-1 mutants show slightly early flowering. | Auxin biosynthesis and transport, starch metabolism in response to cold stress | [6,35] |
| At2g01940 (AtIDD15)/SGR5 | idd15-5 increases angles between inflorescence stem or branches and siliques. | Auxin biosynthesis and transport | [6,80,82] |
| At1g25250 (AtIDD16)/FALCON | IDD16-RNAi transgenic plant has the same phenotype as idd15-5. idd14-1 and IDD16-RNAi transgenic plants have enlarged floral organs and infertile siliques. | Auxin biosynthesis and transport | [6] |
| Rice | |||
| LOC_Os10g28330 (OsID)/SID1/RID1 | rid1 causes the never-flowering phenotype and gain of function of SID1, OsIDD1 or OsIDD6 restore the rid1 phenotype. | Flowering transition | [23,32] |
| LOC_Os01g09850 (OsIDD2) | OsIDD2-overexpression plants showed severe dwarfism with height reaching about half that of the wild-type plants, whereas OsIDD2-RNAi and osidd2 rescued the phenotype. | Secondary cell wall structure | [44] |
| LOC_Os09g38340 (OsIDD3)/ROC1 | roc1 mutant shows hypersensitivity to chilling stress. | Cold response | [36] |
| LOC_Os03g13400 (OsIDD14)/LPA1 | lpa1 mutant causes loose plant architecture, reduces shoot gravitropism. | Shoot gravitropism | [47] |
| LOC_Os04g47860 (OsIDD10) | idd10 mutant roots show hypersensitive to exogenous ammonium | Ammonium uptake and nitrogen metabolism | [10,75,76] |
| Zea mays | |||
| Zm2g011357 (ZmID1) | id1 mutant did not undergo a healthy transition to flowering and remained in a prolonged vegetative growth. | Flowering transition | [4,89,90] |
| Zm2g129261 (ZmIDDveg9)/NKD1 Zm5g884137 (ZmIDD9)/NKD2 | nkd1 and nkd2 mutants are important for differentiation, cell patterning, seed maturation, and resource reserve deposition | Endosperm development | [9,43,91] |
| Domesticated barley | |||
| AMM06558 (DbBLF1) | Overexpression of LEAF1 reduced leaf width, leaf1-1 mutant was in contrast. | Regulate barley leaf size | [48] |
| Solanum tuberosum | |||
| X82328 (StPCP1) | Activate uptake of endogenous sucrose | [5] | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kumar, M.; Le, D.T.; Hwang, S.; Seo, P.J.; Kim, H.U. Role of the INDETERMINATE DOMAIN Genes in Plants. Int. J. Mol. Sci. 2019, 20, 2286. https://doi.org/10.3390/ijms20092286
Kumar M, Le DT, Hwang S, Seo PJ, Kim HU. Role of the INDETERMINATE DOMAIN Genes in Plants. International Journal of Molecular Sciences. 2019; 20(9):2286. https://doi.org/10.3390/ijms20092286
Chicago/Turabian StyleKumar, Manu, Dung Thi Le, Seongbin Hwang, Pil Joon Seo, and Hyun Uk Kim. 2019. "Role of the INDETERMINATE DOMAIN Genes in Plants" International Journal of Molecular Sciences 20, no. 9: 2286. https://doi.org/10.3390/ijms20092286