Proteomic Screening for Prediction and Design of Antimicrobial Peptides with AmpGram
Abstract
1. Introduction
2. Results and Discussion
2.1. Benchmark Analysis of AMP Predictors
2.2. Prediction of Potential AMP Regions and Fragments
3. Materials and Methods
3.1. Datasets
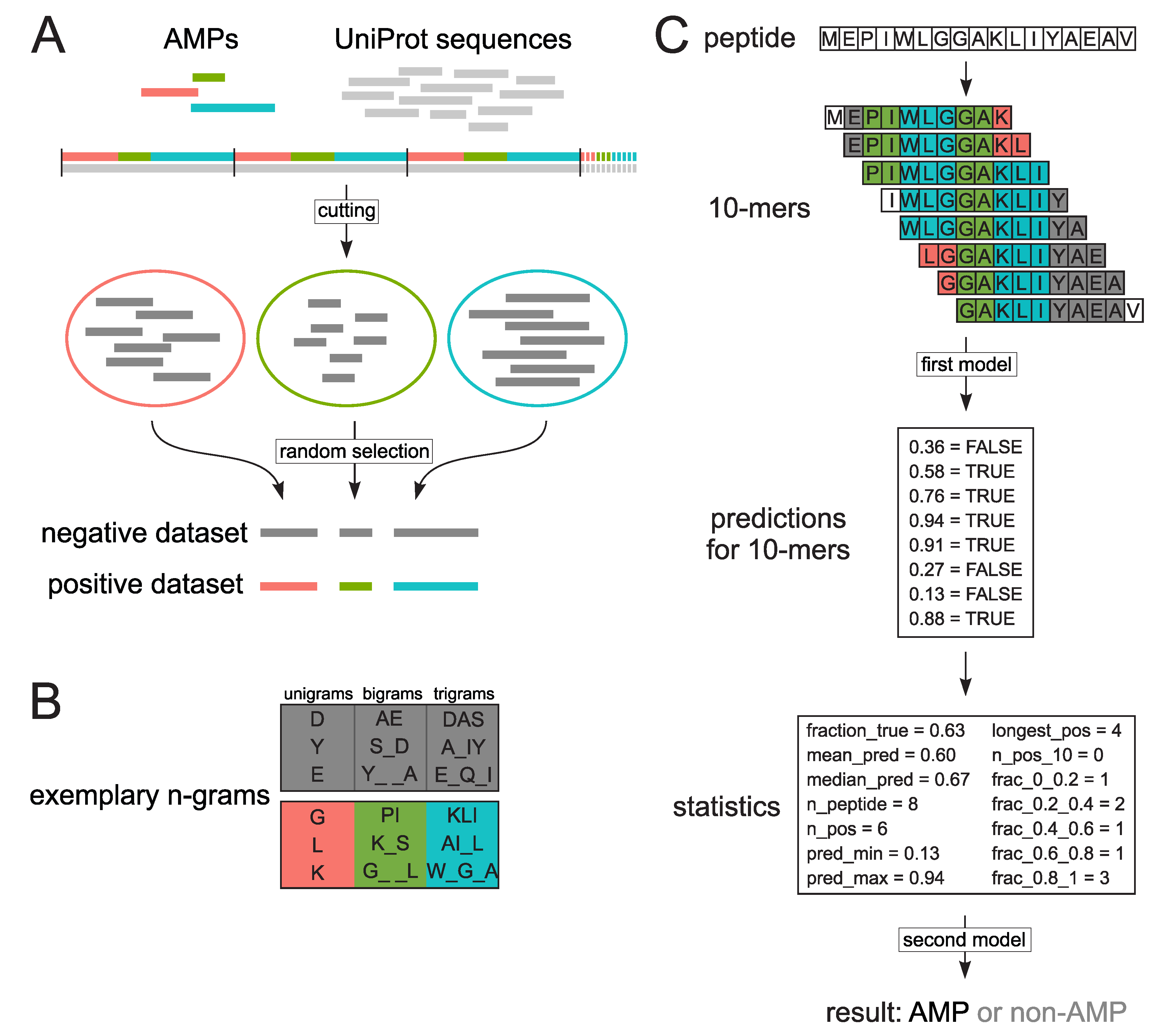
3.2. Extraction of Encoded N-Grams
3.3. Model Training with Random Forests
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| AMP | Anti-microbial peptide |
Appendix A. Availability and Implementation
References
- Laxminarayan, R.; Duse, A.; Wattal, C.; Zaidi, A.K.; Wertheim, H.F.; Sumpradit, N.; Vlieghe, E.; Hara, G.L.; Gould, I.M.; Goossens, H.; et al. Antibiotic resistance—The need for global solutions. Lancet Infect. Dis. 2013, 13, 1057–1098. [Google Scholar] [CrossRef]
- Singh, N.; Abraham, J. Ribosomally synthesized peptides from natural sources. J. Antibiot. 2014, 67, 277–289. [Google Scholar] [CrossRef]
- Cassini, A.; Högberg, L.D.; Plachouras, D.; Quattrocchi, A.; Hoxha, A.; Simonsen, G.S.; Colomb-Cotinat, M.; Kretzschmar, M.E.; Devleesschauwer, B.; Cecchini, M.; et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: A population-level modelling analysis. Lancet Infect. Dis. 2019, 19, 56–66. [Google Scholar] [CrossRef]
- CDC. Antibiotic Resistance Threats in the United States, 2019; Centres for Disease Control and Prevention, US Department of Health and Human Services: Washington, DC, USA, 2019.
- Maróti, G.; Kereszt, A.; Kondorosi, E.; Mergaert, P. Natural roles of antimicrobial peptides in microbes, plants and animals. Res. Microbiol. 2011, 162, 363–374. [Google Scholar] [CrossRef]
- Ahmed, A.; Siman-Tov, G.; Hall, G.; Bhalla, N.; Narayanan, A. Human antimicrobial peptides as therapeutics for viral infections. Viruses 2019, 11, 704. [Google Scholar] [CrossRef]
- Mookherjee, N.; Anderson, M.A.; Haagsman, H.P.; Davidson, D.J. Antimicrobial host defence peptides: Functions and clinical potential. Nat. Rev. Drug Discov. 2020, 19, 311–332. [Google Scholar] [CrossRef]
- Hancock, R.E.; Haney, E.F.; Gill, E.E. The immunology of host defence peptides: Beyond antimicrobial activity. Nat. Rev. Immunol. 2016, 16, 321. [Google Scholar] [CrossRef]
- Mahlapuu, M.; Håkansson, J.; Ringstad, L.; Björn, C. Antimicrobial peptides: An emerging category of therapeutic agents. Front. Cell. Infect. Microbiol. 2016, 6, 194. [Google Scholar] [CrossRef] [PubMed]
- De la Fuente-Núñez, C.; Silva, O.N.; Lu, T.K.; Franco, O.L. Antimicrobial peptides: Role in human disease and potential as immunotherapies. Pharmacol. Ther. 2017, 178, 132–140. [Google Scholar] [CrossRef]
- Schierack, P.; Rödiger, S.; Kuhl, C.; Hiemann, R.; Roggenbuck, D.; Li, G.; Weinreich, J.; Berger, E.; Nolan, L.K.; Nicholson, B.; et al. Porcine E. coli: Virulence-Associated Genes, Resistance Genes and Adhesion and Probiotic Activity Tested by a New Screening Method. PLoS ONE 2013, 8, e59242. [Google Scholar] [CrossRef]
- Raffatellu, M. Learning from bacterial competition in the host to develop antimicrobials. Nat. Med. 2018, 24, 1097–1103. [Google Scholar] [CrossRef]
- Suneja, G.; Nain, S.; Sharma, R. Microbiome: A Source of Novel Bioactive Compounds and Antimicrobial Peptides. In Microbial Diversity in Ecosystem Sustainability and Biotechnological Applications; Springer: Singapore, 2019; pp. 615–630. [Google Scholar]
- Travkova, O.G.; Moehwald, H.; Brezesinski, G. The interaction of antimicrobial peptides with membranes. Adv. Colloid Interface Sci. 2017, 247, 521–532. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, T.A.; Hammami, R. Recent insights into structure–function relationships of antimicrobial peptides. J. Food Biochem. 2019, 43, e12546. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Kizhakkedathu, J.N.; Straus, S.K. Antimicrobial peptides: Diversity, mechanism of action and strategies to improve the activity and biocompatibility in vivo. Biomolecules 2018, 8, 4. [Google Scholar] [CrossRef]
- Li, J.; Koh, J.J.; Liu, S.; Lakshminarayanan, R.; Verma, C.S.; Beuerman, R.W. Membrane active antimicrobial peptides: Translating mechanistic insights to design. Front. Neurosci. 2017, 11, 73. [Google Scholar] [CrossRef] [PubMed]
- Marquette, A.; Bechinger, B. Biophysical investigations elucidating the mechanisms of action of antimicrobial peptides and their synergism. Biomolecules 2018, 8, 18. [Google Scholar] [CrossRef]
- Le, C.F.; Fang, C.M.; Sekaran, S.D. Intracellular targeting mechanisms by antimicrobial peptides. Antimicrob. Agents Chemother. 2017, 61, e02340-16. [Google Scholar] [CrossRef]
- Andersson, D.I.; Hughes, D.; Kubicek-Sutherland, J.Z. Mechanisms and consequences of bacterial resistance to antimicrobial peptides. Drug Resist. Updates 2016, 26, 43–57. [Google Scholar] [CrossRef]
- Lázár, V.; Martins, A.; Spohn, R.; Daruka, L.; Grézal, G.; Fekete, G.; Számel, M.; Jangir, P.K.; Kintses, B.; Csörgő, B.; et al. Antibiotic-resistant bacteria show widespread collateral sensitivity to antimicrobial peptides. Nat. Microbiol. 2018, 3, 718. [Google Scholar] [CrossRef]
- Spohn, R.; Daruka, L.; Lázár, V.; Martins, A.; Vidovics, F.; Grézal, G.; Méhi, O.; Kintses, B.; Számel, M.; Jangir, P.K.; et al. Integrated evolutionary analysis reveals antimicrobial peptides with limited resistance. Nat. Commun. 2019, 10, 1–13. [Google Scholar] [CrossRef]
- Kosikowska, P.; Lesner, A. Antimicrobial peptides (AMPs) as drug candidates: A patent review (2003–2015). Expert Opin. Ther. Patents 2016, 26, 689–702. [Google Scholar] [CrossRef] [PubMed]
- Veltri, D.; Kamath, U.; Shehu, A. Deep learning improves antimicrobial peptide recognition. Bioinformatics 2018, 34, 2740–2747. [Google Scholar] [CrossRef]
- Lee, H.T.; Lee, C.C.; Yang, J.R.; Lai, J.Z.; Chang, K.Y. A large-scale structural classification of antimicrobial peptides. BioMed Res. Int. 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Xiao, X.; Wang, P.; Lin, W.Z.; Jia, J.H.; Chou, K.C. iAMP-2L: A two-level multi-label classifier for identifying antimicrobial peptides and their functional types. Anal. Biochem. 2013, 436, 168–177. [Google Scholar] [CrossRef] [PubMed]
- Waghu, F.H.; Barai, R.S.; Gurung, P.; Idicula-Thomas, S. CAMPR3: A database on sequences, structures and signatures of antimicrobial peptides. Nucleic Acids Res. 2016, 44, D1094–D1097. [Google Scholar] [CrossRef] [PubMed]
- Meher, P.K.; Sahu, T.K.; Saini, V.; Rao, A.R. Predicting antimicrobial peptides with improved accuracy by incorporating the compositional, physico-chemical and structural features into Chou’s general PseAAC. Sci. Rep. 2017, 7, 1–12. [Google Scholar] [CrossRef]
- Gabere, M.N.; Noble, W.S. Empirical comparison of web-based antimicrobial peptide prediction tools. Bioinformatics 2017, 33, 1921–1929. [Google Scholar] [CrossRef]
- Dziuba, B.; Dziuba, M. New milk protein-derived peptides with potential antimicrobial activity: An approach based on bioinformatic studies. Int. J. Mol. Sci. 2014, 15, 14531–14545. [Google Scholar] [CrossRef]
- Park, Y.W.; Nam, M.S. Bioactive peptides in milk and dairy products: A review. Korean J. Food Sci. Anim. Resour. 2015, 35, 831. [Google Scholar] [CrossRef]
- Bruni, N.; Capucchio, M.T.; Biasibetti, E.; Pessione, E.; Cirrincione, S.; Giraudo, L.; Corona, A.; Dosio, F. Antimicrobial activity of lactoferrin-related peptides and applications in human and veterinary medicine. Molecules 2016, 21, 752. [Google Scholar] [CrossRef]
- Mohanty, D.; Jena, R.; Choudhury, P.K.; Pattnaik, R.; Mohapatra, S.; Saini, M.R. Milk derived antimicrobial bioactive peptides: A review. Int. J. Food Prop. 2016, 19, 837–846. [Google Scholar] [CrossRef]
- Primon-Barros, M.; José Macedo, A. Animal venom peptides: Potential for new antimicrobial agents. Curr. Top. Med. Chem. 2017, 17, 1119–1156. [Google Scholar] [CrossRef] [PubMed]
- De Barros, E.; Gonçalves, R.M.; Cardoso, M.H.; Santos, N.C.; Franco, O.L.; Cândido, E.D.S. Snake Venom Cathelicidins as Natural Antimicrobial Peptides. Front. Pharmacol. 2019, 10, 1415. [Google Scholar] [CrossRef] [PubMed]
- Okubo, B.M.; Silva, O.N.; Migliolo, L.; Gomes, D.G.; Porto, W.F.; Batista, C.L.; Ramos, C.S.; Holanda, H.H.; Dias, S.C.; Franco, O.L.; et al. Evaluation of an antimicrobial L-amino acid oxidase and peptide derivatives from Bothropoides mattogrosensis pitviper venom. PLoS ONE 2012, 7, e33639. [Google Scholar] [CrossRef]
- Papareddy, P.; Rydengård, V.; Pasupuleti, M.; Walse, B.; Mörgelin, M.; Chalupka, A.; Malmsten, M.; Schmidtchen, A. Proteolysis of human thrombin generates novel host defense peptides. PLoS Pathog. 2010, 6, e1000857. [Google Scholar] [CrossRef]
- Consortium, U. UniProt: The universal protein knowledgebase. Nucleic Acids Res. 2018, 46, 2699. [Google Scholar]
- Jhong, J.H.; Chi, Y.H.; Li, W.C.; Lin, T.H.; Huang, K.Y.; Lee, T.Y. dbAMP: An integrated resource for exploring antimicrobial peptides with functional activities and physicochemical properties on transcriptome and proteome data. Nucleic Acids Res. 2018, 47, D285–D297. [Google Scholar] [CrossRef]
- Burdukiewicz, M.; Sobczyk, P.; Rödiger, S.; Duda-Madej, A.; Mackiewicz, P.; Kotulska, M. Amyloidogenic motifs revealed by n-gram analysis. Sci. Rep. 2017, 7, 12961. [Google Scholar] [CrossRef]
- Burdukiewicz, M.; Sobczyk, P.; Chilimoniuk, J.; Gagat, P.; Mackiewicz, P. Prediction of Signal Peptides in Proteins from Malaria Parasites. Int. J. Mol. Sci. 2018, 19, 3709. [Google Scholar] [CrossRef]
- Burdukiewicz, M.; Gagat, P.; Jabłoński, S.; Chilimoniuk, J.; Gaworski, M.; Mackiewicz, P.; Marcin, Ł. PhyMet2: A database and toolkit for phylogenetic and metabolic analyses of methanogens. Environ. Microbiol. Rep. 2018, 10, 378–382. [Google Scholar]
- Thennarasu, S.; Nagaraj, R. Specific antimicrobial and hemolytic activities of 18-residue peptides derived from the amino terminal region of the toxin pardaxin. Protein Eng. Des. Sel. 1996, 9, 1219–1224. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Li, X.; Wang, Z. APD3: The antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 2015, 44, D1087–D1093. [Google Scholar] [CrossRef] [PubMed]
- Seshadri Sundararajan, V.; Gabere, M.N.; Pretorius, A.; Adam, S.; Christoffels, A.; Lehväslaiho, M.; Archer, J.A.; Bajic, V.B. DAMPD: A manually curated antimicrobial peptide database. Nucleic Acids Res. 2012, 40, D1108–D1112. [Google Scholar] [CrossRef] [PubMed]
- Lang, M.; Binder, M.; Richter, J.; Schratz, P.; Pfisterer, F.; Coors, S.; Au, Q.; Casalicchio, G.; Kotthoff, L.; Bischl, B. mlr3: A modern object-oriented machine learning framework in R. J. Open Source Softw. 2019. [Google Scholar] [CrossRef]
- Hand, D.J. Measuring Classifier Performance: A Coherent Alternative to the Area under the ROC Curve. Mach. Learn. 2009, 77, 103–123. [Google Scholar] [CrossRef]
- Seyfert, H.M.; Tuckoricz, A.; Interthal, H.; Koczan, D.; Hobom, G. Structure of the bovine lactoferrin-encoding gene and its promoter. Gene 1994, 143, 265–269. [Google Scholar] [CrossRef]
- Hammami, R.; Ben Hamida, J.; Vergoten, G.; Fliss, I. PhytAMP: A database dedicated to antimicrobial plant peptides. Nucleic Acids Res. 2008, 37, D963–D968. [Google Scholar] [CrossRef] [PubMed]
- Fjell, C.D.; Hancock, R.E.; Cherkasov, A. AMPer: A database and an automated discovery tool for antimicrobial peptides. Bioinformatics 2007, 23, 1148–1155. [Google Scholar] [CrossRef] [PubMed]
- Lata, S.; Mishra, N.K.; Raghava, G.P. AntiBP2: Improved version of antibacterial peptide prediction. BMC Bioinform. 2010, 11, S19. [Google Scholar] [CrossRef]
- Hammami, R.; Zouhir, A.; Le Lay, C.; Hamida, J.B.; Fliss, I. BACTIBASE second release: A database and tool platform for bacteriocin characterization. Bmc Microbiol. 2010, 10, 22. [Google Scholar] [CrossRef]
- Zhao, X.; Wu, H.; Lu, H.; Li, G.; Huang, Q. LAMP: A database linking antimicrobial peptides. PLoS ONE 2013, 8, e66557. [Google Scholar] [CrossRef] [PubMed]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef] [PubMed]
- Wollman, F.A. An antimicrobial origin of transit peptides accounts for early endosymbiotic events. Traffic 2016, 17, 1322–1328. [Google Scholar] [CrossRef] [PubMed]
- Wright, M.N.; Ziegler, A. Ranger: A Fast Implementation of Random Forests for High Dimensional Data in C++ and R. J. Stat. Softw. 2017, 77, 1–17. [Google Scholar] [CrossRef]
- Bell, J.; Larson, M.; Kutzler, M.; Bionaz, M.; Löhr, C.V.; Hendrix, D. miRWoods: Enhanced Precursor Detection and Stacked Random Forests for the Sensitive Detection of microRNAs. PLoS Comput. Biol. 2019, 15, e1007309. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.M.; Edwards, M.A.; Li, J.; Yip, C.M.; Deber, C.M. Roles of hydrophobicity and charge distribution of cationic antimicrobial peptides in peptide-membrane interactions. J. Biol. Chem. 2012, 287, 7738–7745. [Google Scholar] [CrossRef] [PubMed]

| Length Range | UniProt | dbAMP |
|---|---|---|
| 0.85 < 10 | 1119 | 508 |
| 11–19 | 1862 | 1894 |
| 0.8520–26 | 1016 | 1634 |
| 27–36 | 2439 | 1779 |
| 0.8537–60 | 9810 | 2049 |
| 61–710 | 482,852 | 4520 |
| 0.85 > 710 | 45,178 | 5 |
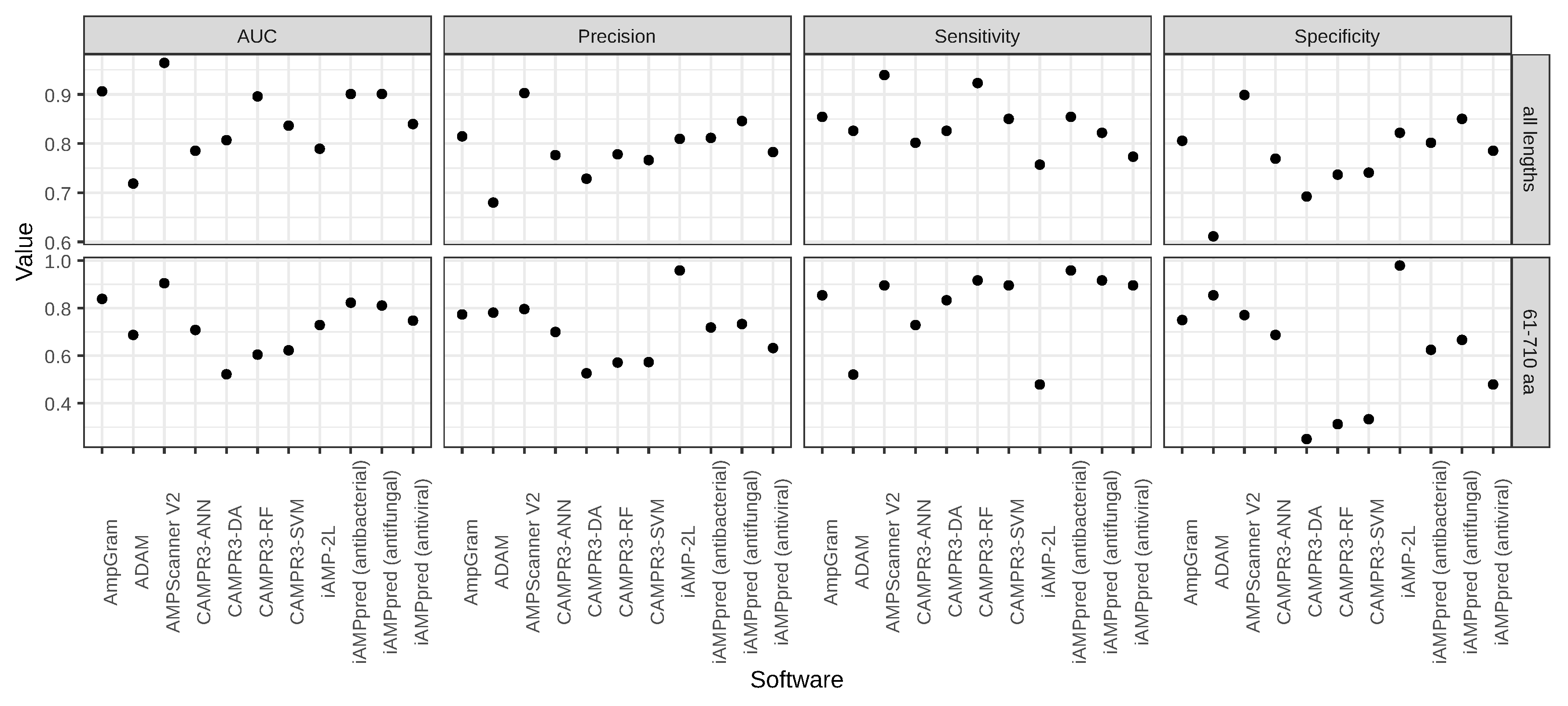
| Software | AUC | Precision | Sensitivity | Specificity |
|---|---|---|---|---|
| AmpGram | 0.9062 | 0.8147 | 0.8543 | 0.8057 |
| ADAM * | 0.7186 | 0.6800 | 0.8259 | 0.6113 |
| 0.85AMPScanner V2 | 0.9641 | 0.9027 | 0.9393 | 0.8988 |
| CAMPR3-ANN * | 0.7854 | 0.7765 | 0.8016 | 0.7692 |
| 0.85CAMPR3-DA | 0.8069 | 0.7286 | 0.8259 | 0.6923 |
| CAMPR3-RF | 0.8958 | 0.7782 | 0.9231 | 0.7368 |
| 0.85CAMPR3-SVM | 0.8363 | 0.7664 | 0.8502 | 0.7409 |
| iAMP-2L * | 0.7895 | 0.8095 | 0.7571 | 0.8219 |
| 0.85iAMPpred (antibacterial) | 0.9008 | 0.8115 | 0.8543 | 0.8016 |
| iAMPpred (antifungal) | 0.9009 | 0.8458 | 0.8219 | 0.8502 |
| 0.85iAMPpred (antiviral) | 0.8397 | 0.7828 | 0.7733 | 0.7854 |
| Software | AUC | Precision | Sensitivity | Specificity |
|---|---|---|---|---|
| AmpGram | 0.8390 | 0.7736 | 0.8542 | 0.7500 |
| ADAM * | 0.6875 | 0.7812 | 0.5208 | 0.8542 |
| AMPScanner V2 | 0.9049 | 0.7963 | 0.8958 | 0.7708 |
| CAMPR3-ANN * | 0.7083 | 0.7000 | 0.7292 | 0.6875 |
| CAMPR3-DA | 0.5221 | 0.5263 | 0.8333 | 0.2500 |
| CAMPR3-RF | 0.6048 | 0.5714 | 0.9167 | 0.3125 |
| CAMPR3-SVM | 0.6228 | 0.5733 | 0.8958 | 0.3333 |
| iAMP-2L * | 0.7292 | 0.9583 | 0.4792 | 0.9792 |
| iAMPpred (antibacterial) | 0.8229 | 0.7188 | 0.9583 | 0.6250 |
| iAMPpred (antifungal) | 0.8110 | 0.7333 | 0.9167 | 0.6667 |
| iAMPpred (antiviral) | 0.7476 | 0.6324 | 0.8958 | 0.4792 |
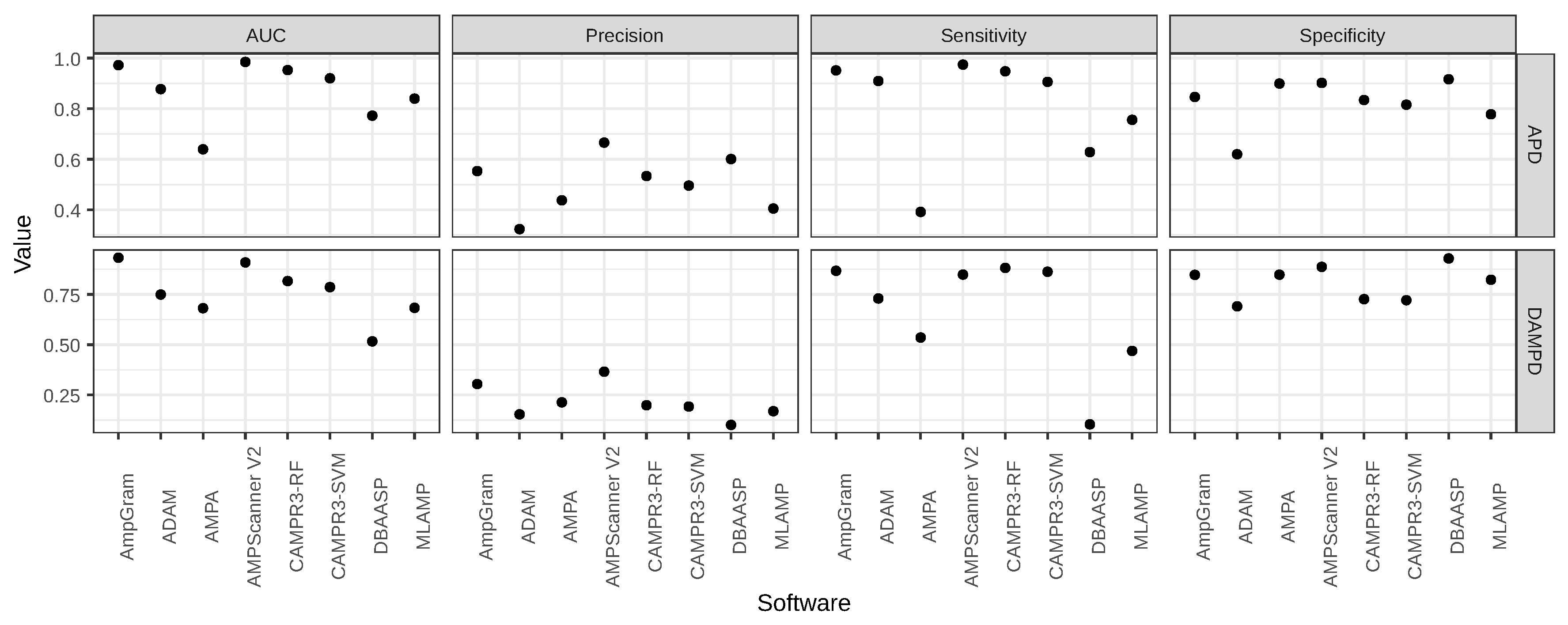
| Software | AUC | Precision | Sensitivity | Specificity |
|---|---|---|---|---|
| AmpGram | 0.9723 | 0.5531 | 0.9515 | 0.8462 |
| ADAM | 0.8774 | 0.3236 | 0.9095 | 0.6198 |
| AMPA | 0.6394 | 0.4377 | 0.3917 | 0.8994 |
| AMPScanner V2 | 0.9848 | 0.6657 | 0.9743 | 0.9022 |
| CAMPR3-RF | 0.9528 | 0.5337 | 0.9480 | 0.8343 |
| CAMPR3-SVM | 0.9202 | 0.4958 | 0.9060 | 0.8158 |
| DBAASP | 0.7723 | 0.6008 | 0.6281 | 0.9165 |
| MLAMP | 0.8397 | 0.4052 | 0.7560 | 0.7781 |
| Software | AUC | Precision | Sensitivity | Specificity |
|---|---|---|---|---|
| AmpGram | 0.9321 | 0.3045 | 0.8673 | 0.8472 |
| ADAM | 0.7494 | 0.1540 | 0.7299 | 0.6907 |
| AMPA | 0.6813 | 0.2136 | 0.5355 | 0.8479 |
| AMPScanner V2 | 0.9088 | 0.3661 | 0.8483 | 0.8867 |
| CAMPR3-RF | 0.8162 | 0.1991 | 0.8815 | 0.7265 |
| CAMPR3-SVM | 0.7862 | 0.1926 | 0.8626 | 0.7210 |
| DBAASP | 0.5165 | 0.1014 | 0.1043 | 0.9287 |
| MLAMP | 0.6833 | 0.1695 | 0.4692 | 0.8227 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Burdukiewicz, M.; Sidorczuk, K.; Rafacz, D.; Pietluch, F.; Chilimoniuk, J.; Rödiger, S.; Gagat, P. Proteomic Screening for Prediction and Design of Antimicrobial Peptides with AmpGram. Int. J. Mol. Sci. 2020, 21, 4310. https://doi.org/10.3390/ijms21124310
Burdukiewicz M, Sidorczuk K, Rafacz D, Pietluch F, Chilimoniuk J, Rödiger S, Gagat P. Proteomic Screening for Prediction and Design of Antimicrobial Peptides with AmpGram. International Journal of Molecular Sciences. 2020; 21(12):4310. https://doi.org/10.3390/ijms21124310
Chicago/Turabian StyleBurdukiewicz, Michał, Katarzyna Sidorczuk, Dominik Rafacz, Filip Pietluch, Jarosław Chilimoniuk, Stefan Rödiger, and Przemysław Gagat. 2020. "Proteomic Screening for Prediction and Design of Antimicrobial Peptides with AmpGram" International Journal of Molecular Sciences 21, no. 12: 4310. https://doi.org/10.3390/ijms21124310
APA StyleBurdukiewicz, M., Sidorczuk, K., Rafacz, D., Pietluch, F., Chilimoniuk, J., Rödiger, S., & Gagat, P. (2020). Proteomic Screening for Prediction and Design of Antimicrobial Peptides with AmpGram. International Journal of Molecular Sciences, 21(12), 4310. https://doi.org/10.3390/ijms21124310





