Identification of Long-Distance Transmissible mRNA between Scion and Rootstock in Cucurbit Seedling Heterografts
Abstract
:1. Introduction
2. Results
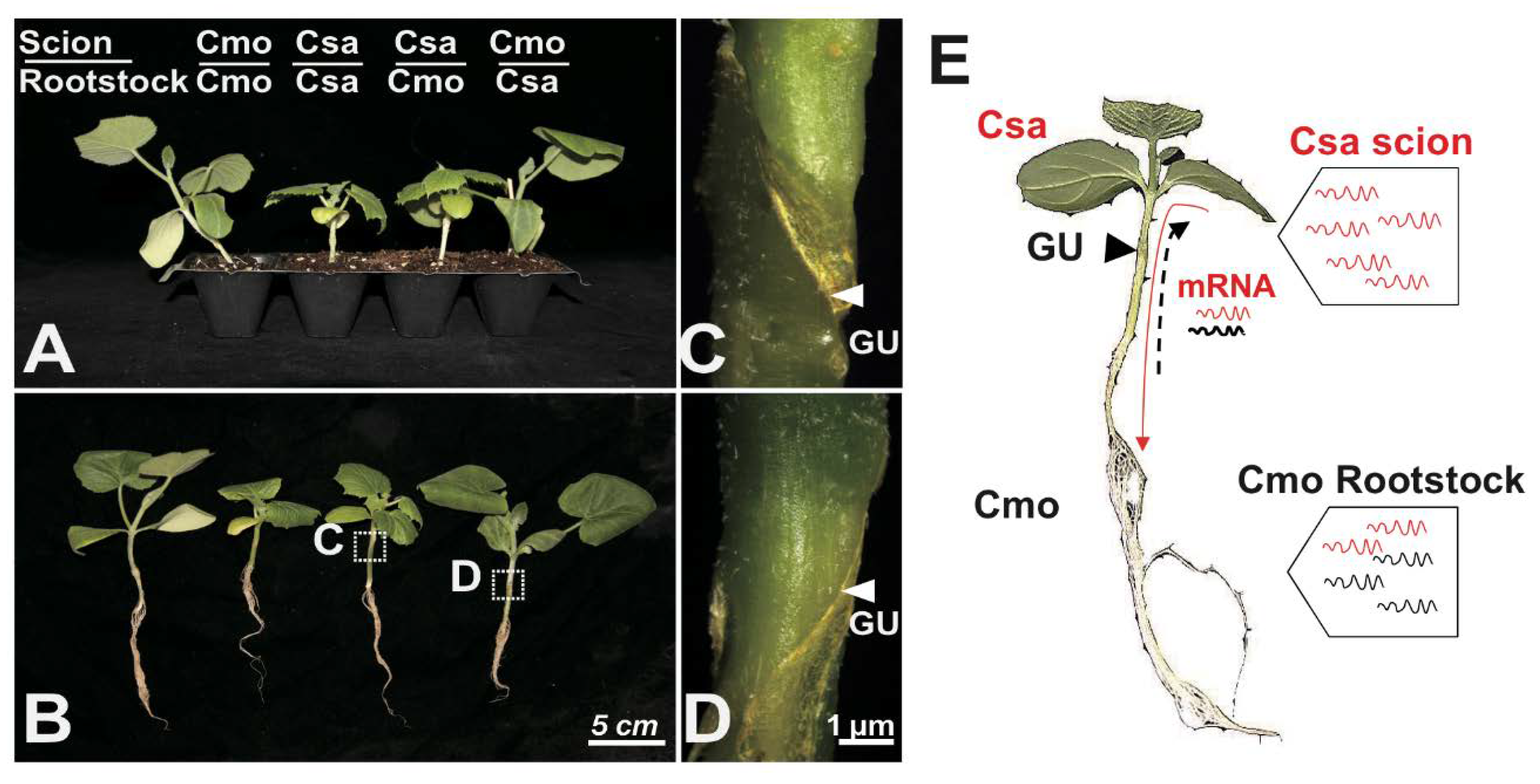
2.1. A High Number of Transcripts Directionally Movement in Grafted Cucumber and Pumpkin Seedlings
2.2. Function Characterization of Mobile mRNAs of Cucumber and Pumpkin Are Universal Common
2.3. Function Characterization of Mobile mRNAs of Cucumber and Pumpkin Are Individually Different
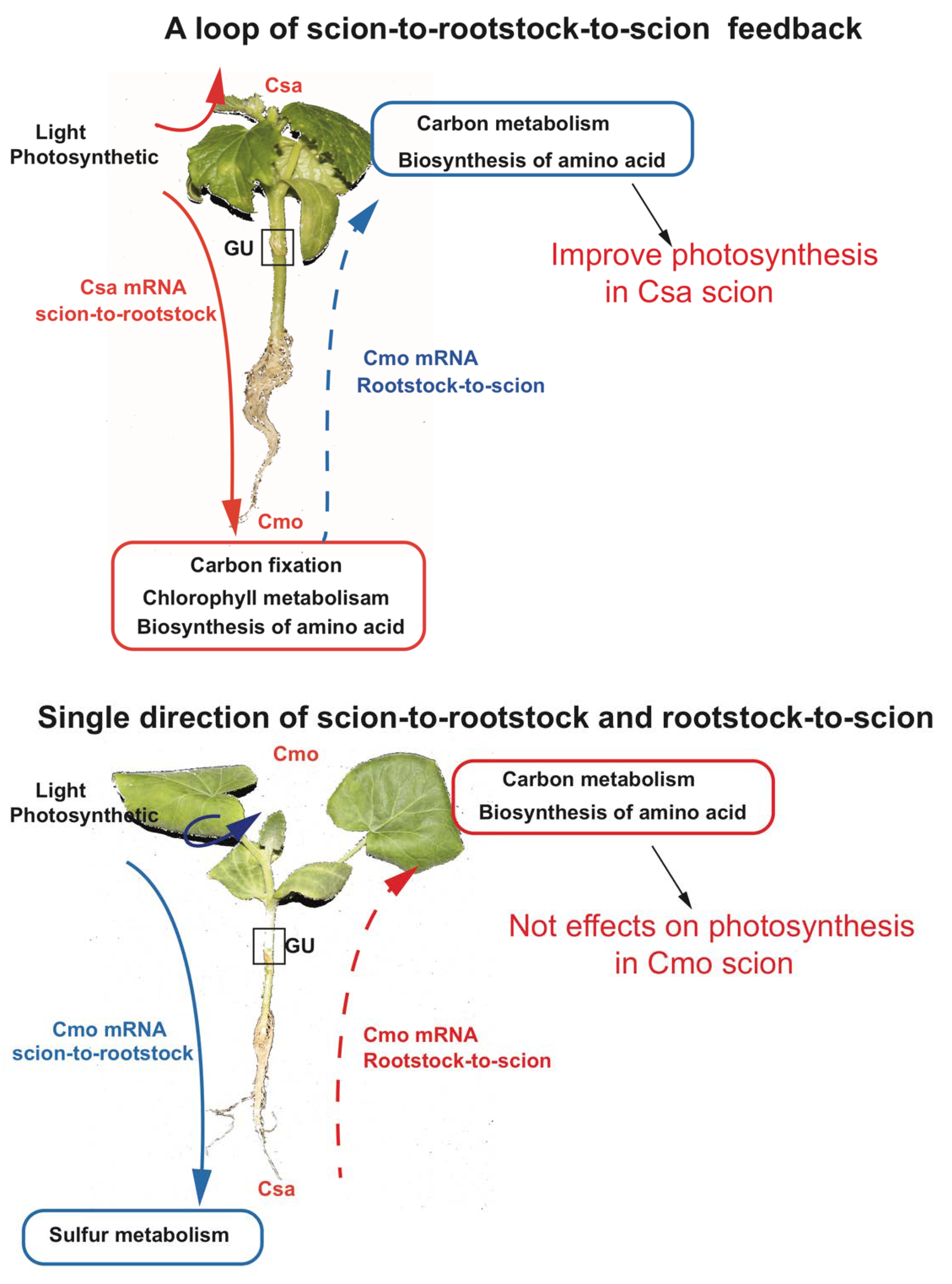
2.4. Identification of Long-Distance Trafficking Between Rootstock and Scion of Csa/Cmo mRNA
3. Discussion
4. Materials and Methods
4.1. Plant Materials, Growth Conditions, and Grafting
4.2. Genomic DNA Isolation and Genomic Purity Verification
4.3. RNA Isolation and cDNA Synthesis
4.4. Construction and Colony Sequencing
4.5. RNA and DNA Library Construction and Sequencing
4.6. Consensus Genome Construction
4.7. Mobile RNA Identification by RNA-seq
4.8. GO (Gene Ontology) Enrichment and KEGG (Kyoto Encyclopedia of Genes and Genomes) Pathway Analysis
4.9. Orthology Analysis
4.10. Quantitative RT-PCR
4.11. Data Statistical Analysis
4.12. Data Availability
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Albacete, A.; Martinez-Andujar, C.; Martinez-Perez, A.; Thompson, A.J.; Dodd, I.C.; Perez-Alfocea, F. Unravelling rootstock × scion interactions to improve food security. J. Exp. Bot. 2015, 66, 2211–2226. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Albacete, A.; Andujar, C.; Perez-Alfocea, F.; Lozano, J.; Asins, M. Rootstock-mediated variation in tomato vegetative growth under low potassium or phosphorous supplies. I International Symposium on Vegetable Grafting, Wuhan, China. Acta. Hortic. 2015, 1086, 147–152. [Google Scholar] [CrossRef]
- Louws, F.J.; Rivard, C.L.; Kubota, C. Grafting fruiting vegetables to manage soilborne pathogens, foliar pathogens, arthropods and weeds. Sci. Hortic. 2010, 127, 127–146. [Google Scholar] [CrossRef]
- Haroldsen, V.M.; Szczerba, M.W.; Aktas, H.; Lopez-Baltazar, J.; Odias, M.J.; Chi-Ham, C.L.; Labavitch, J.M.; Bennett, A.B.; Powell, A.L.T. Mobility of transgenic nucleic acids and proteins within grafted rootstocks for agricultural improvement. Front. Plant. Sci. 2012, 3, 39. [Google Scholar] [CrossRef] [Green Version]
- Yin, L.K.; Zhao, W.C.; Shu, C.; Li, X.M.; Fan, J.W.; Wang, S.H. Role of protective enzymes in tomato rootstocks to resist root knot nematodes. I International Symposium on Vegetable Grafting, Wuhan, China. Acta. Hortic. 2015, 1086, 213–218. [Google Scholar] [CrossRef]
- Turnbull, C.G.N.; Lopez-Cobollo, R.M. Heavy traffic in the fast lane: Long-distance signalling by macromolecules. New Phytol. 2013, 198, 33–51. [Google Scholar] [CrossRef]
- Lucas, W.J.; Yoo, B.C.; Kragler, F. RNA as a long-distance information macromolecule in plants. Nat. Rev. Mol. Cell Bio. 2011, 2, 849–857. [Google Scholar] [CrossRef]
- Lough, T.J.; Lee, R.H.; Emerson, S.J.; Forster, R.L.; Lucas, W.J. Functional analysis of the 5′ untranslated region of potexvirus RNA reveals a role in viral replication and cell-to-cell movement. Virology 2006, 351, 455–465. [Google Scholar] [CrossRef] [Green Version]
- Lough, T.J.; Lucas, W.J. Integrative plant biology: Role of phloem long-distance macromolecular trafficking. Annu. Rev. Plant Biol. 2006, 57, 203–232. [Google Scholar] [CrossRef] [Green Version]
- Ruiz-Medrano, R.; Xoconostle-Cázares, B.; Lucas, W.J. Phloem long-distance transport of CmNACP mRNA: Implications for supracellular regulation in plants. Development 1999, 126, 4405–4419. [Google Scholar]
- Kim, M.; Canio, W.; Kessler, S.; Sinha, N. Developmental changes due to long-distance movement of a homeobox fusion transcript in tomato. Science 2001, 293, 287–289. [Google Scholar] [CrossRef] [PubMed]
- Haywood, V.; Yu, T.S.; Huang, N.C.; Lucas, W.J. Phloem long-distance trafficking of GIBBERELLIC ACID-INSENSITIVE RNA regulates leaf development. Plant J. 2005, 42, 49–68. [Google Scholar] [CrossRef]
- Banerjee, A.K.; Lin, T.; Hannapel, D.J. Untranslated regions of a mobile transcript mediate RNA metabolism. Plant Physiol. 2009, 151, 1831–1843. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ham, B.K.; Brandom, J.L.; Xoconostle-Cazares, B.; Ringgold, V.; Lough, T.; Lucas, W.J. A polypyrimidine tract binding protein, pumpkin RBP50, forms the basis of a phloem-mobile ribonucleoprotein complex. Plant Cell 2009, 21, 197–215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hu, C.Y.; Ham, B.K.; El-Shabrawi, H.M.; Alexander, D.; Zhang, D.B.; Ryals, J.; Lucas, W.J. Proteomics and metabolomics analyses reveal the cucurbit sieve tube system as a complex metabolic space. Plant J. 2016, 87, 442–454. [Google Scholar] [CrossRef]
- Paultre, D.S.G.; Gustin, M.P.; Molnar, A.; Oparka, K.J. Lost in transit: Long-distance trafficking and phloem unloading of protein signals in Arabidopsis homografts. Plant Cell 2016, 28, 2016–2025. [Google Scholar] [CrossRef] [Green Version]
- Kehr, J.; Kragler, F. Long distance RNA movement. New Phytol. 2018, 218, 29–40. [Google Scholar] [CrossRef] [Green Version]
- Dreher, T.W.; Rao, A.L.N.; Hall, T.C. Replication in vivo of mutant brome mosaic virus RNAs defective in aminoacylation. J. Mol. Biol. 1989, 206, 425–438. [Google Scholar] [CrossRef]
- Fechter, P.; Rudinger-Thirion, J.; Florentz, C.; Giegé, R. Novel features in the tRNA-like world of plant viral RNAs. Cell. Mol. Life Sci. 2001, 58, 1547–1561. [Google Scholar] [CrossRef]
- Barends, S.; Rudinger-Thirion, J.; Florentz, C.; Giege, R.; Pleij, C.W.A.; Kraal, B. tRNA-like structure regulates translation of Brome mosaic virus RNA. J. Virol. 2004, 78, 4003–4010. [Google Scholar] [CrossRef] [Green Version]
- Yang, L.; Perrera, V.; Saplaoura, E.; Apelt, F.; Bahin, M.; Kramdi, A.; Olas, J.; Mueller-Roeber, B.; Sokolowska, E.; Zhang, W.N. m5C Methylation Guides Systemic Transport of Messenger RNA over Graft Junctions in Plants. Curr. Biol. 2019, 29, 2465–2476. [Google Scholar] [CrossRef] [Green Version]
- Ding, B. The biology of viroid-host interactions. Annu. Rev. Phytopathol. 2009, 47, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.; LeBlanc, M.L.; Wafula, E.K.; de Pamphilis, C.W.; Westwood, J.H. Genomic-scale exchange of mRNA between a parasitic plant and its hosts. Science 2014, 345, 808–811. [Google Scholar] [CrossRef] [PubMed]
- Notaguchi, M.; Higashiyama, T.; Suzuki, T. Identification of mRNAs that move over long distances using an RNA-Seq analysis of Arabidopsis/Nicotiana benthamiana heterografts. Plant Cell Physiol. 2015, 56, 311–321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thieme, C.J.; Rojas-Triana, M.; Stecyk, E.; Schudoma, C.; Zhang, W.N.; Yang, L.; Minambres, M.; Walther, D.; Schulze, W.X.; Paz-Ares, J.; et al. Endogenous Arabidopsis messenger RNAs transported to distant tissues. Nat. Plants 2015, 1, 15025. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.Z.; Mao, L.Y.; Jittayasothorn, Y.; Kang, Y.M.; Jiao, C.; Fei, Z.J.; Zhong, G.Y. Messenger RNA exchange between scions and rootstocks in grafted grapevines. BMC Plant Biol. 2015, 15, 251. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Z.L.; Zheng, Y.; Ham, B.K.; Chen, J.Y.; Yoshida, A.; Kochian, L.V.; Fei, Z.J.; Lucas, W.J. Vascular-mediated signalling involved in early phosphate stress response in plants. Nat. Plants 2016, 2, 16033. [Google Scholar] [CrossRef]
- Xia, C.; Zheng, Y.; Huang, J.; Zhou, X.J.; Li, R.; Zha, M.R.; Wang, S.J.; Huang, Z.Q.; Lan, H.; Turgeon, R.; et al. Elucidation of the mechanisms of long-distance mRNA movement in a Nicotiana benthamiana/Tomato heterograft system. Plant Physiol. 2018, 177, 745–758. [Google Scholar] [CrossRef] [Green Version]
- Omid, A.; Keilin, T.; Glass, A.; Leshkowitz, D.; Wolf, S. Characterization of phloem-sap transcription profile in melon plants. J. Exp. Bot. 2007, 58, 3645–3656. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.; Bie, Z.L.; Huang, Y.; Han, X.Y. Effect of grafting on the growth and ion concentrations of cucumber seedlings under NaCl stress (Plant nutrition). Soil Sci. Plant Nutr. 2008, 54, 895–902. [Google Scholar] [CrossRef]
- Lee, J.M.; Kubota, C.; Tsao, S.J.; Bie, Z.L.; Echevarria, P.H.; Morra, L.; Oda, M. Current status of vegetable grafting: Diffusion, grafting techniques, automation. Sci. Hortic. 2010, 127, 93–105. [Google Scholar] [CrossRef]
- Li, H.; Liu, S.S.; Yi, C.Y.; Wang, F.; Zhou, J.; Xia, X.J.; Shi, K.; Zhao, Y.H.; Yu, J.Q. Hydrogen peroxide mediates abscisic acid-induced HSP 70 accumulation and heat tolerance in grafted cucumber plants. Plant Cell Environ. 2014, 37, 2768–2780. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Guo, S.R.; Li, H.; Sun, H.Z.; Lu, N.; Shu, S.; Sun, J. Resistance of Cucumber Grafting Rootstock Pumpkin Cultivars to Chilling and Salinity Stresses. Korean J. Hortic. Sci. Technol. 2017, 35, 220–231. [Google Scholar] [CrossRef]
- Huang, Y.; Bie, Z.L.; Liu, P.; Niu, M.L.; Zhen, A.; Liu, Z.X.; Lei, B.; Gu, D.J.; Lu, C.; Wang, B.T. Reciprocal grafting between cucumber and pumpkin demonstrates the roles of the rootstock in the determination of cucumber salt tolerance and sodium accumulation. Sci. Hortic. 2012, 149, 47–54. [Google Scholar] [CrossRef]
- Xing, W.W.; Li, L.; Gao, P.; Li, H.; Shao, Q.S.; Shu, S.; Sun, J.; Guo, S.R. Effects of grafting with pumpkin rootstock on carbohydrate metabolism in cucumber seedlings under Ca(NO3)2 stress. Plant Physiol. Biochem. 2015, 84, 124–132. [Google Scholar] [CrossRef] [PubMed]
- Niu, M.L.; Sun, S.T.; Nawaz, M.A.; Sun, J.Y.; Cao, H.S.; Lu, J.Y.; Huang, Y.; Bie, Z.L. Grafting Cucumber Onto Pumpkin Induced Early Stomatal Closure by Increasing ABA Sensitivity Under Salinity Conditions. Front. Plant Sci. 2019, 10, 1290. [Google Scholar] [CrossRef]
- Banerjee, A.K.; Chatterjee, M.; Yu, Y.Y.; Suh, S.G.; Miller, W.A.; Hannapel, D.J. Dynamics of a mobile RNA of potato involved in a long-distance signaling pathway. Plant Cell 2006, 18, 3443–3457. [Google Scholar] [CrossRef] [Green Version]
- Ham, B.K.; Lucas, W.J. Phloem-Mobile RNAs as Systemic Signaling Agents. Annu. Rev. Plant Biol. 2017, 68, 173–195. [Google Scholar] [CrossRef]
- Warschefsky, E.J.; Klein, L.L.; Frank, M.H.; Chitwood, D.H.; Londo, J.P.; von Wettberg, E.J.B.; Miller, A.J. Rootstocks: Diversity, domestication, and impacts on shoot phenotypes. Trends Plant Sci. 2016, 21, 418–437. [Google Scholar] [CrossRef]
- Lu, X.H.; Liu, W.Q.; Wang, T.; Zhang, J.L.; Li, X.J.; Zhang, W.N. Systemic long-distance signaling and communication between the rootstock and scion in grafted vegetables. Front. Plant Sci. 2020, 11, 460. [Google Scholar] [CrossRef]
- Notaguchi, M.; Okamoto, S. Dynamics of long-distance signaling via plant vascular tissues. Front. Plant Sci. 2015, 6, 161. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gaion, L.A.; Carvalho, R.F. Long-Distance Signaling: What Grafting has Revealed? J. Plant. Growth Regul. 2018, 37, 694–704. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, F.; Shinozaki, K. Long-distance signaling in plant stress response. Curr. Opin. Plant Biol. 2019, 47, 106–111. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, L.P.; Xing, N.L.; Wu, X.H.; Wu, X.Y.; Wang, B.G.; Lu, Z.F.; Xu, P.; Tao, Y.; Li, G.J.; et al. A universal pipeline for mobile mRNA detection and insights into heterografting advantages under chilling stress. Hortic Res. 2020, 7, 13. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence alignment/map (SAM) format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [Green Version]
- Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatica 2011, 27, 2987–2993. [Google Scholar] [CrossRef] [Green Version]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [Green Version]
- Shen, W.; Le, S.; Li, Y.; Hu, F. SeqKit: A Cross-Platform and Ultrafast Toolkit for FASTA/Q File Manipulation. PLoS ONE 2016, 11, e0163962. [Google Scholar] [CrossRef]










© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, W.; Xiang, C.; Li, X.; Wang, T.; Lu, X.; Liu, Z.; Gao, L.; Zhang, W. Identification of Long-Distance Transmissible mRNA between Scion and Rootstock in Cucurbit Seedling Heterografts. Int. J. Mol. Sci. 2020, 21, 5253. https://doi.org/10.3390/ijms21155253
Liu W, Xiang C, Li X, Wang T, Lu X, Liu Z, Gao L, Zhang W. Identification of Long-Distance Transmissible mRNA between Scion and Rootstock in Cucurbit Seedling Heterografts. International Journal of Molecular Sciences. 2020; 21(15):5253. https://doi.org/10.3390/ijms21155253
Chicago/Turabian StyleLiu, Wenqian, Chenggang Xiang, Xiaojun Li, Tao Wang, Xiaohong Lu, Zixi Liu, Lihong Gao, and Wenna Zhang. 2020. "Identification of Long-Distance Transmissible mRNA between Scion and Rootstock in Cucurbit Seedling Heterografts" International Journal of Molecular Sciences 21, no. 15: 5253. https://doi.org/10.3390/ijms21155253




