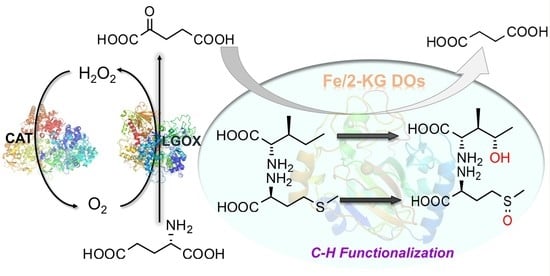
2-Ketoglutarate-Generated In Vitro Enzymatic Biosystem Facilitates Fe(II)/2-Ketoglutarate-Dependent Dioxygenase-Mediated C–H Bond Oxidation for (2s,3r,4s)-4-Hydroxyisoleucine Synthesis
Abstract
:1. Introduction
2. Results and Discussion
2.1. Design of a Fe(II)/2-KG DO-Based In Vitro Enzymatic Biosystem for Hydroxy Amino Acid Synthesis
2.2. Enzyme Kinetics
2.3. Effects of Reaction Components on IDO and LGOX Activity
2.4. Enzymatic Cascade for 4-HIL Synthesis
2.5. Fed-Batch 4-HIL Synthesis Reaction
2.6. In Vitro Enzymatic Biosystem for Hydroxy Amino Acids Synthesis
3. Materials and Methods
3.1. Reagents
3.2. Preparation of Recombinant Enzymes
3.3. Enzymatic Activity Assays
3.4. Determination of Km and Vmax
3.5. Effects of Reaction Components on Enzyme Activities
3.6. One-Pot Synthesis of 4-HIL
3.7. Two-Step Process for Hydroxy Amino Acid Synthesis
3.8. Fed-Batch Reaction for 4-HIL Synthesis
3.9. Analysis of Organic Acids
3.10. Analysis of Amino Acids
3.11. Nuclear Magnetic Resonance (NMR) Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Newhouse, T.; Baran, P.S. If C-H bonds could talk: Selective C-H Bond oxidation. Angew. Chem. Int. Ed. Engl. 2011, 50, 3362–3374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davies, H.M.L.; Morton, D. Recent advances in C-H functionalization. J. Org. Chem. 2016, 81, 343–350. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qiu, Y.; Gao, S. Trends in applying C-H oxidation to the total synthesis of natural products. Nat. Prod. Rep. 2016, 33, 562–581. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Fernandez-Fueyo, E.; Hollmann, F.; Paul, C.E.; Pesic, M.; Schmidt, S.; Wang, Y.; Younes, S.; Zhang, W. Biocatalytic oxidation reactions: A chemist’s perspective. Angew. Chem. Int. Ed. Engl. 2018, 57, 9238–9261. [Google Scholar] [CrossRef] [PubMed]
- Martinez, S.; Hausinger, R.P. Catalytic mechanisms of Fe(II)- and 2-oxoglutarate-dependent oxygenases. J. Biol. Chem. 2015, 290, 20702–20711. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jia, B.; Jia, X.; Kim, K.H.; Jeon, C.O. Integrative view of 2-oxoglutarate/Fe(II)-dependent oxygenase diversity and functions in bacteria. Biochim. Biophys. Act. Gen. Subj. 2017, 1861, 323–334. [Google Scholar] [CrossRef]
- Hara, R.; Yamagata, K.; Miyake, R.; Kawabata, H.; Uehara, H.; Kino, K. Discovery of Lysine hydroxylases in the clavaminic acid synthase-like superfamily for efficient hydroxylysine bioproduction. Appl. Environ. Microbiol. 2017, 83, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Strieker, M.; Kopp, F.; Mahlert, C.; Essen, L.-O.; Marahiel, M.A. Mechanistic and structural basis of stereospecific Cβ-hydroxylation in calcium-dependent antibiotic, a daptomycin-type lipopeptide. ACS Chem. Biol. 2007, 2, 187–196. [Google Scholar] [CrossRef]
- Kodera, T.; Smirnov, S.V.; Samsonova, N.N.; Kozlov, Y.I.; Koyama, R.; Hibi, M.; Ogawa, J.; Yo9kozeki, K.; Shimizu, S. A novel L-isoleucine hydroxylating enzyme, L-isoleucine dioxygenase from Bacillus thuringiensis, produces (2S,3R,4S)-4-hydroxyisoleucine. Biochem. Biophys. Res. Commun. 2009, 390, 506–510. [Google Scholar] [CrossRef] [PubMed]
- Lukat, P.; Katsuyama, Y.; Wenzel, S.; Binz, T.; König, C.; Blankenfeldt, W.; Brönstrup, M.; Müller, R. Biosynthesis of methyl-proline containing griselimycins, natural products with anti-tuberculosis activity. Chem. Sci. 2017, 8, 7521–7527. [Google Scholar] [CrossRef] [Green Version]
- Shibasaki, T.; Mori, H.; Chiba, S.; Ozaki, A. Microbial proline 4-hydroxylase screening and gene cloning. Appl. Environ. Microbiol. 1999, 65, 4028–4031. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koketsu, K.; Shomura, Y.; Moriwaki, K.; Hayashi, M.; Mitsuhashi, S.; Hara, R.; Kino, K.; Higuchi, Y. Refined regio- and stereoselective hydroxylation of L-pipecolic acid by protein engineering of L-proline cis-4-hydroxylase based on the X-ray crystal structure. ACS Synth. Biol. 2015, 4, 383–392. [Google Scholar] [CrossRef]
- Hibi, M.; Kasahara, T.; Kawashima, T.; Yajima, H.; Kozono, S.; Smirnov, S.V.; Kodera, T.; Sugiyama, M.; Shimizu, S.; Yokozeki, K. Multi-enzymatic synthesis of optically pure β-Hydroxy α-amino acids. Adv. Synth. Catal. 2015, 354, 741–745. [Google Scholar] [CrossRef]
- Li, P.; Wu, G. Roles of dietary glycine, proline, and hydroxyproline in collagen synthesis and animal growth. Amino Acids 2018, 50, 29–38. [Google Scholar] [CrossRef]
- Reddy, V.P.; Kumar, A.V.; Rao, K.R. New strategy for the synthesis of N-aryl pyrroles: Cu-catalyzed C–N cross-coupling reaction of trans-4-hydroxy-L-proline with aryl halides. Tetrahedron Lett. 2011, 52, 777–780. [Google Scholar] [CrossRef]
- Poisson, J.-F.; Orellana, A.; Greene, A.E. Stereocontrolled synthesis of (−)-kainic acid from trans-4-hydroxy-L-proline. J. Org. Chem. 2005, 70, 10860–10863. [Google Scholar] [CrossRef]
- Zafar, M.I.; Gao, F. 4-Hydroxyisoleucine: A potential new treatment for type 2 diabetes mellitus. BioDrugs 2016, 30, 255–262. [Google Scholar] [CrossRef]
- Fowden, L. Isolation of γ-hydroxynorvaline from Lathyrus odoratus Seed. Nature 1966, 209, 807–808. [Google Scholar] [CrossRef]
- Broca, C.; Manteghetti, M.; Gross, R.; Baissac, Y.; Jacob, M.; Petit, P.; Sauvaire, Y.; Ribes, G. 4-Hydroxyisoleucine: Effects of synthetic and natural analogues on insulin secretion. Eur. J. Pharmacol. 2000, 390, 339–345. [Google Scholar] [CrossRef]
- Smirnov, S.V.; Kodera, T.; Samsonova, N.N.; Kotlyarovа, V.А.; Rushkevich, N.Y.; Kivero, A.D.; Sokolov, P.M.; Hibi, M.; Ogawa, J.; Shimizu, S. Metabolic engineering of Escherichia coli to produce (2S, 3R, 4S)-4-hydroxyisoleucine. Appl. Microbiol. Biotechnol. 2010, 88, 719–726. [Google Scholar] [CrossRef]
- Zhang, C.; Li, Y.; Ma, J.; Liu, Y.; He, J.; Li, Y.; Zhu, F.; Meng, J.; Zhan, J.; Li, Z.; et al. High production of 4-hydroxyisoleucine in Corynebacterium glutamicum by multistep metabolic engineering. Metab. Eng. 2018, 49, 287–298. [Google Scholar] [CrossRef] [PubMed]
- Baixue, L.; Keqiang, F.; Jian, Z.; Junjie, J.; Linjun, W.; Keqian, Y.; Yong, T. Reconstitution of TCA cycle with DAOCS to engineer Escherichia coli into an efficient whole cell catalyst of penicillin G. Proc. Natl. Acad. Sci. USA 2015, 112, 9855–9859. [Google Scholar]
- Liu, L.; Hossain, G.S.; Shin, H.; Li, J.; Du, G.; Chen, J. One-step production of α-ketoglutaric acid from glutamic acid with an engineered L-amino acid deaminase from Proteus mirabilis. J. Biotechnol. 2013, 164, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Hossain, G.S.; Li, J.; Shin, H.; Chen, R.R.; Du, G.; Liu, L.; Chen, J. Bioconversion of L-glutamic acid to α-ketoglutaric acid by an immobilized whole-cell biocatalyst expressing L-amino acid deaminase from Proteus mirabilis. J. Biotechnol. 2014, 169, 112–120. [Google Scholar] [CrossRef]
- Niu, P.; Dong, X.; Wang, Y.; Liu, L. Enzymatic production of α-ketoglutaric acid from L-glutamic acid via L-glutamate oxidase. J. Biotechnol. 2014, 179, 56–62. [Google Scholar] [CrossRef]
- Wu, J.; Fan, X.; Liu, J.; Luo, Q.; Xu, J.; Chen, X. Promoter engineering of cascade biocatalysis for α-ketoglutaric acid production by coexpressing L-glutamate oxidase and catalase. Appl. Microbiol. Biotechnol. 2018, 102, 4755–4764. [Google Scholar] [CrossRef]
- Schrittwieser, J.H.; Velikogne, S.; Hall, M.; Kroutil, W. Artificial biocatalytic linear cascades for preparation of organic molecules. Chem. Rev. 2017, 118, 270–348. [Google Scholar] [CrossRef] [Green Version]
- Hibi, M.; Kawashima, T.; Kodera, T.; Smirnov, S.V.; Sokolov, P.M.; Sugiyama, M.; Shimizu, S.; Yokozeki, K.; Ogawa, J. Characterization of Bacillus thuringiensis L-isoleucine dioxygenase for production of useful amino acids. Appl. Environ. Microbiol. 2011, 77, 6926–6930. [Google Scholar] [CrossRef] [Green Version]
- Hausinger, R.P. Fe(II)/α-ketoglutarate-dependent hydroxylases and related Enzymes. Crit. Rev. Biochem. Mol. Biol. 2004, 39, 21–68. [Google Scholar] [CrossRef]
- Guterl, J.K.; Garbe, D.; Carsten, J.; Steffler, F.; Sommer, B.; Reiße, S.; Philipp, A.; Haack, M.; Rühmann, B.; Koltermann, A. Cell-free metabolic engineering: Production of chemicals by minimized reaction cascades. ChemSusChem 2012, 5, 2165–2172. [Google Scholar] [CrossRef]
- Qiao, Z.; Xu, M.; Shao, M.; Zhao, Y.; Long, M.; Yang, T.; Zhang, X.; Yang, S.; Nakanishi, H.; Rao, Z. Engineered disulfide bonds improve thermostability and activity of L-isoleucine hydroxylase for efficient 4-HIL production in Bacillus subtilis 168. Eng. Life Sci. 2019, 20, 7–16. [Google Scholar] [CrossRef]
- Hold, C.; Billerbeck, S.; Panke, S. Forward design of a complex enzyme cascade reaction. Nat. Commun. 2016, 7, 12971. [Google Scholar] [CrossRef] [Green Version]
- Beer, B.; Pick, A.; Sieber, V. In vitro metabolic engineering for the production of α-ketoglutarate. Metab. Eng. 2017, 40, 5–13. [Google Scholar] [CrossRef]
- Rose, N.R.; Mcdonough, M.A.; King, O.N.; Kawamura, A.; Schofield, C.J. Inhibition of 2-oxoglutarate dependent oxygenases. Chem. Soc. Rev. 2011, 40, 4364–4397. [Google Scholar] [CrossRef]
- Hara, R.; Kino, K. Characterization of novel 2-oxoglutarate dependent dioxygenases converting L-proline to cis-4-hydroxy-l-proline. Biochem. Biophys. Res. Commun. 2009, 379, 882–886. [Google Scholar] [CrossRef]
- Kusakabe, H.; Midorikawa, Y.; Fujishima, T.; Kuninaka, A.; Yoshino, H. Purification and properties of a new enzyme, L-glutamate oxidase, from Streptomyces sp. X-119-6 grown on wheat bran. Agric. Biol. Chem. 1983, 47, 1323–1328. [Google Scholar] [CrossRef]
- Ferreira, L.A.; Pinho, S.P.; Macedo, E.A. Solubility of L-serine, L-threonine and L-isoleucine in aqueous aliphatic alcohol solutions. Fluid Phase Equilib. 2008, 270, 1–9. [Google Scholar] [CrossRef]
- Shi, F.; Niu, T.; Fang, H. 4-Hydroxyisoleucine production of recombinant Corynebacterium glutamicum ssp. lactofermentumunder optimal corn steep liquor limitation. Appl. Microbiol. Biotechnol. 2015, 99, 3851–3863. [Google Scholar] [CrossRef]
- Jing, X.; Wang, X.; Zhang, W.; An, J.; Luo, P.; Nie, Y.; Xu, Y. Highly regioselective and stereoselective hydroxylation of free amino acids by a 2-oxoglutarate-dependent dioxygenase from Kutzneria albida. ACS Omega 2019, 4, 8350–8358. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Nie, Y.; Xu, Y. Improvement of the activity and stability of starch-debranching pullulanase from Bacillus naganoensis via tailoring of the active sites lining the catalytic pocket. J. Agric. Food Chem. 2018, 66, 13236–13242. [Google Scholar] [CrossRef]
- Xiao, R.; Anderson, S.; Aramini, J.; Belote, R.; Buchwald, W.A.; Ciccosanti, C.; Conover, K.; Everett, J.K.; Hamilton, K.; Huang, Y.J. The high-throughput protein sample production platform of the northeast structural genomics consortium. J. Struct. Biol. 2010, 172, 21–33. [Google Scholar] [CrossRef] [Green Version]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Falcioni, F.; Blank, L.M.; Frick, O.; Karau, A.; Bühler, B.; Schmid, A. Proline availability regulates proline-4-hydroxylase synthesis and substrate uptake in proline-hydroxylating recombinant Escherichia coli. Appl. Environ. Microbiol. 2013, 79, 3091–3100. [Google Scholar] [CrossRef] [Green Version]
- Herbert, P.; Santos, L.; Alves, A. Simultaneous quantification of primary, secondary amino acids, and biogenic amines in musts and wines using OPA/3-MPA/FMOC-CI fluorescent derivatives. J. Food Sci. 2010, 66, 1319–1325. [Google Scholar] [CrossRef]
- Sheiner, L.B.; Beal, S.L. Evaluation of methods for estimating population pharmacokinetic parameters. I. Michaelis-menten model: Routine clinical pharmacokinetic data. J. Pharmacokinet. Biopharm. 1980, 8, 553–571. [Google Scholar] [CrossRef]
- Mori, H.; Shibasaki, T.; Uozaki, Y.; Ochiai, K.; Ozaki, A. Detection of novel proline 3-hydroxylase activities in Streptomyces and Bacillus spp. by regio- and stereospecific hydroxylation of L-proline. Appl. Environ. Microbiol. 1996, 62, 1903–1907. [Google Scholar] [CrossRef] [Green Version]







| Enzyme | Substrate | Km (mM) | Vmax (U·mg−1) | kcat (s−1) | kcat/Km (s−1·m·M−1) |
|---|---|---|---|---|---|
| IDO | l-Ile | 6.34 ± 0.12 | 8.99 ± 0.09 | 4.18 ± 0.08 | 0.66 ± 0.04 |
| 2-KG | 15.12 ± 0.08 | 19.79 ± 0.08 | 9.18 ± 0.07 | 0.61 ± 0.07 | |
| LGOX | l-Glu | 2.65 ± 0.11 | 2.93 ± 0.06 | 3.38 ± 0.05 | 1.28 ± 0.07 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jing, X.-R.; Liu, H.; Nie, Y.; Xu, Y. 2-Ketoglutarate-Generated In Vitro Enzymatic Biosystem Facilitates Fe(II)/2-Ketoglutarate-Dependent Dioxygenase-Mediated C–H Bond Oxidation for (2s,3r,4s)-4-Hydroxyisoleucine Synthesis. Int. J. Mol. Sci. 2020, 21, 5347. https://doi.org/10.3390/ijms21155347
Jing X-R, Liu H, Nie Y, Xu Y. 2-Ketoglutarate-Generated In Vitro Enzymatic Biosystem Facilitates Fe(II)/2-Ketoglutarate-Dependent Dioxygenase-Mediated C–H Bond Oxidation for (2s,3r,4s)-4-Hydroxyisoleucine Synthesis. International Journal of Molecular Sciences. 2020; 21(15):5347. https://doi.org/10.3390/ijms21155347
Chicago/Turabian StyleJing, Xiao-Ran, Huan Liu, Yao Nie, and Yan Xu. 2020. "2-Ketoglutarate-Generated In Vitro Enzymatic Biosystem Facilitates Fe(II)/2-Ketoglutarate-Dependent Dioxygenase-Mediated C–H Bond Oxidation for (2s,3r,4s)-4-Hydroxyisoleucine Synthesis" International Journal of Molecular Sciences 21, no. 15: 5347. https://doi.org/10.3390/ijms21155347