Subchromosome-Scale Nuclear and Complete Mitochondrial Genome Characteristics of Morchella crassipes
Abstract
:1. Introduction
2. Results
2.1. Genomic De Novo Assembly
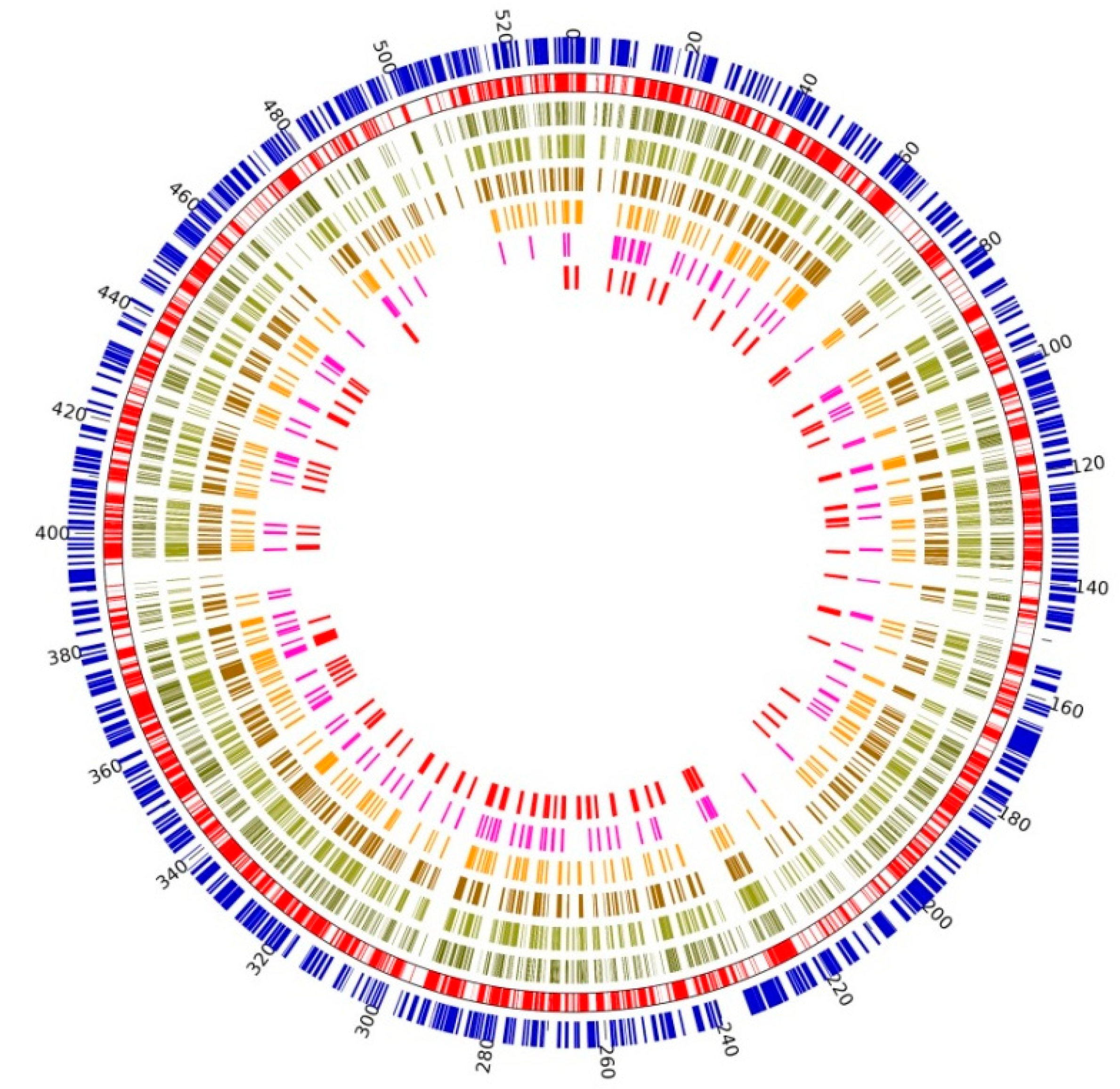
2.2. Genomic Structure and Characteristic Analysis
2.3. Protein Prediction and Functional Annotation
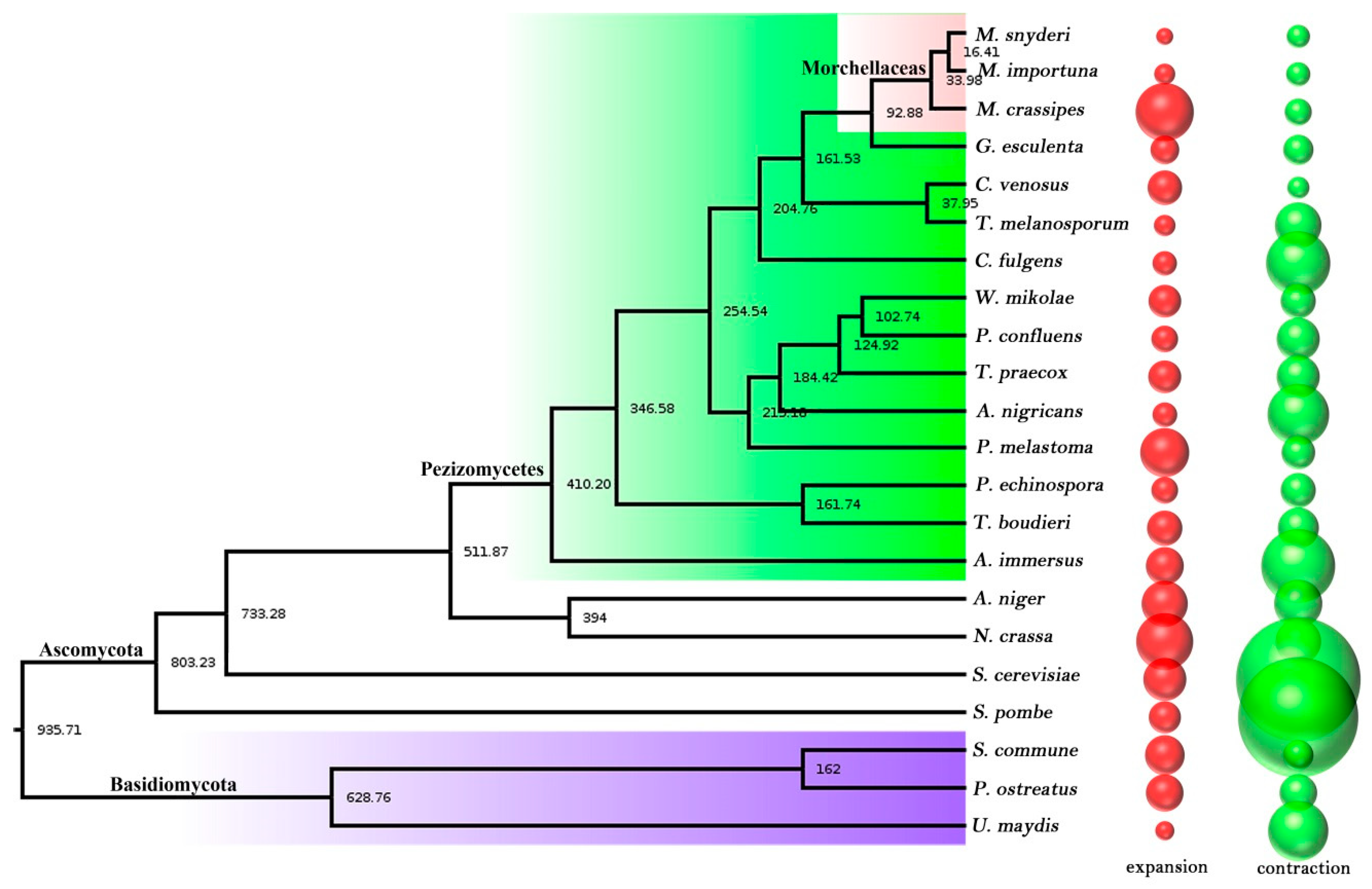
2.4. Phylogenetic Tree Construction and Evolution Analysis
2.5. Contraction and Expansion of Gene Families
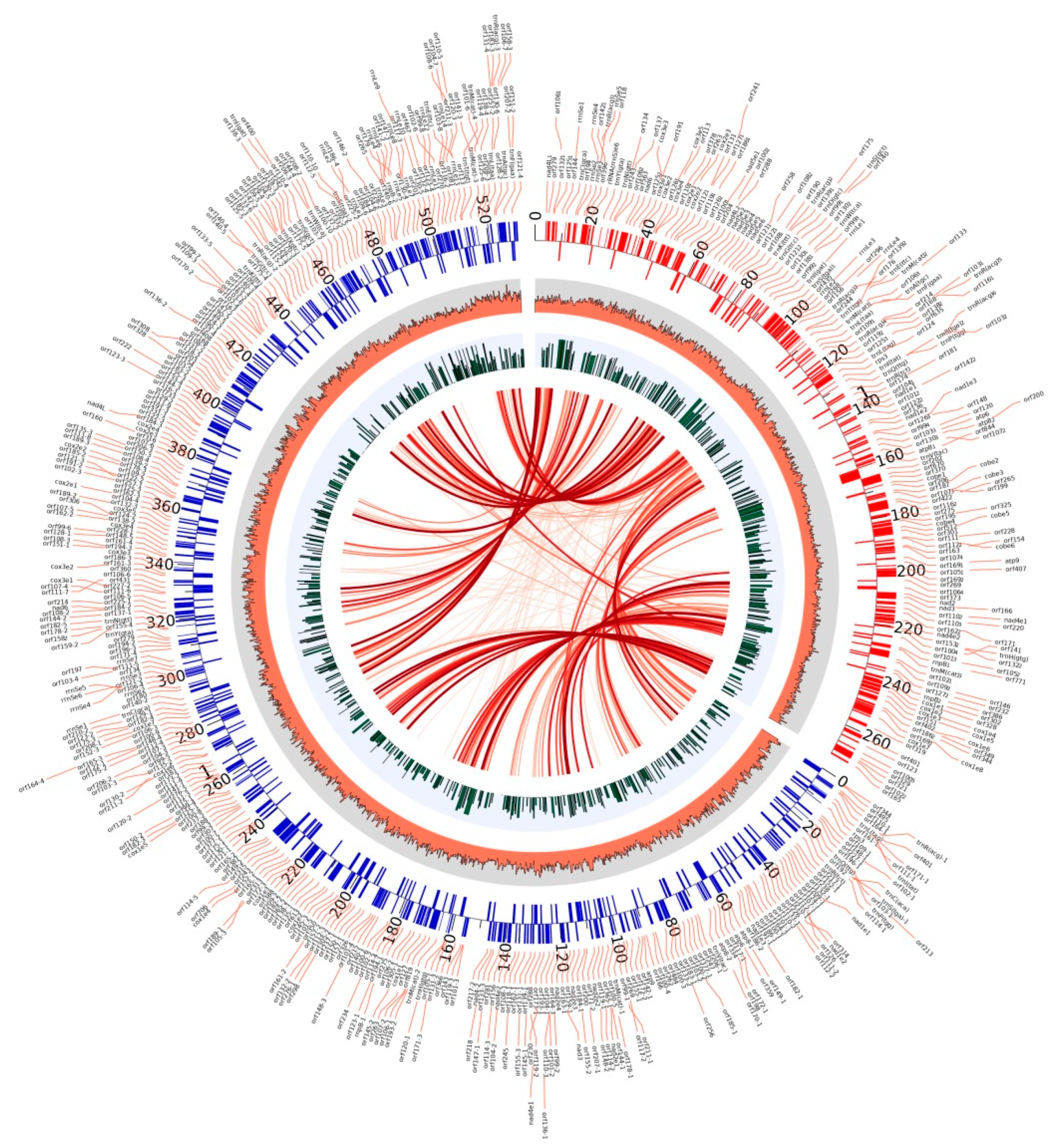
2.6. Mitochondrial Genome Assembly and Annotation of M. crassipes
2.7. Mitochondrial Genome Characteristic Analysis of M. crassipes
3. Discussion
3.1. Comparative Genome Analysis of M. crassipes
3.2. Comparative Analysis of CAZYme Enzymes
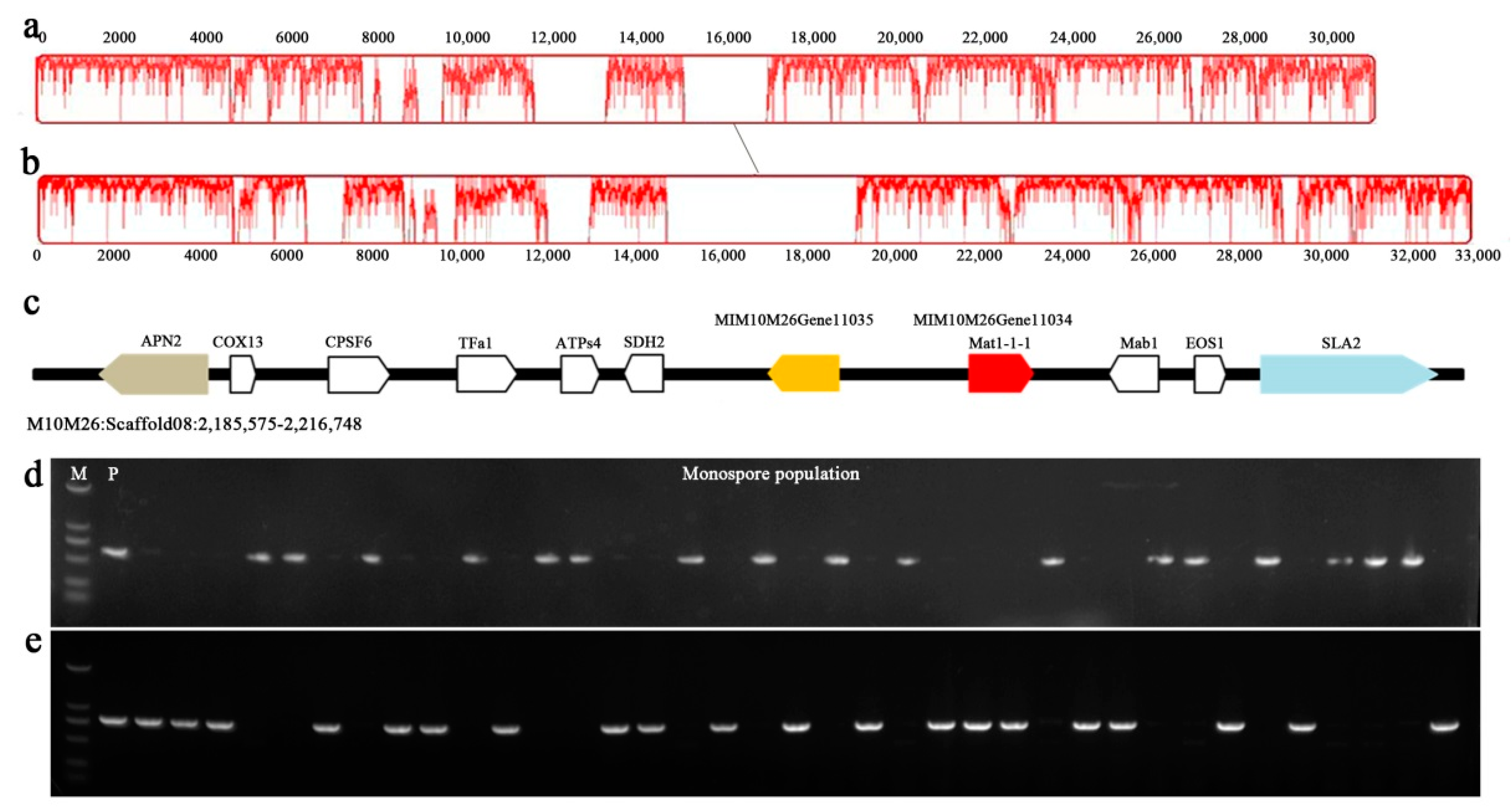
3.3. M. crassipes Is a Heterothalic Fungus
3.4. Comparative Analysis of Mitochondrial Genome
4. Materials and Methods
4.1. Strain Selection and Material Preparation
4.2. Genome and Transcriptome Sequencing
4.3. Genome Assembly and Annotation
4.4. Mitochondrial Genome Assembly, Gene Annotation and Bioinformatics Analysis
4.5. Phylogenetic Tree Construction and Comparative Genomic Analysis
4.6. DNA and RNA Extraction and PCR Amplification
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Liu, W.; Zhang, Y.; He, P. Morel Biology and Cultivation; Jilin science and Technology Press: Changchun, China, 2017. [Google Scholar]
- Pilz, D.; Rebecca, M.L.; Susan, A.; Luis, V.R.; Shannon, B.; Tricia, W.; Parks, C.G.; Erika, M.F.; Blaze, B. Ecology and Management of Morels Harvested from the Forests of Western North America; United States Department of Agriculture: Washington, DC, USA, 2007. [Google Scholar]
- O’Donnell, K.; Rooney, A.P.; Mills, G.L.; Kuo, M.; Weber, N.S.; Rehner, S.A. Phylogeny and historical biogeography of true morels (Morchella) reveals an early Cretaceous origin and high continental endemism and provincialism in the Holarctic. Fungal Genet. Biol. 2011, 48, 252–265. [Google Scholar] [CrossRef] [PubMed]
- Kuo, M. Morels; University of Michigan Press: Ann Arbor, MI, USA, 2005. [Google Scholar]
- Nitha, B.; Meera, C.R.; Janardhanan, K.K. Anti-inflammatory and antitumour activities of cultured mycelium of morel mushroom, Morchella esculenta. Curr. Sci. India. 2007, 92, 235–239. [Google Scholar]
- He, P.X.; Geng, L.J.; Mao, D.B.; Xu, C.P. Production, characterization and antioxidant activity of exopolysaccharides from submerged culture of Morchella crassipes. Bioproc. Biosyst. Eng. 2012, 35, 1325–1332. [Google Scholar] [CrossRef] [PubMed]
- Stojkovic, D.S.; Davidovic, S.; Zivkovic, J.; Glamoclija, J.; Ciric, A.; Stevanovic, M.; Ferreira, I.C.F.R.; Sokovic, M. Comparative evaluation of antimutagenic and antimitotic effects of Morchella esculenta extracts and protocatechuic acid. Front. Life Sci. 2013, 7, 218–223. [Google Scholar] [CrossRef]
- Ower, R. Notes on the development of the morel ascocarp: Morchella esculenta. Mycologia 1982, 74, 142–144. [Google Scholar] [CrossRef]
- He, P.X.; Wei, L.; Yingli, C.; Xin-sheng, H. Strain identification and phylogenetic analysis of cultivated and wild strains of Morchella belonging to elata clade in China. J. Zhengzhou Univ. Light Ind. 2015, 30, 26–29. [Google Scholar]
- Liu, W.; Cai, Y.; He, P.; Ma, X.; Bian, Y. Occurrence and control of pests and diseases in field cultivation of Morchella mushrooms. Acta Edulis Fungi 2019, 26, 7. [Google Scholar]
- He, P.X.; Wang, K.; Cai, Y.L.; Liu, W. Live cell confocal laser imaging studies on the nuclear behavior during meiosis and ascosporogenesis in Morchella importuna under artificial cultivation. Micron 2017, 101, 108–113. [Google Scholar] [CrossRef]
- Liu, W.; Chen, L.; Cai, Y.; Zhang, Q.; Bian, Y. Opposite polarity monospore genome de novo sequencing and comparative analysis reveal the possible heterothallic life cycle of Morchella importuna. Int. J. Mol. Sci. 2018, 19, 2525. [Google Scholar] [CrossRef]
- Liu, W.; Cai, Y.; He, P.; Bian, Y. Cultivation of monosporic and hybrid populations and polarity analysis of Morchella importuna. J. Fungal Res. 2019, 17, 7. [Google Scholar]
- Chai, H.; Chen, L.; Chen, W.; Zhao, Q.; Zhang, X.; Su, K.; Zhao, Y. Characterization of mating-type idiomorphs suggests that Morchella importuna, Mel-20 and M. sextelata are heterothallic. Mycol. Prog. 2017, 16, 743–752. [Google Scholar] [CrossRef]
- Du, X.H.; Zhao, Q.; Xia, E.H.; Gao, L.Z.; Richard, F.; Yang, Z.L. Mixed-reproductive strategies, competitive mating-type distribution and life cycle of fourteen black morel species. Sci. Rep. 2017, 7, 1493. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Volk, T.J.; Leonard, T.J. Physiological and environmental studies of sclerotium formation and maturation in isolates of Morchella crassipes. Appl. Environ. Microbiol. 1989, 55, 3095–3100. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- He, P.; Liu, W. Molecular identification of Morchella crassipes and phylogenetic analysis of Morchella fungi. Jiangsu Agric. Sci. 2010, 26, 5. [Google Scholar]
- Buscot, F.; Roux, J. Association between living roots and ascocarps of Morchella rotunda. Trans. Br. Mycol. Soc. 1987, 89, 249–252. [Google Scholar] [CrossRef]
- Buscot, F.; Kottke, I. The association of Morchella rotunda (Pers.) Boudier with roots of Picea abies (L.) Karst. New Phytol. 1990, 116, 425–430. [Google Scholar] [CrossRef]
- Buscot, F. Synthesis of two types of association between Morchella esculenta and Picea abies under controlled culture conditions. J. Plant Physiol. 1993, 141, 12–17. [Google Scholar] [CrossRef]
- Dahlstrom, J.L.; Smith, J.E.; Weber, N.S. Mycorrhiza-like interaction by Morchella with species of the Pinaceae in pure culture synthesis. Mycorrhiza 2000, 9, 279–285. [Google Scholar] [CrossRef]
- Hobbie, E.A.; Rice, S.F.; Weber, N.S.; Smith, J.E. Isotopic evidence indicates saprotrophy in post-fire Morchella in Oregon and Alaska. Mycologia 2016, 108, 638–645. [Google Scholar] [CrossRef]
- Hervey, A.; Bistis, G.; Leong, I. Cultural studies of single ascospore isolates of Morchella esculenta. Mycologia 1978, 70, 1269–1274. [Google Scholar] [CrossRef]
- Martin, F.; Kohler, A.; Murat, C.; Balestrini, R.; Coutinho, P.M.; Jaillon, O.; Montanini, B.; Morin, E.; Noel, B.; Percudani, R.; et al. Périgord black truffle genome uncovers evolutionary origins and mechanisms of symbiosis. Nature 2010, 464, 1033. [Google Scholar] [CrossRef] [PubMed]
- Labbé, J.; Zhang, X.; Yin, T.; Schmutz, J.; Grimwood, J.; Martin, F.; Tuskan, G.A.; Le Tacon, F. A genetic linkage map for the ectomycorrhizal fungus Laccaria bicolor and its alignment to the whole-genome sequence assemblies. New Phytol. 2008, 180, 316–328. [Google Scholar]
- Wingfield, B.D.; Bills, G.F.; Dong, Y.; Huang, W.; Nel, W.J.; Swalarsk-Parry, B.S.; Vaghefi, N.; Wilken, P.M.; An, Z.; de Beer, Z.W. Draft genome sequence of Annulohypoxylon stygium, Aspergillus mulundensis, Berkeleyomyces basicola (syn. Thielaviopsis basicola), Ceratocystis smalleyi, two Cercospora beticola strains, Coleophoma cylindrospora, Fusarium fracticaudum, Phialophora cf. hyalina, and Morchella septimelata. Ima Fungus 2018, 9, 199. [Google Scholar] [PubMed]
- Burger, G.; Gray, M.W.; Franz Lang, B. Mitochondrial genomes: Anything goes. Trends Genet. 2003, 19, 709–716. [Google Scholar] [CrossRef] [PubMed]
- Wai, T.; Langer, T. Mitochondrial dynamics and metabolic regulation. Trends Endocrinol. Metab. 2016, 27, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Cai, Y.; Zhang, Q.; Chen, L.; Shu, F.; Ma, X.; Bian, Y. The mitochondrial genome of Morchella importuna (272.2 kb) is the largest among fungi and contains numerous introns, mitochondrial non-conserved open reading frames and repetitive sequences. Int. J. Biol. Macromol. 2020, 143, 373–381. [Google Scholar] [CrossRef]
- Trappe, M.J.; Trappe, J.M.; Bonito, G.M. Kalapuya brunnea gen. & sp nov and its relationship to the other sequestrate genera in Morchellaceae. Mycologia 2010, 102, 1058–1065. [Google Scholar]
- He, P.; Wang, K.; Cai, Y.; Hu, X.; Zheng, Y.; Zhang, J.; Liu, W. Involvement of autophagy and apoptosis and lipid accumulation in sclerotial morphogenesis of Morchella importuna. Micron 2018, 109, 34–40. [Google Scholar] [CrossRef]
- Aubert, G.; Lansdorp, P.M. Telomeres and aging. Physiol. Rev. 2008, 88, 557–579. [Google Scholar] [CrossRef]
- He, P.; Cai, Y.; Liu, S.; Han, L.; Huang, L.; Liu, W. Morphological and ultrastructural examination of senescence in Morchella elata. Micron 2015, 78, 79–84. [Google Scholar] [CrossRef]
- He, P.; Yu, M.; Cai, Y.; Liu, W.; Wang, W.; Wang, S.; Li, J. Effect of ageing on culture and cultivation of the culinary-medicinal mushrooms, Morchella importuna and M. sextelata (Ascomycetes). Int. J. Med. Mushrooms 2019, 21, 1089–1098. [Google Scholar] [CrossRef]
- Gibson, J.L.; Kimbrough, J.W. Ultrastructural observations on Helvellaceae (Pezizales). ii. ascosporogenesis of Gyromitra esculenta. Can. J. Bot. 1988, 66, 1743–1749. [Google Scholar] [CrossRef]
- Losada, L.; Pakala, S.B.; Fedorova, N.D.; Joardar, V.; Shabalina, S.A.; Hostetler, J.; Pakala, S.M.; Zafar, N.; Thomas, E.; Rodriguez-Carres, M.; et al. Mobile elements and mitochondrial genome expansion in the soil fungus and potato pathogen Rhizoctonia solani AG-3. FEMS Microbiol. Lett. 2014, 352, 165–173. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Salavirta, H.; Oksanen, I.; Kuuskeri, J.; Makela, M.; Laine, P.; Paulin, L.; Lundell, T. Mitochondrial genome of Phlebia radiata is the second largest (156 kbp) among fungi and features signs of genome flexibility and recent recombination events. PLoS ONE 2014, 9, e97141. [Google Scholar] [CrossRef]
- Mardanov, A.V.; Beletsky, A.V.; Kadnikov, V.V.; Ignatov, A.N.; Ravin, N.V. The 203 kbp mitochondrial genome of the phytopathogenic fungus Sclerotinia borealis reveals multiple invasions of introns and genomic duplications. PLoS ONE 2014, 9, e107536. [Google Scholar] [CrossRef]
- Li, Y.; Hu, X.D.; Yang, R.H.; Hsiang, T.; Wang, K.; Liang, D.Q.; Liang, F.; Cao, D.M.; Zhou, F.; Wen, G.; et al. Complete mitochondrial genome of the medicinal fungus Ophiocordyceps sinensis. Sci. Rep. 2015, 5, 13892. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Q.; Liu, W.; Cai, Y.; Lan, A.-F.; Bian, Y. Validation of internal control genes for quantitative real-time PCR gene expression analysis in Morchella. Molecules 2018, 23, 2331. [Google Scholar] [CrossRef] [Green Version]
- Butler, J.; MacCallum, I.; Kleber, M.; Shlyakhter, I.A.; Belmonte, M.K.; Lander, E.S.; Nusbaum, C.; Jaffe, D.B. ALLPATHS: De novo assembly of whole-genome shotgun microreads. Genome Res. 2008, 18, 810–820. [Google Scholar] [CrossRef] [Green Version]
- Ye, C.; Hill, C.M.; Wu, S.; Ruan, J.; Ma, Z. DBG2OLC: Efficient assembly of large genomes using long erroneous reads of the third generation sequencing technologies. Sci. Rep. 2016, 6, 31900. [Google Scholar] [CrossRef]
- Chakraborty, M.; Baldwin-Brown, J.G.; Long, A.D.; Emerson, J.J. Contiguous and accurate de novo assembly of metazoan genomes with modest long read coverage. Nucleic Acids Res. 2016, 44, e147. [Google Scholar]
- Boetzer, M.; Henkel, C.V.; Jansen, H.J.; Butler, D.; Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 2011, 27, 578–579. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luo, R.; Liu, B.; Xie, Y.; Li, Z.; Huang, W.; Yuan, J.; He, G.; Chen, Y.; Pan, Q.; Liu, Y. SOAPdenovo2: An empirically improved memory-efficient short-read de novo assembler. Gigascience 2012, 1, 18. [Google Scholar] [CrossRef] [PubMed]
- Nansheng, C. Using RepeatMasker to Identify Repetitive Elements in Genomic Sequences. Curr. Protoc. Bioinform. 2004, 5. [Google Scholar] [CrossRef]
- Smit, A.F.; Hubley, R. RepeatModeler Open-1.0. Available online: http://www.repeatmasker.org (accessed on 31 July 2015).
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357. [Google Scholar] [CrossRef] [Green Version]
- Birney, E.; Clamp, M.; Durbin, R. GeneWise and genomewise. Genome Res. 2004, 14, 988–995. [Google Scholar] [CrossRef] [Green Version]
- Stanke, M.; Keller, O.; Gunduz, I.; Hayes, A.; Waack, S.; Morgenstern, B. AUGUSTUS: Ab initio prediction of alternative transcripts. Nucleic Acids Res. 2006, 34, W435–W439. [Google Scholar] [CrossRef] [Green Version]
- Punta, M.; Coggill, P.C.; Eberhardt, R.Y.; Mistry, J.; Tate, J.; Boursnell, C.; Pang, N.; Forslund, K.; Ceric, G.; Clements, J. The Pfam protein families database. Nucleic Acids Res. 2011, 40, D290–D301. [Google Scholar] [CrossRef]
- Conesa, A.; Götz, S.; García-Gómez, J.M.; Terol, J.; Talón, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [Green Version]
- Yin, Y.; Mao, X.; Yang, J.; Chen, X.; Mao, F.; Xu, Y. dbCAN: A web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 2012, 40, W445–W451. [Google Scholar] [CrossRef]
- Zdobnov, E.M.; Apweiler, R. InterProScan-an integration platform for the signature-recognition methods in InterPro. Bioinformatics 2001, 17, 847–848. [Google Scholar] [CrossRef] [Green Version]
- Beier, S.; Thiel, T.; Münch, T.; Scholz, U.; Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 2017, 33, 2583–2585. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simão, F.A.; Waterhouse, R.M.; Ioannidis, P.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 2015, 31, 3210–3212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef] [PubMed]
- Darling, A.C.E.; Mau, B.; Blattner, F.R.; Perna, N.T. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 2004, 14, 1394–1403. [Google Scholar] [CrossRef] [Green Version]
- Lohse, M.; Drechsel, O.; Kahlau, S.; Bock, R. OrganellarGenomeDRAW-a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res. 2013, 41, W575–W581. [Google Scholar] [CrossRef]
- Li, L.; Stoeckert, C.J.; Roos, D.S. OrthoMCL: Identification of ortholog groups for eukaryotic genomes. Genome Res. 2003, 13, 2178–2189. [Google Scholar] [CrossRef] [Green Version]
- Katoh, K.; Misawa, K.; Kuma, K.I.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [Green Version]
- Darriba, D.; Taboada, G.L.; Doallo, R.; Posada, D. ProtTest 3: Fast selection of best-fit models of protein evolution. Bioinformatics 2011, 27, 1164–1165. [Google Scholar] [CrossRef] [Green Version]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Taylor, J.W.; Berbee, M.L. Dating divergences in the Fungal Tree of Life: Review and new analyses. Mycologia 2006, 98, 838–849. [Google Scholar] [CrossRef]
- Douzery, E.J.P.; Snell, E.A.; Bapteste, E.; Delsuc, F.; Philippe, H. The timing of eukaryotic evolution: Does a relaxed molecular clock reconcile proteins and fossils? Proc. Natl. Acad. Sci. USA 2004, 101, 15386–15391. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Bie, T.; Cristianini, N.; Demuth, J.P.; Hahn, M.W. CAFE: A computational tool for the study of gene family evolution. Bioinformatics 2006, 22, 1269–1271. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Description | Characteristics |
|---|---|
| the genome scaffold number (n) | 23 |
| the genome contig number (n) | 97 |
| the longest length (bp) | 7,341,407 |
| the shortest length (bp) | 544,727 |
| the genome size estimate (bp) | 58,642,780 |
| the genome scaffold size (bp) | 56,756,124 |
| the genome contig size (bp) | 56,627,527 |
| the rate of N | 0.0023 |
| the rate of GC | 0.4734 |
| the scaffold N50 (bp) | 2,999,225 |
| the contig N50 (bp) | 814,084 |
| the scaffold N90 (bp) | 1,236,361 |
| the contig N90 (bp) | 363,112 |
| the BUSCO genome integrity (%) | 98.62 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, W.; Cai, Y.; Zhang, Q.; Shu, F.; Chen, L.; Ma, X.; Bian, Y. Subchromosome-Scale Nuclear and Complete Mitochondrial Genome Characteristics of Morchella crassipes. Int. J. Mol. Sci. 2020, 21, 483. https://doi.org/10.3390/ijms21020483
Liu W, Cai Y, Zhang Q, Shu F, Chen L, Ma X, Bian Y. Subchromosome-Scale Nuclear and Complete Mitochondrial Genome Characteristics of Morchella crassipes. International Journal of Molecular Sciences. 2020; 21(2):483. https://doi.org/10.3390/ijms21020483
Chicago/Turabian StyleLiu, Wei, Yingli Cai, Qianqian Zhang, Fang Shu, Lianfu Chen, Xiaolong Ma, and Yinbing Bian. 2020. "Subchromosome-Scale Nuclear and Complete Mitochondrial Genome Characteristics of Morchella crassipes" International Journal of Molecular Sciences 21, no. 2: 483. https://doi.org/10.3390/ijms21020483






