miRNAs and lncRNAs in Echinococcus and Echinococcosis
Abstract
:1. Introduction
2. Widespread Expression of miRNAs in Echinococcus spp. According to Transcriptome Analysis
2.1. MiRNAs Expressed in Different Developmental Stages of E. granulosus Sensu Stricto
2.2. MiRNA Expression Profiles in E. canadensis
2.3. MiRNA Expression Patterns in E. multilocularis
2.4. Common miRNAs in Echinococcus spp.
3. Non-Coding RNAs in Intermediate Hosts during Infection with Echinococcus spp.
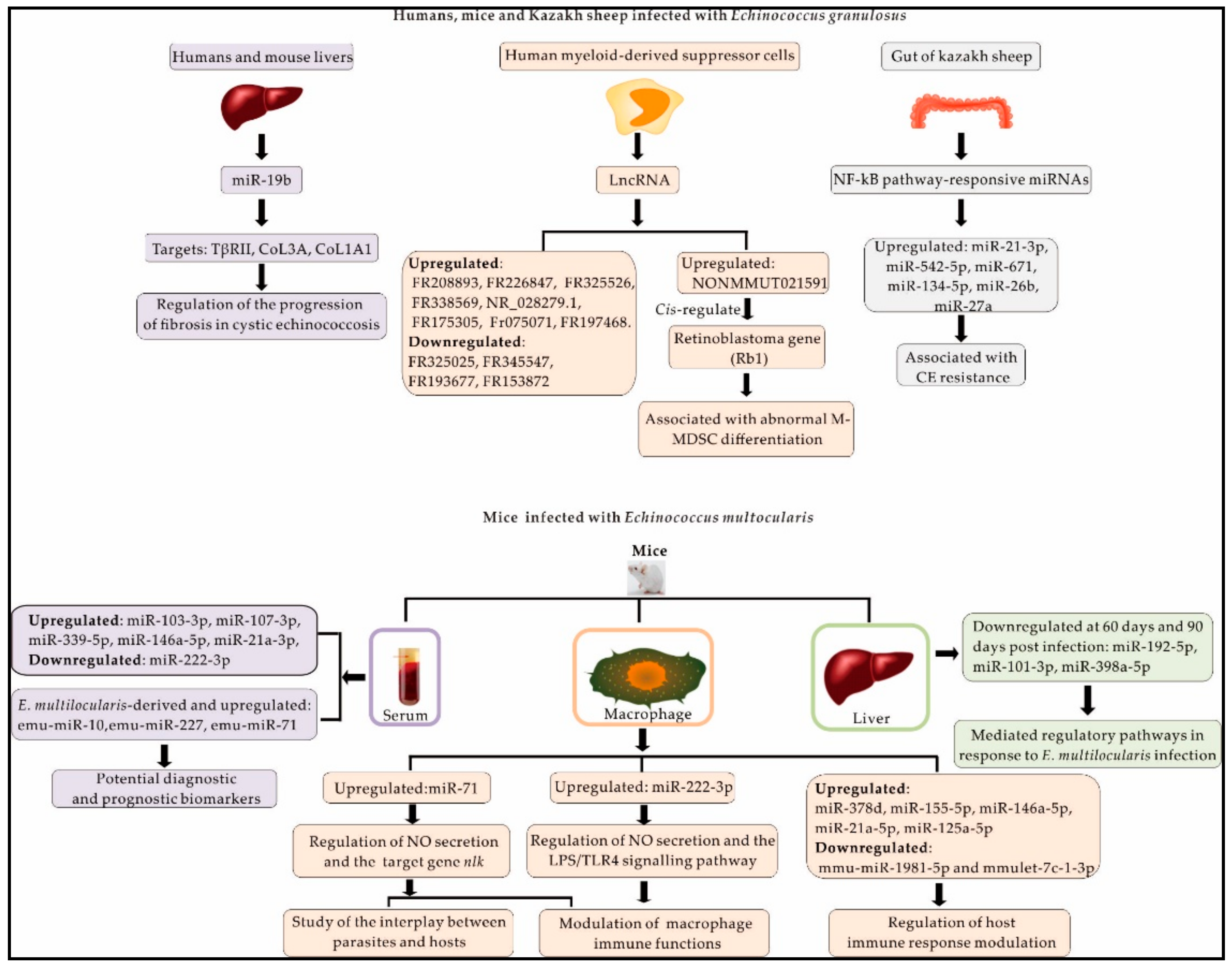
3.1. MiRNAs and lncRNAs in Host Responses to E. granulosus
3.2. Mouse miRNAs Dysregulated during Infection with E. multilocularis
3.3. Common miRNA Families in the Host Model during Infection with Echinococcus spp.
4. MiRNAs Mainly Associated with Immune and Pathological Processes during Host Infection with Echinococcus spp.
4.1. MiR-71 as an Innate Immune Regulator in Echinococcosis
4.2. miR-19b as an Effective Treatment Biomarker
4.3. miR-222-3p Modulates Macrophage Immunity
5. Echinococcus miRNA-Related Databases
6. Techniques and Methods Used in miRNA Studies in Echinococcus and Echinococcosis
6.1. MiRNA Identification
6.2. Verification of the Functions and Exploration of the Mechanisms
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Cucher, M.; Macchiaroli, N.; Kamenetzky, L.; Maldonado, L.; Brehm, K.; Rosenzvit, M.C. High-throughput characterization of Echinococcus spp. metacestode miRNomes. Int. J. Parasitol. 2015, 45, 253–267. [Google Scholar] [CrossRef]
- Yan, B.; Liu, X.; Wu, J.; Zhao, S.; Yuan, W.; Wang, B.; Wureli, H.; Tu, C.; Chen, C.; Wang, Y. Genetic Diversity of Echinococcus granulosus Genotype G1 in Xinjiang, Northwest of China. Korean J. Parasitol. 2018, 56, 391–396. [Google Scholar] [CrossRef] [Green Version]
- McManus, D.P.; Gray, D.J.; Zhang, W.; Yang, Y. Diagnosis, treatment, and management of echinococcosis. BMJ 2012, 344, e3866. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Monteiro, K.M.; De Carvalho, M.O.; Zaha, A.; Ferreira, H.B. Proteomic analysis of the Echinococcus granulosus metacestode during infection of its intermediate host. Proteomics 2010, 10, 1985–1999. [Google Scholar] [CrossRef]
- Zhang, C.; Wang, J.; Lü, G.; Li, J.; Lu, X.; Mantion, G.; Vuitton, D.A.; Wen, H.; Lin, R. Hepatocyte Proliferation/Growth Arrest Balance in the Liver of Mice during E. multilocularis Infection: A Coordinated 3-Stage Course. PLoS ONE 2012, 7, e30127. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, Y. Suppression of mouse miRNA-222-3p in response to Echinococcus multilocularis infection. Int. Immunopharmacol. 2018, 64, 252–255. [Google Scholar] [CrossRef] [PubMed]
- Craig, P.S.; Hegglin, D.; Lightowlers, M.W.; Torgerson, P.R.; Wang, Q. Echinococcosis: Control and Prevention. Adv. Parasitol. 2017, 96, 55–158. [Google Scholar]
- Deplazes, P.; Rinaldi, L.; Rojas, C.A.; Torgerson, P.; Harandi, M.; Romig, T.; Antolova, D.; Schurer, J.; Lahmar, S.; Cringoli, G.; et al. Global Distribution of Alveolar and Cystic Echinococcosis; Elsevier BV: Amsterdam, The Netherlands, 2017; Volume 95, pp. 315–493. [Google Scholar]
- Wang, Z.; Wang, X.; Liu, X. Echinococcosis in China, a review of the epidemiology of Echinococcus spp. Ecohealth 2008, 5, 115–126. [Google Scholar] [PubMed]
- Giraudoux, P.; Zhao, Y.; Afonso, E.; Yan, H.; Knapp, J.; Rogan, M.T.; Shi, D.; Jia, W.; Craig, P.S. Long-term retrospective assessment of a transmission hotspot for human alveolar echinococcosis in mid-west China. PLOS Neglected Trop. Dis. 2019, 13, e0007701. [Google Scholar] [CrossRef] [Green Version]
- Nakao, M.; Lavikainen, A.; Yanagida, T.; Ito, A. Phylogenetic systematics of the genus Echinococcus (Cestoda: Taeniidae). Int. J. Parasitol. 2013, 43, 1017–1029. [Google Scholar] [CrossRef]
- Macchiaroli, N.; Cucher, M.; Zarowiecki, M.; Maldonado, L.; Kamenetzky, L.; Rosenzvit, M.C. microRNA profiling in the zoonotic parasite Echinococcus canadensis using a high-throughput approach. Parasit. Vectors 2015, 8, 83. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lekka, E.; Hall, J. Noncoding RNAs in disease. FEBS Lett. 2018, 592, 2884–2900. [Google Scholar] [CrossRef] [PubMed]
- Tsai, I.J.; The Taenia Solium Genome Consortium; Zarowiecki, M.; Holroyd, N.; Garciarrubio, A.; Sánchez-Flores, A.; Brooks, K.L.; Tracey, A.; Bobes, R.J.; Fragoso, G.; et al. The genomes of four tapeworm species reveal adaptations to parasitism. Nature 2013, 496, 57–63. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, H.; Zhang, W.; Zhang, L.; Zhang, Z.; Li, J.; Lu, G.; Zhu, Y.; Wang, Y.; Huang, Y.; Liu, J.; et al. The genome of the hydatid tapeworm Echinococcus granulosus. Nat. Genet. 2013, 45, 1168–1175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartel, D.P. Metazoan MicroRNAs. Cell 2018, 173, 20–51. [Google Scholar] [CrossRef] [PubMed]
- Barbagallo, D.; Vittone, G.; Romani, M.; Purrello, M. Noncoding RNAs in Health and Disease. Int. J. Genom. 2018, 2018, 1–2. [Google Scholar] [CrossRef] [Green Version]
- Ancarola, M.E.; Marcilla, A.; Herz, M.; Macchiaroli, N.; Pérez, M.; Asurmendi, S.; Brehm, K.; Poncini, C.; Rosenzvit, M.; Cucher, M. Cestode parasites release extracellular vesicles with microRNAs and immunodiagnostic protein cargo. Int. J. Parasitol. 2017, 47, 675–686. [Google Scholar] [CrossRef]
- Yu, A.; Wang, Y.; Yin, J.; Zhang, J.; Cao, S.; Cao, J.; Shen, Y. Microarray analysis of long non-coding RNA expression profiles in monocytic myeloid-derived suppressor cells in Echinococcus granulosus-infected mice. Parasit. Vectors 2018, 11, 327. [Google Scholar] [CrossRef]
- Ameres, S.L.; Zamore, P.D. Diversifying microRNA sequence and function. Nat. Rev. Mol. Cell Boil. 2013, 14, 475–488. [Google Scholar] [CrossRef]
- Bian, E.-B.; Xiong, Z.-G.; Li, J. New advances of lncRNAs in liver fibrosis, with specific focus on lncRNA–miRNA interactions. J. Cell. Physiol. 2018, 234, 2194–2203. [Google Scholar] [CrossRef]
- Zhang, Z.; Yang, T.; Xiao, J. Circular RNAs: Promising Biomarkers for Human Diseases. EBioMedicine 2018, 34, 267–274. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Macchiaroli, N.; Maldonado, L.L.; Zarowiecki, M.; Cucher, M.; Gismondi, M.I.; Kamenetzky, L.; Rosenzvit, M.C. Genome-wide identification of microRNA targets in the neglected disease pathogens of the genus Echinococcus. Mol. Biochem. Parasitol. 2017, 214, 91–100. [Google Scholar] [CrossRef] [PubMed]
- Judice, C.C.; Bourgard, C.; Kayano, A.C.A.V.; Albrecht, L.; Costa, F.T.M. MicroRNAs in the Host-Apicomplexan Parasites Interactions: A Review of Immunopathological Aspects. Front. Microbiol. 2016, 6, 499. [Google Scholar] [CrossRef] [Green Version]
- Carrière, J.; Barnich, N.; Nguyen, H.T.T. Exosomes: From Functions in Host-Pathogen Interactions and Immunity to Diagnostic and Therapeutic Opportunities. Rev. Physiol. Biochem. Pharmacol. 2016, 172, 39–75. [Google Scholar] [PubMed]
- Cai, P.; Gobert, G.N.; McManus, D.P.; Information, P.E.K.F.C. MicroRNAs in Parasitic Helminthiases: Current Status and Future Perspectives. Trends Parasitol. 2016, 32, 71–86. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y. The novel regulatory role of lncRNA-miRNA-mRNA axis in cardiovascular diseases. J. Cell. Mol. Med. 2018, 22, 5768–5775. [Google Scholar] [CrossRef]
- Zampetaki, A.; Albrecht, A.; Steinhofel, K. Long Non-coding RNA Structure and Function: Is There a Link? Front. Physiol. 2018, 9, 9. [Google Scholar] [CrossRef]
- Yue, Y.; Guo, T.; Yuan, C.; Liu, J.; Guo, J.; Feng, R.; Niu, C.; Sun, X.; Yang, B. Integrated analysis of the roles of long noncoding RNA and coding RNA expression in sheep (Ovis aries) skin during initiation of secondary hair follicle. PLoS ONE 2016, 11, e0156890. [Google Scholar] [CrossRef] [Green Version]
- Zhan, S.; Dong, Y.; Zhao, W.; Guo, J.; Zhong, T.; Wang, L.; Li, L.; Zhang, H. Genome-wide identification and characterization of long non-coding RNAs in developmental skeletal muscle of fetal goat. BMC Genom. 2016, 17, 666. [Google Scholar] [CrossRef] [Green Version]
- Kern, C.; Wang, Y.; Chitwood, J.; Korf, I.; Delany, M.; Cheng, H.; Medrano, J.F.; Van Eenennaam, A.L.; Ernst, C.; Ross, P.; et al. Genome-wide identification of tissue-specific long non-coding RNA in three farm animal species. BMC Genom. 2018, 19, 684. [Google Scholar] [CrossRef] [Green Version]
- Gao, P.F.; Guo, X.H.; Du, M.; Cao, G.Q.; Yang, Q.C.; Pu, Z.D.; Wang, Z.Y.; Zhang, Q.; Li, M.; Jin, Y.S.; et al. LncRNA profiling of skeletal muscles in Large White pigs and Mashen pigs during development. J. Anim. Sci. 2017, 95, 4239–4250. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scott, E.Y.; Mansour, T.; Bellone, R.R.; Brown, C.T.; Mienaltowski, M.J.; Penedo, M.C.; Ross, P.J.; Valberg, S.J.; Murray, J.D.; Finno, C.J. Identification of long non-coding RNA in the horse transcriptome. BMC Genom. 2017, 18, 511. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Bai, J.; Wu, A.; Wang, Y.; Zhang, J.; Wang, Z.; Li, Y.; Xu, J.; Li, X. Co-LncRNA: Investigating the lncRNA combinatorial effects in GO annotations and KEGG pathways based on human RNA-Seq data. Database 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Le Béguec, C.; Wucher, V.; Lagoutte, L.; Cadieu, E.; Botherel, N.; Hédan, B.; De Brito, C.; Guillory, A.-S.; André, C.; Derrien, T.; et al. Characterisation and functional predictions of canine long non-coding RNAs. Sci. Rep. 2018, 8, 13444. [Google Scholar] [CrossRef] [Green Version]
- Woehle, C.; Kusdian, G.; Radine, C.; Graur, D.; Landan, G.; Gould, S.B. The parasite Trichomonas vaginalis expresses thousands of pseudogenes and long non-coding RNAs independently from functional neighbouring genes. BMC Genom. 2014, 15, 906. [Google Scholar] [CrossRef] [Green Version]
- Liu, W.; Huang, L.; Wei, Q.; Zhang, Y.; Zhang, S.; Zhang, W.; Cai, L.; Liang, S. Microarray analysis of long non-coding RNA expression profiles uncovers a Toxoplasma-induced negative regulation of host immune signaling. Parasit. Vectors 2018, 11, 174. [Google Scholar] [CrossRef]
- Kim, V.N. MicroRNA biogenesis: Coordinated cropping and dicing. Nat. Rev. Mol. Cell Boil. 2005, 6, 376–385. [Google Scholar] [CrossRef]
- Kwak, P.B.; Tomari, Y. The N domain of Argonaute drives duplex unwinding during RISC assembly. Nat. Struct. Mol. Boil. 2012, 19, 145–151. [Google Scholar] [CrossRef]
- Cheloufi, S.; Dos Santos, C.O.; Chong, M.M.W.; Hannon, G.J. A dicer-independent miRNA biogenesis pathway that requires Ago catalysis. Nature 2010, 465, 584–589. [Google Scholar] [CrossRef] [Green Version]
- Du, W.W.; Zhang, C.; Yang, W.; Yong, T.; Awan, F.M.; Yang, B.B. Identifying and Characterizing circRNA-Protein Interaction. Theranostics 2017, 7, 4183–4191. [Google Scholar] [CrossRef]
- Zhang, X.; Wang, W.; Zhu, W.; Dong, J.; Cheng, Y.; Yin, Z.; Shen, F. Mechanisms and Functions of Long Non-Coding RNAs at Multiple Regulatory Levels. Int. J. Mol. Sci. 2019, 20, 5573. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, X.; Zheng, Y. Expression profiling of circulating miRNAs in mouse serum in response to Echinococcus multilocularis infection. Parasitology 2017, 144, 1079–1087. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Zhang, Z.; Jin, L.; Kang, H.; Zhu, Y.; Zhang, L.; Li, X.; Ma, F.; Zhao, L.; Shi, B.; et al. Genome-wide sequencing of small RNAs reveals a tissue-specific loss of conserved microRNA families in Echinococcus granulosus. BMC Genom. 2014, 15, 736. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cucher, M.; Prada, L.; Mourglia-Ettlin, G.; Dematteis, S.; Camicia, F.; Asurmendi, S.; Rosenzvit, M. Identification of Echinococcus granulosus microRNAs and their expression in different life cycle stages and parasite genotypes. Int. J. Parasitol. 2011, 41, 439–448. [Google Scholar] [CrossRef]
- Kamenetzky, L.; Stegmayer, G.; Maldonado, L.; Macchiaroli, N.; Yones, C.; Milone, D. MicroRNA discovery in the human parasite Echinococcus multilocularis from genome-wide data. Genomics 2016, 107, 274–280. [Google Scholar] [CrossRef]
- Wang, Z.R.; Bo, X.W.; Zhang, Y.Y.; MA, X.; Lu, P.P.; Xu, M.F.; Meng, J.M. microRNA profile analyses of the protoscoleces in Echinococcus granulosus. Acta Vet. et Zootech. Sin. 2018, 49, 2477–2485. [Google Scholar]
- Jin, X.; Lu, L.; Su, H.; Lou, Z.; Wang, F.; Zheng, Y.; Xu, G.-T. Comparative analysis of known miRNAs across platyhelminths. FEBS J. 2013, 280, 3944–3951. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mortezaei, S.; Afgar, A.; Mohammadi, M.A.; Mousavi, S.M.; Sadeghi, B.; Harandi, M.F. The effect of albendazole sulfoxide on the expression of miR-61 and let-7 in different in vitro developmental stages of Echinococcus granulosus. Acta Trop. 2019, 195, 97–102. [Google Scholar] [CrossRef]
- Buck, A.H.; Coakley, G.; Simbari, F.; McSorley, H.J.; Quintana, J.F.; Le Bihan, T.; Kumar, S.; Abreu-Goodger, C.; Lear, M.; Harcus, Y.; et al. Exosomes secreted by nematode parasites transfer small RNAs to mammalian cells and modulate innate immunity. Nat. Commun. 2014, 5, 5488. [Google Scholar] [CrossRef]
- Quintana, J.F.; Makepeace, B.L.; Babayan, S.A.; Ivens, A.; Pfarr, K.M.; Blaxter, M.; Debrah, A.; Wanji, S.; Ngangyung, H.F.; Bah, G.S.; et al. Extracellular Onchocerca-derived small RNAs in host nodules and blood. Parasit. Vectors 2015, 8, 58. [Google Scholar] [CrossRef] [Green Version]
- Juvvuna, P.K.; Khandelia, P.; Lee, L.M.; Makeyev, E.V. Argonaute identity defines the length of mature mammalian microRNAs. Nucleic Acids Res. 2012, 40, 6808–6820. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Guo, X.; He, W.; Shao, Z.; Zhang, X.; Yang, J.; Shen, Y.; Luo, X.; Cao, J. Effects of Echinococcus multilocularis miR-71 mimics on murine macrophage RAW264.7 cells. Int. Immunopharmacol. 2016, 34, 259–262. [Google Scholar] [CrossRef]
- Guo, X.; Zhang, X.; Yang, J.; Jin, X.; Ding, J.; Xiang, H.; Ayaz, M.; Luo, X.; Zheng, Y. Suppression of nemo-like kinase by miR-71 in Echinococcus multilocularis. Exp. Parasitol. 2017, 183, 1–5. [Google Scholar] [CrossRef]
- Zheng, Y.; Cai, X.; Bradley, J.E. microRNAs in parasites and parasite infection. RNA Boil. 2013, 10, 371–379. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, X.; Zheng, Y. MicroRNA expression profile in RAW264.7 macrophage cells exposed to Echinococcus multilocularis metacestodes. Parasitology 2018, 145, 416–423. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Li, X.; Wang, X.; Ban, Q.; Hui, W.; Jia, B. MicroRNA profiling of the intestinal tissue of Kazakh sheep after experimental Echinococcus granulosus infection, using a high-throughput approach. Parasite 2016, 23, 23. [Google Scholar] [CrossRef] [Green Version]
- Schmid, M.; Zimara, N.; Wege, A.K.; Ritter, U. Myeloid-derived suppressor cell functionality and interaction with Leishmania major parasites differ in C57BL/6 and BALB/c mice. Eur. J. Immunol. 2014, 44, 3295–3306. [Google Scholar] [CrossRef]
- Pan, W.; Zhou, H.-J.; Shen, Y.-J.; Wang, Y.; Xu, Y.-X.; Hu, Y.; Jiang, Y.-Y.; Yuan, Z.-Y.; Ugwu, C.E.; Cao, J.-P. Surveillance on the Status of Immune Cells after Echinnococcus granulosus Protoscoleces Infection in Balb/c Mice. PLoS ONE 2013, 8, e59746. [Google Scholar] [CrossRef] [Green Version]
- Jin, X.; Guo, X.; Zhu, N.; Ayaz, M.; Zheng, Y. miRNA profiling in the mice in response to Echinococcus multilocularis infection. Acta Trop. 2017, 166, 39–44. [Google Scholar] [CrossRef]
- Jiang, M.-M.; Mai, Z.-T.; Wan, S.-Z.; Chi, Y.-M.; Zhang, X.; Sun, B.-H.; Di, Q.-G. Microarray profiles reveal that circular RNA hsa_circ_0007385 functions as an oncogene in non-small cell lung cancer tumorigenesis. J. Cancer Res. Clin. Oncol. 2018, 144, 667–674. [Google Scholar] [CrossRef]
- Wang, X.; Yong, C.; Yu, K.; Yu, R.; Zhang, R.; Yu, L.; Li, S.; Cai, S. Long Noncoding RNA (lncRNA) n379519 Promotes Cardiac Fibrosis in Post-Infarct Myocardium by Targeting miR-30. Med Sci. Monit. 2018, 24, 3958–3965. [Google Scholar] [CrossRef] [PubMed]
- Gao, M.; Li, C.; Xu, M.; Liu, Y.; Cong, M.; Liu, S. LncRNA MT1DP Aggravates Cadmium-Induced Oxidative Stress by Repressing the Function of Nrf2 and is Dependent on Interaction with miR-365. Adv. Sci. 2018, 5, 1800087. [Google Scholar] [CrossRef] [PubMed]
- Diao, L.; Wang, S.; Sun, Z. Long noncoding RNA GAPLINC promotes gastric cancer cell proliferation by acting as a molecular sponge of miR-378 to modulate MAPK1 expression. OncoTargets Ther. 2018, 11, 2797–2804. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qu, L.; Ding, J.; Chen, C.; Wu, Z.-J.; Liu, B.; Gao, Y.; Chen, W.; Liu, F.; Sun, W.; Li, X.-F.; et al. Exosome-Transmitted lncARSR Promotes Sunitinib Resistance in Renal Cancer by Acting as a Competing Endogenous RNA. Cancer Cell 2016, 29, 653–668. [Google Scholar] [CrossRef]
- Yan, T.H.; Qiu, C.; Sun, J.; Li, W.H. MiR-877-5p suppresses cell growth, migration and invasion by targeting cyclin dependent kinase 14 and predicts prognosis in hepatocellular carcinoma. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 3038–3046. [Google Scholar]
- Yu, M.J.; Zhao, N.; Shen, H.; Wang, H. Long Noncoding RNA MRPL39 Inhibits Gastric Cancer Proliferation and Progression by Directly Targeting miR-130. Genet. Test. Mol. Biomark. 2018, 22, 656–663. [Google Scholar] [CrossRef]
- Xie, Z.-Z.; Xiao, Z.-C.; Song, Y.-X.; Li, W.; Tan, G.-L. Long non-coding RNA Dleu2 affects proliferation, migration and invasion ability of laryngeal carcinoma cells through triggering miR-16-1 pathway. Eur. Rev. Med Pharmacol. Sci. 2018, 22, 1963–1970. [Google Scholar]
- Momen-Heravi, F.; Bala, S. miRNA regulation of innate immunity. J. Leukoc. Boil. 2018, 103, 1205–1217. [Google Scholar] [CrossRef]
- Cruz, L.O.; Hashemifar, S.S.; Wu, C.-J.; Cho, S.; Nguyen, D.T.; Lin, L.-L.; Khan, A.A.; Lu, L.-F. Excessive expression of miR-27 impairs Treg-mediated immunological tolerance. J. Clin. Investig. 2017, 127, 530–542. [Google Scholar] [CrossRef] [Green Version]
- Liu, H.; Chen, N.; Bi, J.; Han, J.; Yang, M.; Dong, W.; Lin, T.; Huang, J. Circular RNA circUBXN7 represses cell growth and invasion by sponging miR-1247-3p to enhance B4GALT3 expression in bladder cancer. Aging 2018, 10, 2606–2623. [Google Scholar] [CrossRef]
- Yang, J.; Liu, Q.; Cao, S.; Xu, T.; Li, X.; Zhou, D.; Pan, L.; Li, C.; Huang, C.; Meng, X.; et al. MicroRNA-145 increases the apoptosis of activated hepatic stellate cells induced by TRAIL through NF-kappaB signaling pathway. Front. Pharmacol. 2017, 8, 980. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghorbani, S.; Talebi, F.; Chan, W.F.; Masoumi, F.; Vojgani, M.; Power, C.; Noorbakhsh, F. MicroRNA-181 Variants Regulate T Cell Phenotype in the Context of Autoimmune Neuroinflammation. Front. Immunol. 2017, 8, 758. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gantier, M.P. X-chromosome-encoded microRNA-19 and -18 are possible modulators of female immunity. BioEssays 2013, 35, 671. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Cheng, Y. Inhibition of miR-20 promotes proliferation and autophagy in articular chondrocytes by PI3K/AKT/mTOR signaling pathway. Biomed. Pharmacother. 2018, 97, 607–615. [Google Scholar] [CrossRef]
- Luo, L.-J.; Zhang, L.-P.; Duan, C.-Y.; Wang, B.; He, N.-N.; Abulimiti, P.; Lin, Y. The inhibition role of miR-22 in hepatocellular carcinoma cell migration and invasion via targeting CD147. Cancer Cell Int. 2017, 17, 17. [Google Scholar] [CrossRef] [Green Version]
- Chen, M.; Hu, W.; Xiong, C.-L.; Qu, Z.; Yin, C.-Q.; Wang, Y.-H.; Luo, C.-L.; Guan, Q.; Yuan, C.-H.; Wang, F.-B. miR-22 targets YWHAZ to inhibit metastasis of hepatocellular carcinoma and its down-regulation predicts a poor survival. Oncotarget 2016, 7, 80751–80764. [Google Scholar] [CrossRef] [Green Version]
- Dong, Z.; Qi, R.; Guo, X.; Zhao, X.; Li, Y.; Zeng, Z.; Bai, W.; Chang, X.; Hao, L.; Chen, Y.; et al. MiR-223 modulates hepatocellular carcinoma cell proliferation through promoting apoptosis via the Rab1-mediated mTOR activation. Biochem. Biophys. Res. Commun. 2017, 483, 630–637. [Google Scholar] [CrossRef] [Green Version]
- Schueller, F.; Roy, S.; Loosen, S.H.; Alder, J.; Koppe, C.; Schneider, A.T.; Wandrer, F.; Bantel, H.; Vucur, M.; Mi, Q.-S.; et al. miR-223 represents a biomarker in acute and chronic liver injury. Clin. Sci. 2017, 131, 1971–1987. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.-F.; Li, S.-H.; Liu, Y.; Luo, Y.-T. Long Noncoding RNA CIR Promotes Chondrocyte Extracellular Matrix Degradation in Osteoarthritis by Acting as a Sponge For Mir-27b. Cell. Physiol. Biochem. 2017, 43, 602–610. [Google Scholar] [CrossRef]
- Wu, Y.; Sun, Q.; Dai, L. Immune regulation of miR-30 on the Mycobacterium tuberculosis-induced TLR/MyD88 signaling pathway in THP-1 cells. Exp. Ther. Med. 2017, 14, 3299–3303. [Google Scholar] [CrossRef] [Green Version]
- Ren, H.; Zhang, Y.; Zhu, H. MiR-339 depresses cell proliferation via directly targeting S-phase kinase-associated protein 2 mRNA in lung cancer. Thorac. Cancer 2018, 9, 408–414. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ying, X.; Zhang, W.; Fang, M.; Zhang, W.; Wang, C.; Han, L. miR-345-5p regulates proliferation, cell cycle, and apoptosis of acute myeloid leukemia cells by targeting AKT2. J. Cell. Biochem. 2018, 120, 1620–1629. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, J.; Chen, X.; Yang, Y.; Wang, F.; Li, W.; Awuti, M.; Sun, Y.; Lian, C.; Li, Z.; et al. miR-365 promotes diabetic retinopathy through inhibiting Timp3 and increasing oxidative stress. Exp. Eye Res. 2018, 168, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.-Z.; Wang, D.; Sun, X.-N.; Sheng, L. MicroRNA-378 acts as a prognosis marker and inhibits cell migration, invasion and epithelial-mesenchymal transition in human glioma by targeting IRG1. Eur. Rev. Med Pharmacol. Sci. 2018, 22, 3837–3846. [Google Scholar] [PubMed]
- Garros, R.F.; Paul, R.; Connolly, M.; Lewis, A.; Garfield, B.E.; Natanek, S.A.; Bloch, S.; Mouly, V.; Griffiths, M.J.; Polkey, M.I.; et al. MicroRNA-542 Promotes Mitochondrial Dysfunction and SMAD Activity and Is Elevated in Intensive Care Unit-acquired Weakness. Am. J. Respir. Crit. Care Med. 2017, 196, 1422–1433. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, Z.; Hou, Y.; Zhang, K.; Peng, X. gga-miR-99a targets SMARCA5 to regulate Mycoplasma gallisepticum (HS strain) infection by depressing cell proliferation in chicken. Gene 2017, 627, 239–247. [Google Scholar] [CrossRef]
- Pan, Y.; Wu, Y.; Hu, J.; Shan, Y.; Ma, J.; Ma, H.; Qi, X.; Jia, L. Long noncoding RNA HOTAIR promotes renal cell carcinoma malignancy through alpha-2, 8-sialyltransferase 4 by sponging microRNA-124. Cell Prolif. 2018, 51, e12507. [Google Scholar] [CrossRef] [Green Version]
- Dong, N.; Xu, B.; Shi, H. Long noncoding RNA MALAT1 acts as a competing endogenous RNA to regulate Amadori-glycated albumin-induced MCP-1 expression in retinal microglia by a microRNA-124-dependent mechanism. Inflamm. Res. 2018, 67, 913–925. [Google Scholar] [CrossRef]
- Wang, R.; Zhang, S.; Chen, X.; Li, N.; Li, J.; Jia, R.; Pan, Y.; Liang, H. EIF4A3-induced circular RNA MMP9 (circMMP9) acts as a sponge of miR-124 and promotes glioblastoma multiforme cell tumorigenesis. Mol. Cancer 2018, 17, 166. [Google Scholar] [CrossRef]
- Lakner, A.M.; Steuerwald, N.M.; Walling, T.L.; Ghosh, S.; Li, T.; McKillop, I.H.; Russo, M.W.; Bonkovsky, H.L.; Schrum, L.W. Inhibitory effects of microRNA 19b in hepatic stellate cell-mediated fibrogenesis. Hepatol. 2012, 56, 300–310. [Google Scholar] [CrossRef] [Green Version]
- Xueyong, Z. Primary Studies on the emu-miR-71 as Inhibiting Molecule of the Potential Drug Target Nemo-Like Kinase in Echinococcus Multilocularis Metacestode; Lanzhou Veterinary Research Institute: Lanzhou, China, 2015. [Google Scholar]
- Song, J.; Ouyang, Y.; Che, J.; Li, X.; Zhao, Y.; Yang, K.; Zhao, X.; Chen, Y.; Fan, C.; Yuan, W. Potential Value of miR-221/222 as Diagnostic, Prognostic, and Therapeutic Biomarkers for Diseases. Front. Immunol. 2017, 8, 29. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, Y. Strategies of Echinococcus species responses to immune attacks: Implications for therapeutic tool development. Int. Immunopharmacol. 2013, 17, 495–501. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yang, G.; Zhao, D.; Wang, J.; Bai, Y.; Peng, Q.; Wang, H.; Fang, R.; Chen, G.; Wang, Z.; et al. CD103-positive CSC exosome promotes EMT of clear cell renal cell carcinoma: Role of remote MiR-19b-3p. Mol. Cancer 2019, 18, 86. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.-X.; Guo, L.-X.; Pan, H.-S.; Liang, Y.-M.; Li, X. Circ_ANKIB1 stabilizes the regulation of miR-19b on SOCS3/STAT3 pathway to promote osteosarcoma cell growth and invasion. Hum. Cell 2019, 33, 252–260. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.; Liu, D.; Zhao, L.; Wang, L.; Xin, M.; Li, X. Long noncoding RNA MALAT1 alleviates lipopolysaccharide-induced inflammatory injury by upregulating microRNA-19b in murine chondrogenic ATDC5 cells. J. Cell. Biochem. 2018, 119, 10165–10175. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Wang, L.; Ali, T.; Li, L.; Bi, X.; Wang, J.; Lu, G.; Shao, Y.; Vuitton, D.A.; Wen, H.; et al. Hydatid cyst fluid promotes peri-cystic fibrosis in cystic echinococcosis by suppressing miR-19 expression. Parasit. Vectors 2016, 9, 278. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Zhang, C.; Wei, X.; Blagosklonov, O.; Lv, G.; Lu, X.; Mantion, G.; Vuitton, D.A.; Wen, H.; Lin, R. TGF-beta and TGF-beta/Smad signaling in the interactions between Echinococcus multilocularis and its hosts. PLoS ONE 2013, 8, e55379. [Google Scholar]
- Wang, J.; Lin, R.; Zhang, W.; Li, L.; Gottstein, B.; Blagosklonov, O.; Lü, G.; Zhang, C.; Lu, X.; Vuitton, D.A.; et al. Transcriptional Profiles of Cytokine/Chemokine Factors of Immune Cell-Homing to the Parasitic Lesions: A Comprehensive One-Year Course Study in the Liver of E. multilocularis-Infected Mice. PLoS ONE 2014, 9, e91638. [Google Scholar] [CrossRef] [Green Version]
- Kozomara, A.; Birgaoanu, M.; Griffiths-Jones, S. miRBase: From microRNA sequences to function. Nucleic Acids Res. 2019, 47, D155–D162. [Google Scholar] [CrossRef]
- Nawrocki, E.P.; Burge, S.W.; Bateman, A.; Daub, J.; Eberhardt, R.Y.; Eddy, S.R.; Floden, E.W.; Gardner, P.P.; Jones, T.A.; Tate, J.; et al. Rfam 12.0: Updates to the RNA families database. Nucleic Acids Res. 2015, 43, D130–D137. [Google Scholar] [CrossRef]
- Betel, D.; Koppal, A.; Agius, P.; Sander, C.; Leslie, C. Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome Boil. 2010, 11, R90. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miranda, K.C.; Huynh, T.; Tay, Y.; Ang, Y.-S.; Tam, W.-L.; Thomson, A.M.; Lim, B.; Rigoutsos, I. A Pattern-Based Method for the Identification of MicroRNA Binding Sites and Their Corresponding Heteroduplexes. Cell 2006, 126, 1203–1217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krüger, J.; Rehmsmeier, M. RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res. 2006, 34, W451–W454. [Google Scholar] [CrossRef] [PubMed]
- Vlachos, I.S.; Paraskevopoulou, M.D.; Karagkouni, D.; Georgakilas, G.; Vergoulis, T.; Kanellos, I.; Anastasopoulos, I.L.; Maniou, S.; Karathanou, K.; Kalfakakou, D.; et al. DIANA-TarBase v7.0: Indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res. 2015, 43, D153–D159. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhou, X.; Hao, L.; Piao, X.; Hou, N.; Chen, Q. Genome-Wide Transcriptome Analysis Reveals Extensive Alternative Splicing Events in the Protoscoleces of Echinococcus granulosus and Echinococcus multilocularis. Front Microbiol. 2017, 8, 929. [Google Scholar] [CrossRef] [PubMed]
- Shanghai Chinese Human Genome Center. Echinococcus Granulosus Small RNA Transcriptome; Yun Bai: Shanghai, China, 2013. [Google Scholar]
- Friedlander, M.R.; Mackowiak, S.D.; Li, N.; Chen, W.; Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012, 40, 37–52. [Google Scholar] [CrossRef] [PubMed]
- Roote, J.; Prokop, A. How to Design a Genetic Mating Scheme: A Basic Training Package for Drosophila Genetics. G3 Genes Genomes Genet. 2013, 3, 353–358. [Google Scholar]

| Species | Tissue | Source | Genotype | Clean Reads | Identified miRNAs | Technology | Country | Ref. |
|---|---|---|---|---|---|---|---|---|
| E. granulosus | Protoscoleces | Porcine liver hydatid cysts and bovine lung hydatid cysts | G1 | 182 | 34 conserved and four new candidate miRNAs | Sanger | Argentina and Uruguay | [45] |
| Protoscoleces | Pig | G7 | miR-125, miR-2, miR-71, miR-9, miR-10, let-7 and miR-277 | |||||
| Protoscoleces | Sheep | G1 | miR-125, miR-2, miR-71, miR-9, miR-10, let-7 and miR-277 | |||||
| Germinal layer of secondary cysts | Murine | G1 | miR-2, miR-71, miR-9, miR-10, let-7 and miR-277 | |||||
| Pre-microcysts | Porcine liver and bovine lung | G7 | miR-125, miR-2 | |||||
| Microcysts | Porcine liver and bovine lung | G7 | miR-71 | |||||
| Protoscoleces | Sheep liver | __ | 21,708,040 | 109 known and 260 novel miRNAs | Illumina HiSeqTM 250 | China | [47] | |
| Adult | Sheep liver hydatid cysts | __ | 10,069,724 | 46 known miRNAs, 92 novel mature miRNAs | Illumina Genome Analyzer II | China | [44] | |
| Protoscoleces | Sheep liver | 11,775,532 | 45 known miRNAs, 91 novel mature miRNAs | |||||
| Cyst membrane | Dog | 8,025,262 | 45 known miRNAs, 103 novel mature miRNAs | |||||
| Protoscoleces | Naturally infected sheep liver | G1 | 1,642,112/1,956,161 (two biological replicates) | 36 miRNAs | Illumina Genome Analyzer II | China | [12] | |
| E. canadensis | Protoscoleces | Naturally infected swine liver | G7 | 4,065,356/1,882,945 (two biological replicates) | 35 miRNAs | Illumina Genome Analyzer II | China | [12] |
| Cyst walls | 2,487,372/2,117,367 (two biological replicates) | 35 miRNAs | ||||||
| Cyst walls | Swine liver | 16,431,381/16,364,826 (two biological replicates) | 42 miRNAs | |||||
| E. multilocularis | Metacestodes | Female CF1 mice (6–8 weeks old) | __ | 24,703,158/20,396,074 (two biological replicates) | 37 miRNAs | High-throughput small RNA sequencing | Argentina | [1] |
| Metacestodes | Naturally infected crab-eating macaque (Macaca fascicularis) liver | __ | The complete E. multilocularis genome (https://www.genedb.org/#/species/Emultilocularis) | 24 miRNAs | Self-organizing map analysis | Northern Germany | [46] |
| MiRNA | Description of Target Genes | Biological Function | |
|---|---|---|---|
| MiR-71 | Nemo-like kinase | Involved in protoscolex development and regulates host macrophage functions in Echinococcus | [52,54] |
| Let-7 | Unknown | May be associated with the capability of E. granulosus for bi-directional development, can be significantly affected in the microcyst stage of E. granulosus and can exhibit different changes in expression in response to albendazole sulfoxide | [44,49] |
| MiR-61 | Unknown | Significantly affected in the microcyst stage of E. granulosus and exhibits different changes in expression in response to albendazole sulfoxide | [49] |
| MiR-10 | MAPKs; ECANG7_04447; ECANG7_01705; ECANG7_09658 | May be involved in regulating the MAPK and Wnt signalling pathways in Echinococcus | [23] |
| MiR-124 | ECANG7_04102; ECANG7_10164; ECANG7_00514; ECANG7_02390; ECANG7_01054 | May regulate development, host-parasite interactions, and stem cell pluripotency; related to the MAPK and TGF-beta signalling pathways in Echinococcus | [23] |
| MiR-184 | ECANG7_02390; ECANG7_09002 (casein kinase II); ECANG7_05735 (phosphatidylinositol phospholipase C gene); ECANG7_00867 (calcium/calmodulin-dependent protein kinase gene) | May act in a regulatory loop in miRNA biogenesis in Echinococcus | [23] |
| MiR-277 | ECANG7_01278; ECANG7_02522 | May be involved in the regulation of Wnt signalling pathways regulating the pluripotency of stem cells in Echinococcus | [23] |
| MiR-281 | ECANG7_04919 and ECANG7_00818 (glypicas); Nos-1 | Potentially involved in the developmental morphogenesis of Echinococcus. | [23] |
| MiR-2 | ECANG7_10172; ECANG7_02390; Nos-1; ECANG7_02601; ECANG7_05326 | Potentially involved in ubiquitin-mediated proteolysis and herpes simplex infection signalling pathways, TAFH/NHR1 transcription initiation, and segmentation in Echinococcus | [23] |
| MiR-307 | Ubiquitin-conjugating enzyme, E2 | May be involved in the ubiquitin-mediated proteolysis and herpes simplex infection signalling pathways in Echinococcus | [23] |
| MiR-7 | ECANG7_04919 and ECANG7_00818; ECANG7_03238 | Potentially involved in the developmental morphogenesis of Echinococcus | [23] |
| MiR-9 | ECANG7_02182; ECANG7_05944 | Bromodomain-containing protein is an orthologue of Caenorhabditis elegans lin-49, which is involved in nematode larval development | [23] |
| MiR-96 | ECANG7_06901 | Unknown, but has a high level of expression, particularly in the Echinococcus protoscolex stage. | [23] |
| MiR-125 | ECANG7_01292; ECANG7_01524 | May regulate developmental genes in Echinococcus | [23] |
| MiR-36 | Unknown | May correspond with the increased regenerative capacity of E. multilocularis with respect to that of E. granulosus s.l. | [1] |
| MiR-745 | G2:M phase-specific E3 ubiquitin protein ligase | Unknown | [23] |
| MiR-8 | Occludin/RNA polymerase II elongation factor, ELL domain RNA polymerase II elongation factor ELL | Unknown | [23] |
| MiR-87 | Zinc finger, C2H2 | Unknown | [23] |
| Bantam | Ribosomal protein S2 | Unknown | [23] |
| MiRNA | Expression Level | Target | Biological Function | Ref. | |||
|---|---|---|---|---|---|---|---|
| Sheep Gut | Mouse | ||||||
| Macrophage | Liver | Serum | |||||
| MiR-1247 | ↑ | - | - | ↓ | CircUBXN7 | Represses cell growth and invasion in human bladder cancer. | [71] |
| MiR-145 | ↑ | - | ↓ | - | ZEB2 | Increases the apoptosis of activated hepatic stellate cells induced by TRAIL via the NF-κB signalling pathway. | [72] |
| MiR-181 | ↑ | - | - | ↓ | Smad7 Hsa_circ_0007385 | Influences the differentiation of T helper cells and the activation of macrophages, controls T cell sensitivity to antigens during development | [73] |
| MiR-18 | ↑ | - | - | ↑ | Unknown | As the female immunity regulator, miR-18 controls the expression of A20/Tnfaip3 and exacerbating NF-κB-driven inflammation in fibroblast-like synoviocytes of rheumatoid arthritis | [74] |
| MiR-20 | ↑ | - | ↓ | - | ATG10 | Inhibits autophagy and chondrocyte proliferation by targeting ATG10 through the PI3K/AKT/mTOR signalling pathway. | [75] |
| MiR-21 | ↑ | ↑ | - | ↑ | Different targets engaged in each cell type and at each time point | As the one of the master regulators of innate immunity, miR-21 plays a myriad of roles in various cellular processes via regulating genes involved in signalling pathways, such as p53, FOXO1, TGF-α, apoptosis (PDCD4),P13K/Akt/mTOR, VEGF, and NF-αB | [69] |
| MiR-22 | ↑ | - | ↓ | - | CD147, YWHAZ | Inhibits hepatocellular carcinoma cell invasion, migration, and proliferation, miR-22 downregulation predicts poor survival. | [76,77] |
| MiR-223 | ↑ | - | ↓ | - | Ras-related protein Rab-1 (Rab1) | May promote apoptosis and suppress cell growth through Rab1-mediated mTOR activation in hepatocellular carcinoma cells. In addition, miR-223 is a biomarker of acute and chronic liver injury | [78,79] |
| MiR-27 | ↑ | - | ↓ | ↑ | GATA3, c-Rel, Smad2, Smad3, lncRNA-CIR | MiR-27 plays the important roles for safeguarding Treg-mediated immunological tolerance | [70,80] |
| MiR-30 | ↑ | ↑ | ↓ | ↓ | MyD88, lncRNA n379519, lncRNA CNALPTC1 | Inhibiting cytokine expression and TLR/MyD88 activation in THP-1 cells during Mycobacterium tuberculosis H37Rv infection | [62,81] |
| MiR-339 | ↑ | ↓ | - | ↑ | Skp2 | Binding to the 3′-UTR of Skp2 mRNA to inhibit the lung cancer cells proliferation | [82] |
| MiR-345 | ↑ | - | ↑ | ↓ | AKT2 | Regulates the cell cycle, apoptosis, and proliferation of acute myeloid leukaemia cells by targeting AKT2 | [83] |
| MiR-365 | ↑ | - | ↑ | ↑ | LncRNA MT1DP, Timp3 | The lncRNA MT1DP exacerbates cadmium-induced oxidative stress by suppressing the function of Nrf2 acting as ceRNA of miR-365. Then, miR-365 promotes diabetic retinopathy through inhibiting lncRNA Timp3 increasing oxidative stress | [63,84] |
| MiR-378 | ↑ | ↑ | ↑ | - | IRG1, lncGAPLINC | Acts as a prognostic marker and inhibits epithelial-mesenchymal transition in human glioma and acts as a molecular sponge of lncGAPLINC to stimulate gastric cancer cell proliferation | [64,85] |
| MiR-449 | ↑ | - | ↓ | - | LncARSR | LncARSR is competitively binding to miR-449 and thereby promoting sunitinib resistance in renal cancer | [65] |
| MiR-542 | ↑ | - | - | ↓ | SMAD | After activation of SMAD2/3 phosphorylation and the promotion of mitochondrial dysfunction, upregulated miR-542-3p/5p may reduce muscle atrophy in intensive care of patients | [86] |
| MiR-877 | ↑ | - | - | ↓ | Cyclin-dependent kinase 14 | Suppresses cell migration, invasion, and growth, and predicts prognosis in hepatocellular carcinoma | [66] |
| MiR-99 | ↑ | ↑ | - | - | SMARCA5 | Regulates Mycoplasma gallisepticum (HS strain) infection by suppressing cell proliferation in chickens | [87] |
| MiR-124 | ↑ | - | - | ↓ | LncHOTAIR, lncMALAT1, circMMP9 | LncHOTAIR sponged miRNA-124 to promote renal cell carcinoma malignancy through alpha-2,8-sialyltransferase 4. LncRNA MALAT1 acts as a ceRNA to control amadori-glycated albumin-induced MCP-1 expression in retinal microglia through a miRNA-124-dependent mechanism | [88,89,90] |
| MiR-130 | ↑ | - | - | ↓ | LncMRPL39 | LncMRPL39 inhibits gastric cancer progression and proliferation by directly binding to miR-130 | [67] |
| MiR-16 | ↑ | ↓ | LncDleu2 | LncDleu2 influences the invasion, migration, and proliferation of laryngeal cancer cells via miR-16 | [68] | ||
| Name | Website | Description | Reference |
|---|---|---|---|
| Wellcome Sanger institute | https://www.sanger.ac.uk/resources/downloads/helminths/echinococcus-multilocularis.html | Includes E. multilocularis and E. granulosus genomes | [14] |
| Sequence Read Archive | https://www.ncbi.nlm.nih.gov/sra/?term=Echinococcus%20miRNA | Includes Echinococcus miRNA raw sequencing data obtained by second-generation sequencing | [12,107,108] |
| miRBase | http://www.mirbase.org | A database of Echinococcus miRNA sequences and annotations | [101] |
| Rfam | http://rfam.xfam.org/search?q=Echinococcus | Includes known Echinococcus rRNAs, tRNAs, snRNAs and mRNAs | [102] |
| miRDeep2 | https://www.osc.edu/book/export/html/4389 | miRNA prediction | [109] |
| miRanda | http://www.microrna.org/microrna/home.do | Used to predict the target genes of all mature miRNAs | [103] |
| RNA22 | https://cm.jefferson.edu/rna22/ | Used for target predictions for multiple species | [104] |
| RNAhybrid | https://bibiserv.cebitec.uni-bielefeld.de/rnahybrid/ | Used to find the minimum free energy for hybridization of a long and a short RNA for predicting miRNA targets | [105] |
| TarBase v6.0 | http://diana.imis.athena-innovation.gr/DianaTools/index.php?r=tarbase/index | Includes experimentally verified interactions between miRNAs and target genes | [106] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
He, Z.; Yan, T.; Yuan, Y.; Yang, D.; Yang, G. miRNAs and lncRNAs in Echinococcus and Echinococcosis. Int. J. Mol. Sci. 2020, 21, 730. https://doi.org/10.3390/ijms21030730
He Z, Yan T, Yuan Y, Yang D, Yang G. miRNAs and lncRNAs in Echinococcus and Echinococcosis. International Journal of Molecular Sciences. 2020; 21(3):730. https://doi.org/10.3390/ijms21030730
Chicago/Turabian StyleHe, Zhi, Taiming Yan, Ya Yuan, Deying Yang, and Guangyou Yang. 2020. "miRNAs and lncRNAs in Echinococcus and Echinococcosis" International Journal of Molecular Sciences 21, no. 3: 730. https://doi.org/10.3390/ijms21030730




