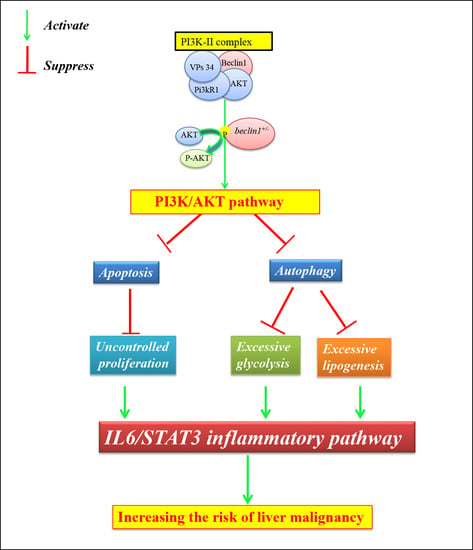
Strategy of Hepatic Metabolic Defects Induced by beclin1 Heterozygosity in Adult Zebrafish
Abstract
:1. Introduction
2. Results
2.1. CRISPR/Cas9-Mediated Targeted Mutagenesis of atg7 and beclin1 in Zebrafish
2.2. Beclin1 Heterozygosity Affects Liver Histology and Causes High Mortality Rate in Male Fish
2.3. Heterozygosity of beclin1 Alters Hepatic Energy Metabolism in Male Zebrafish
2.4. Beclin1+/− Affects Hepatic Glucose/Lipid Flux on the Genetic Level
2.5. Unlike atg7+/−, Heterozygosity of beclin1 Suppresses Autophagy via Activating PI3K/AKT Pathway
2.6. Induction of IL6/STAT3 Inflammatory Pathway and Downregulation of Apoptosis Pathway in beclin1+/− Liver
2.7. Heterozygosity of beclin1 Enhances Hepatocellular Proliferation and Inflammation Probably Due to Apoptosis Suppression
3. Discussion
4. Materials and Methods
4.1. Establishment of atg7 and beclin1 Mutant Lines and Calculation of Survival Rates
4.2. Assessments of Liver Histology
4.3. RNA Extraction and Quantitative Real-Time PCR (qRT-PCR)
4.4. Protein Extraction and Western Blotting
4.5. Immunohistochemistry, TUNEL Assay, and Immunofluorescence
4.6. Image Acquisition and Digital Image Analysis
4.7. Transcriptomic Sequencing (RNA-Seq) and Gene Ontology
4.8. Statistical Analysis and Graphs Preparation
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Rui, L. Energy metabolism in the liver. Compr. Physiol. 2014, 4, 177–197. [Google Scholar]
- Rabinowitz, J.D.; White, E. Autophagy and metabolism. Science 2010, 330, 1344–1348. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Madrigal-Matute, J.; Cuervo, A.M. Regulation of Liver Metabolism by Autophagy. Gastroenterology 2016, 150, 328–339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, Y.; Wang, H.; Wei, C.; Xiang, Y.; Liang, X.; Phang, C.W.; Jiao, R. HDAC6 regulates lipid droplet turnover in response to nutrient deprivation via p62-mediated selective autophagy. J. Genet. Genomics 2019, 46, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Shen, S.; Yang, L.; Li, L.; Bai, Y.; Liu, H. Lipid metabolism in mouse embryonic fibroblast cells in response to autophagy induced by nutrient stress. Anal. Chim. Acta 2018, 1037, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Farah, B.L.; Landau, D.J.; Sinha, R.A.; Brooks, E.D.; Wu, Y.; Fung, S.Y.S.; Tanaka, T.; Hirayama, M.; Bay, B.H.; Koeberl, D.D.; et al. Induction of autophagy improves hepatic lipid metabolism in glucose-6-phosphatase deficiency. J. Hepatol. 2016, 64, 370–379. [Google Scholar] [CrossRef] [PubMed]
- Han, S.L.; Wang, J.; Zhang, Y.X.; Qiao, F.; Chen, L.Q.; Zhang, M.L.; Du, Z.Y. Inhibited autophagy impairs systemic nutrient metabolism in Nile tilapia. Comp. Biochem. Physiol. A Mol. Integr. Physiol. 2019, 236, 110521. [Google Scholar] [CrossRef]
- Feng, Y.; He, D.; Yao, Z.; Klionsky, D.J. The machinery of macroautophagy. Cell Res. 2014, 24, 24–41. [Google Scholar] [CrossRef] [Green Version]
- Heras-Sandoval, D.; Perez-Rojas, J.M.; Hernandez-Damian, J.; Pedraza-Chaverri, J. The role of PI3K/AKT/mTOR pathway in the modulation of autophagy and the clearance of protein aggregates in neurodegeneration. Cell Signal. 2014, 26, 2694–2701. [Google Scholar] [CrossRef]
- Rubinsztein, D.C.; Shpilka, T.; Elazar, Z. Mechanisms of autophagosome biogenesis. Curr. Biol. 2012, 22, R29–R34. [Google Scholar] [CrossRef] [Green Version]
- Yue, Z.Y.; Jin, S.K.; Yang, C.W.; Levine, A.J.; Heintz, N. Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc. Natl. Acad. Sci. USA 2003, 100, 15077–15082. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Russell, S.E.H.; Hickey, G.I.; Lowry, W.S.; White, P.; Atkinson, R.J. Allele Loss from Chromosome-17 in Ovarian-Cancer. Oncogene 1990, 5, 1581–1583. [Google Scholar] [PubMed]
- Eccles, D.M.; Russell, S.E.H.; Haites, N.E.; Atkinson, R.; Bell, D.W.; Gruber, L.; Hickey, I.; Kelly, K.; Kitchener, H.; Leonard, R.; et al. Early Loss of Heterozygosity on 17q in Ovarian-Cancer. Oncogene 1992, 7, 2069–2072. [Google Scholar] [PubMed]
- Futreal, P.A.; Soderkvist, P.; Marks, J.R.; Iglehart, J.D.; Cochran, C.; Barrett, J.C.; Wiseman, R.W. Detection of Frequent Allelic Loss on Proximal Chromosome-17q in Sporadic Breast-Carcinoma Using Microsatellite Length Polymorphisms. Cancer Res. 1992, 52, 2624–2627. [Google Scholar]
- Saito, H.; Inazawa, J.; Saito, S.; Kasumi, F.; Koi, S.; Sagae, S.; Kudo, R.; Saito, J.; Noda, K.; Nakamura, Y. Detailed Deletion Mapping of Chromosome 17q in Ovarian and Breast Cancers—2-Cm Region on 17q21.3 Often and Commonly Deleted in Tumors. Cancer Res. 1993, 53, 3382–3385. [Google Scholar]
- Gao, X.; Zacharek, A.; Salkowski, A.; Grignon, D.J.; Sakr, W.; Porter, A.T.; Honn, K.V. Loss of heterozygosity of the BRCA1 and other loci on chromosome 17q in human prostate cancer. Cancer Res. 1995, 55, 1002–1005. [Google Scholar]
- Liang, X.H.; Jackson, S.; Seaman, M.; Brown, K.; Kempkes, B.; Hibshoosh, H.; Levine, B. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 1999, 402, 672–676. [Google Scholar] [CrossRef]
- Qu, X.; Yu, J.; Bhagat, G.; Furuya, N.; Hibshoosh, H.; Troxel, A.; Rosen, J.; Eskelinen, E.L.; Mizushima, N.; Ohsumi, Y.; et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J. Clin. Investig. 2003, 112, 1809–1820. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Li, H.; Spitsbergen, J.M.; Gong, Z. Males develop faster and more severe hepatocellular carcinoma than females in kras(V12) transgenic zebrafish. Sci. Rep. 2017, 7, 41280. [Google Scholar] [CrossRef]
- Fresno Vara, J.A.; Casado, E.; de Castro, J.; Cejas, P.; Belda-Iniesta, C.; Gonzalez-Baron, M. PI3K/Akt signalling pathway and cancer. Cancer Treat. Rev. 2004, 30, 193–204. [Google Scholar] [CrossRef]
- Golob-Schwarzl, N.; Krassnig, S.; Toeglhofer, A.M.; Park, Y.N.; Gogg-Kamerer, M.; Vierlinger, K.; Schroder, F.; Rhee, H.; Schicho, R.; Fickert, P.; et al. New liver cancer biomarkers: PI3K/AKT/mTOR pathway members and eukaryotic translation initiation factors. Eur. J. Cancer 2017, 83, 56–70. [Google Scholar] [CrossRef]
- Ouyang, L.; Shi, Z.; Zhao, S.; Wang, F.T.; Zhou, T.T.; Liu, B.; Bao, J.K. Programmed cell death pathways in cancer: A review of apoptosis, autophagy and programmed necrosis. Cell Prolif. 2012, 45, 487–498. [Google Scholar] [CrossRef]
- Mizushima, N. Autophagy: Process and function. Genes Dev. 2007, 21, 2861–2873. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elmore, S. Apoptosis: A review of programmed cell death. Toxicol. Pathol. 2007, 35, 495–516. [Google Scholar] [CrossRef]
- Gunther, C.; Neumann, H.; Neurath, M.F.; Becker, C. Apoptosis, necrosis and necroptosis: Cell death regulation in the intestinal epithelium. Gut 2013, 62, 1062–1071. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nikoletopoulou, V.; Markaki, M.; Palikaras, K.; Tavernarakis, N. Crosstalk between apoptosis, necrosis and autophagy. BBA-Mol. Cell Res. 2013, 1833, 3448–3459. [Google Scholar] [CrossRef] [Green Version]
- Macintosh, R.L.; Timpson, P.; Thorburn, J.; Anderson, K.I.; Thorburn, A.; Ryan, K.M. Inhibition of autophagy impairs tumor cell invasion in an organotypic model. Cell Cycle 2012, 11, 2022–2029. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hanel, W.; Marchenko, N.; Xu, S.; Yu, S.X.F.; Weng, W.; Moll, U. Two hot spot mutant p53 mouse models display differential gain of function in tumorigenesis. Cell Death Differ. 2013, 20, 898–909. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muller, P.A.J.; Caswell, P.T.; Doyle, B.; Iwanicki, M.P.; Tan, E.H.; Karim, S.; Lukashchuk, N.; Gillespie, D.A.; Ludwig, R.L.; Gosselin, P.; et al. Mutant p53 Drives Invasion by Promoting Integrin Recycling. Cell 2009, 139, 1327–1341. [Google Scholar] [CrossRef]
- Kurnia, I.; Siregar, B.; Soetopo, S.; Ramli, I.; Kurjana, T.; Andriono, T.K.; Tobing, M.D.L.; Suryawathi, B.; Kisnanto, T.; Tetriana, D. Correlation Between Akt and p53 Protein Expression and Chemoradiotherapy Response in Cervical Cancer Patients. Hayati J. Biosci. 2014, 21, 173–179. [Google Scholar] [CrossRef] [Green Version]
- Morselli, E.; Tasdemir, E.; Maiuri, M.C.; Galluzzi, L.; Kepp, O.; Criollo, A.; Vicencio, J.M.; Soussi, T.; Kroemer, G. Mutant p53 protein localized in the cytoplasm inhibits autophagy. Cell Cycle 2008, 7, 3056–3061. [Google Scholar] [CrossRef] [PubMed]
- Blandino, G.; Valenti, F.; Sacconi, A.; Di Agostino, S. Wild type- and mutant p53 proteins in mitochondrial dysfunction: Emerging insights in cancer disease. Semin. Cell Dev. Biol. 2020, 98, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Petitjean, A.; Achatz, M.I.; Borresen-Dale, A.L.; Hainaut, P.; Olivier, M. TP53 mutations in human cancers: Functional selection and impact on cancer prognosis and outcomes. Oncogene 2007, 26, 2157–2165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Degenhardt, K.; Mathew, R.; Beaudoin, B.; Bray, K.; Anderson, D.; Chen, G.H.; Mukherjee, C.; Shi, Y.F.; Gelinas, C.; Fan, Y.J.; et al. Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell 2006, 10, 51–64. [Google Scholar] [CrossRef] [Green Version]
- Fesus, L.; Demeny, M.A.; Petrovski, G. Autophagy Shapes Inflammation. Antioxid. Redox Sign. 2011, 14, 2233–2243. [Google Scholar] [CrossRef]
- Zhou, J.; Jin, J.O.; Patel, E.S.; Yu, Q. Interleukin-6 inhibits apoptosis of exocrine gland tissues under inflammatory conditions. Cytokine 2015, 76, 244–252. [Google Scholar] [CrossRef] [Green Version]
- Grivennikov, S.; Karin, E.; Terzic, J.; Mucida, D.; Yu, G.Y.; Vallabhapurapu, S.; Scheller, J.; Rose-John, S.; Cheroutre, H.; Eckmann, L.; et al. IL-6 and Stat3 Are Required for Survival of Intestinal Epithelial Cells and Development of Colitis-Associated Cancer (vol 15, pg 103, 2009). Cancer Cell 2009, 15, 241. [Google Scholar] [CrossRef] [Green Version]
- Hodge, D.R.; Hurt, E.M.; Farrar, W.L. The role of IL-6 and STAT3 in inflammation and cancer. Eur. J. Cancer 2005, 41, 2502–2512. [Google Scholar] [CrossRef]
- Slinger, E.; Maussang, D.; Schreiber, A.; Siderius, M.; Rahbar, A.; Fraile-Ramos, A.; Lira, S.A.; Soderberg-Naucler, C.; Smit, M.J. HCMV-Encoded Chemokine Receptor US28 Mediates Proliferative Signaling Through the IL-6-STAT3 Axis. Sci. Signal. 2010, 3, ra58. [Google Scholar] [CrossRef]
- Zhang, L.; Yang, B.; Li, X.L.; Zhang, Y.J.; Zhao, J.P.; Wang, W.B.; Yu, X.H.; Zhai, Z.H.; Sun, H.Z. The targeting of endoglin on vascular endothelial cells affects the infiltration of M2 macrophages into the breast cancer microenvironment by modulating the interleukin-6 (IL-6) level. Transl. Cancer Res. 2018, 7, 912–921. [Google Scholar] [CrossRef]
- Qi, Y.L.; Zhang, M.F.; Li, H.; Frank, J.A.; Dai, L.; Liu, H.J.; Zhang, Z.; Wang, C.; Chen, G. Autophagy Inhibition by Sustained Overproduction of IL6 Contributes to Arsenic Carcinogenesis. Cancer Res. 2014, 74, 3740–3752. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qin, B.B.; Zhou, Z.; He, J.Q.; Yan, C.L.; Ding, S.P. IL-6 Inhibits Starvation-induced Autophagy via the STAT3/Bcl-2 Signaling Pathway. Sci. Rep. 2015, 5, 15701. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Howe, K.; Clark, M.D.; Torroja, C.F.; Torrance, J.; Berthelot, C.; Muffato, M.; Collins, J.E.; Humphray, S.; McLaren, K.; Matthews, L.; et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 2013, 496, 498–503. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lam, S.H.; Wu, Y.L.; Vega, V.B.; Miller, L.D.; Spitsbergen, J.; Tong, Y.; Zhan, H.Q.; Govindarajan, K.R.; Lee, S.; Mathavan, S.; et al. Conservation of gene expression signatures between zebrafish and human liver tumors and tumor progression. Nat. Biotechnol. 2006, 24, 73–75. [Google Scholar] [CrossRef] [PubMed]
- Shang, R.Z.; Qu, S.B.; Wang, D.S. Reprogramming of glucose metabolism in hepatocellular carcinoma: Progress and prospects. World J. Gastroenterol. 2016, 22, 9933–9943. [Google Scholar] [CrossRef]
- Li, C.; Yang, W.; Zhang, J.L.; Zheng, X.; Yao, Y.M.; Tu, K.S.; Liu, Q.G. SREBP-1 Has a Prognostic Role and Contributes to Invasion and Metastasis in Human Hepatocellular Carcinoma. Int. J. Mol. Sci. 2014, 15, 7124–7138. [Google Scholar] [CrossRef]
- Christiaens, V.; Van Hul, M.; Lijnen, H.R.; Scroyen, I. CD36 promotes adipocyte differentiation and adipogenesis. BBA-Gen. Subj. 2012, 1820, 949–956. [Google Scholar] [CrossRef]
- Zhao, T.; Wu, K.; Hogstrand, C.; Xu, Y.H.; Chen, G.H.; Wei, C.C.; Luo, Z. Lipophagy mediated carbohydrate-induced changes of lipid metabolism via oxidative stress, endoplasmic reticulum (ER) stress and ChREBP/PPARgamma pathways. Cell Mol. Life Sci. 2019. [Google Scholar] [CrossRef]
- Vivanco, I.; Sawyers, C.L. The phosphatidylinositol 3-Kinase AKT pathway in human cancer. Nat. Rev. Cancer 2002, 2, 489–501. [Google Scholar] [CrossRef]
- Steelman, L.S.; Abrams, S.L.; Whelan, J.; Bertrand, F.E.; Ludwig, D.E.; Basecke, J.; Libra, M.; Stivala, F.; Milella, M.; Tafuri, A.; et al. Contributions of the Raf/MEK/ERK, PI3K/PTEN/Akt/mTOR and Jak/STAT pathways to leukemia. Leukemia 2008, 22, 686–707. [Google Scholar] [CrossRef] [Green Version]
- Komatsu, M.; Waguri, S.; Ueno, T.; Iwata, J.; Murata, S.; Tanida, I.; Ezaki, J.; Mizushima, N.; Ohsumi, Y.; Uchiyama, Y. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol 2005, 169, 425–434. [Google Scholar] [CrossRef] [PubMed]
- Takamura, A.; Komatsu, M.; Hara, T.; Sakamoto, A.; Kishi, C.; Waguri, S.; Eishi, Y.; Hino, O.; Tanaka, K.; Mizushima, N. Autophagy-deficient mice develop multiple liver tumors. Genes Dev. 2011, 25, 795–800. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rohatgi, R. Autophagy-Independent Role for Beclin 1 in the Regulation of Growth Factor Receptor Signaling: A Dissertation. Ph.D. Thesis, University of Massachusetts Medical School, Worcester, MA, USA, January 2015. [Google Scholar] [CrossRef]
- Coloff, J.L.; Mason, E.F.; Altman, B.J.; Gerriets, V.A.; Liu, T.Y.; Nichols, A.N.; Zhao, Y.X.; Wofford, J.A.; Jacobs, S.R.; Ilkayeva, O.; et al. Akt Requires Glucose Metabolism to Suppress Puma Expression and Prevent Apoptosis of Leukemic T Cells. J. Biol. Chem. 2011, 286, 5921–5933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tangkijvanich, P.; Mahachai, V.; Suwangool, P.; Poovorawan, Y. Gender difference in clinicopathologic features and survival of patients with hepatocellular carcinoma. World J. Gastroenterol. 2004, 10, 1547–1550. [Google Scholar] [CrossRef] [PubMed]
- Dohmen, K.; Shigematsu, H.; Irie, K.; Ishibashi, H. Longer survival in female than male with hepatocellular carcinoma. J. Gastroen. Hepatol. 2003, 18, 267–272. [Google Scholar] [CrossRef] [PubMed]
- Mei, J.; Gui, J.F. Genetic basis and biotechnological manipulation of sexual dimorphism and sex determination in fish. Sci. China Life Sci. 2015, 58, 124–136. [Google Scholar] [CrossRef] [Green Version]
- Bollrath, J.; Phesse, T.J.; von Burstin, V.A.; Putoczki, T.; Bennecke, M.; Bateman, T.; Nebelsiek, T.; Lundgren-May, T.; Canli, O.; Schwitalla, S.; et al. gp130-Mediated Stat3 Activation in Enterocytes Regulates Cell Survival and Cell-Cycle Progression during Colitis-Associated Tumorigenesis. Cancer Cell 2009, 15, 91–102. [Google Scholar] [CrossRef] [Green Version]
- Rohatgi, R.A.; Shaw, L.M. An autophagy-independent function for Beclin 1 in cancer. Mol. Cell Oncol. 2016, 3, e1030539. [Google Scholar] [CrossRef] [Green Version]
- Ganapathy-Kanniappan, S.; Geschwind, J.F.H. Tumor glycolysis as a target for cancer therapy: Progress and prospects. Mol. Cancer 2013, 12, 152. [Google Scholar] [CrossRef] [Green Version]
- Yamashita, T.; Honda, M.; Takatori, H.; Nishino, R.; Minato, H.; Takamura, H.; Ohta, T.; Kaneko, S. Activation of lipogenic pathway correlates with cell proliferation and poor prognosis in hepatocellular carcinoma. J. Hepatol. 2009, 50, 100–110. [Google Scholar] [CrossRef]
- Zaidi, N.; Lupien, L.; Kuemmerle, N.B.; Kinlaw, W.B.; Swinnen, J.V.; Smans, K. Lipogenesis and lipolysis: The pathways exploited by the cancer cells to acquire fatty acids. Prog. Lipid Res. 2013, 52, 585–589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DeFilippis, R.A.; Chang, H.; Dumont, N.; Rabban, J.T.; Chen, Y.Y.; Fontenay, G.V.; Berman, H.K.; Gauthier, M.L.; Zhao, J.X.; Hu, D.L.; et al. CD36 Repression Activates a Multicellular Stromal Program Shared by High Mammographic Density and Tumor Tissues. Cancer Discov. 2012, 2, 826–839. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gudkov, A.V.; Gurova, K.V.; Komarova, E.A. Inflammation and p53: A Tale of Two Stresses. Genes Cancer 2011, 2, 503–516. [Google Scholar] [CrossRef] [PubMed]
- Sugawara, K.; Suzuki, N.N.; Fujioka, Y.; Mizushima, N.; Ohsumi, Y.; Inagaki, F. The crystal structure of microtubule-associated protein light chain 3, a mammalian homologue of Saccharomyces cerevisiae Atg8. Genes Cells 2004, 9, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Bjorkoy, G.; Lamark, T.; Pankiv, S.; Overvatn, A.; Brech, A.; Johansen, T. Monitoring Autophagic Degradation of P62/Sqstm1. Method Enzymol. 2009, 452, 181–197. [Google Scholar]
- Westerfield, M. The Zebrafish Book: A Guide for the Laboratory Use of Zebrafish (Brachydanio Rerio); University of Oregon Press: Eugene, OR, USA, 1995. [Google Scholar]
- Goel, M.K.; Khanna, P.; Kishore, J. Understanding survival analysis: Kaplan-Meier estimate. Int. J. Ayurveda Res. 2010, 1, 274–278. [Google Scholar]
- Van der Velden, Y.U.; Wang, L.Q.; Zevenhoven, J.; van Rooijen, E.; van Lohuizen, M.; Giles, R.H.; Clevers, H.; Haramis, A.P.G. The serine-threonine kinase LKB1 is essential for survival under energetic stress in zebrafish. Proc. Natl. Acad. Sci. USA 2011, 108, 4358–4363. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Zhang, X.M.; Liu, Y.J.; Su, Z.H.; Dawar, F.U.; Dan, H.; He, Y.; Gui, J.F.; Mei, J. Leucine mediates autophagosome-lysosome fusion and improves sperm motility by activating the PI3K/Akt pathway. Oncotarget 2017, 8, 111807–111818. [Google Scholar] [CrossRef] [Green Version]
- Dan, C.; Lin, Q.H.; Gong, G.R.; Yang, T.Y.; Xiong, S.T.; Xiong, Y.; Huang, P.P.; Gui, J.F.; Mei, J. A novel PDZ domain-containing gene is essential for male sex differentiation and maintenance in yellow catfish (Pelteobagrus fulvidraco). Sci. Bull. 2018, 63, 1420–1430. [Google Scholar] [CrossRef] [Green Version]
- Pulkkanen, K.J.; Laukkanen, M.O.; Naarala, J.; Yla-Herttuala, S. False-positive apoptosis signal in mouse kidney and liver detected with TUNEL assay. Apoptosis 2000, 5, 329–333. [Google Scholar] [CrossRef]
- MacAulay, C.; Keyes, M.; Hayes, M.; Lo, A.; Wang, G.; Guillaud, M.; Gleave, M.; Fazli, L.; Korbelik, J.; Collins, C.; et al. Quantification of large scale DNA organization for predicting prostate cancer recurrence. Cytom. Part A 2017, 91a, 1164–1174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trapnell, C.; Hendrickson, D.G.; Sauvageau, M.; Goff, L.; Rinn, J.L.; Pachter, L. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 2013, 31, 46–53. [Google Scholar] [CrossRef] [PubMed]







© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mawed, S.A.; He, Y.; Zhang, J.; Mei, J. Strategy of Hepatic Metabolic Defects Induced by beclin1 Heterozygosity in Adult Zebrafish. Int. J. Mol. Sci. 2020, 21, 1533. https://doi.org/10.3390/ijms21041533
Mawed SA, He Y, Zhang J, Mei J. Strategy of Hepatic Metabolic Defects Induced by beclin1 Heterozygosity in Adult Zebrafish. International Journal of Molecular Sciences. 2020; 21(4):1533. https://doi.org/10.3390/ijms21041533
Chicago/Turabian StyleMawed, Suzan Attia, Yan He, Jin Zhang, and Jie Mei. 2020. "Strategy of Hepatic Metabolic Defects Induced by beclin1 Heterozygosity in Adult Zebrafish" International Journal of Molecular Sciences 21, no. 4: 1533. https://doi.org/10.3390/ijms21041533