Ralstonia solanacearum elicitor RipX Induces Defense Reaction by Suppressing the Mitochondrial atpA Gene in Host Plant
Abstract
:1. Introduction
2. Results
2.1. Expression of RipX1 or RipX3 Induces a Defense Reaction in N. benthamiana
2.2. External Application of RipX1 and RipX3 Proteins Induces Macroscopic HR Reactions
2.3. Isolation and Phylogenetic Analysis of RipX-Interacting Protein ATPA from N. benthamiana
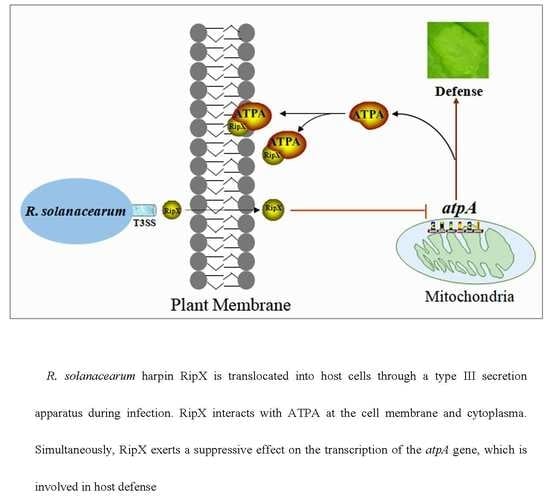
2.4. RipX Interacts with ATPA at the Cell Membrane and Cytoplasm
2.5. Silencing of atpA Gene in N. benthamiana Has No Effect on HR Induction by RipX but Improves Resistance to R. solanacearum
2.6. Expression of RipX1 Suppresses the Transcription of atpA Gene
3. Discussion
4. Materials and Methods
4.1. Plant Material and Cultivation
4.2. Bacterial Strains and Plasmids
4.3. DNA Manipulation
4.4. Agrobacterium-Mediated Transient Expression
4.5. Histochemical Staining
4.6. Confocal Microscopy
4.7. Quantitative Real-Time PCR (qRT-PCR)
4.8. Examination of the HR Reaction Induced by RipX
4.9. Y2H
4.10. Split-Luciferase Complementation Assay
4.11. Virus-Induced Gene Silencing (VIGS)
4.12. Resistance of N. benthamiana to Bacterial Wilt
4.13. Promoter Activity Assays
4.14. Western Blot Analysis
4.15. Sequence and Data Analysis
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Genin, S.; Denny, T.P. Pathogenomics of the Ralstonia solanacearum species complex. Ann. Rev. Phytopathol. 2012, 50, 67–89. [Google Scholar] [CrossRef]
- Coll, N.S.; Valls, M. Current knowledge on the Ralstonia solanacearum type III secretion system. Microb. Biotechnol. 2013, 6, 614–620. [Google Scholar] [PubMed] [Green Version]
- Deslandes, L.; Genin, S. Opening the Ralstonia solanacearum type III effector tool box: Insights into host cell subversion mechanisms. Curr. Opin. Plant. Biol. 2014, 20, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Cunnac, S.; Boucher, C.; Genin, S. Characterization of the cis-acting regulatory element controlling HrpB-mediated activation of the type III secretion system and effector genes in Ralstonia solanacearum. J. Bacteriol. 2004, 186, 2309–2318. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mukaihara, T.; Tamura, N.; Iwabuchi, M. Genome-wide identification of a large repertoire of Ralstonia solanacearum type III effector proteins by a new functional screen. Mol. Plant. Microbe Interact. 2010, 23, 251–262. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lavie, M.; Shillington, E.; Eguiluz, C.; Grimsley, N.; Boucher, C. PopP1, a new member of the YopJ/AvrRxv family of type III effector proteins, acts as a host-specificity factor and modulates aggressiveness of Ralstonia solanacearum. Mol. Plant. Microbe Interact. 2002, 15, 1058–1068. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deslandes, L.; Olivier, J.; Peeters, N.; Feng, D.X.; Khounlotham, M.; Boucher, C.; Somssich, I.; Genin, S.; Marco, Y. Physical interaction between RRS1-R, a protein conferring resistance to, bacterial wilt, and PopP2, a type III effector targeted to the plant nucleus. Proc. Natl. Acad. Sci. USA 2003, 100, 8024–8029. [Google Scholar] [CrossRef] [Green Version]
- Poueymiro, M.; Cunnac, S.; Barberis, P.; Deslandes, L.; Peeters, N.; Cazale-Noel, A.C.; Boucher, C.; Genin, S. Two type III secretion system effectors from Ralstonia solanacearum GMI1000 determine host-range specificity on tobacco. Mol. Plant. Microbe Interact. 2009, 22, 538–550. [Google Scholar] [CrossRef] [Green Version]
- Mukaihara, T.; Hatanaka, T.; Nakano, M.; Oda, K. Ralstonia solanacearum type III effector RipAY is a glutathione-degrading enzyme that is activated by plant cytosolic thioredoxins and suppresses plant immunity. mBio 2016, 7, e00359. [Google Scholar] [CrossRef] [Green Version]
- Nakano, M.; Oda, K.; Mukaihara, T. Ralstonia solanacearum novel E3 ubiquitin ligase (NEL) effectors RipAW and RipAR suppress pattern-triggered immunity in plants. Microbiology 2017, 163, 992–1002. [Google Scholar] [CrossRef]
- Nakano, M.; Mukaihara, T. Ralstonia solanacearum type III effector RipAL targets chloroplasts and induces jasmonic acid production to suppress salicylic acid-mediated defense responses in plants. Plant. Cell Physiol. 2018, 59, 2576–2589. [Google Scholar] [CrossRef]
- Nahar, K.; Matsumoto, I.; Taguchi, F.; Inagaki, Y.; Yamamoto, M.; Toyoda, K.; Shiraishi, T.; Ichinose, Y.; Mukaihara, T. Ralstonia solanacearum type III secretion system effector Rip36 induces a hypersensitive response in the nonhost wild eggplant Solanum torvum. Mol. Plant. Pathol. 2014, 15, 297–303. [Google Scholar] [CrossRef]
- Arlat, M.; Van Gijsegem, F.; Huet, J.C.; Pernollet, J.C.; Boucher, C.A. PopA1, a protein which induces a hypersensitivity-like response on specific Petunia genotypes, is secreted via the Hrp pathway of Pseudomonas solanacearum. EMBO J. 1994, 13, 543–553. [Google Scholar] [CrossRef] [PubMed]
- Peeters, N.; Carrère, S.; Anisimova, M.; Plener, L.; Cazalé, A.-C.; Genin, S. Repertoire, unified nomenclature and evolution of the Type III effector gene set in the Ralstonia solanacearum species complex. BMC Genomics 2013, 14, 859. [Google Scholar] [CrossRef] [PubMed]
- Guéneron, M.; Timmers, A.C.; Boucher, C.; Arlat, M. Two novel proteins, PopB, which has functional nuclear localization signals, and PopC, which has a large leucine-rich repeat domain, are secreted through the hrp-secretion apparatus of Ralstonia solanacearum. Mol. Microbiol. 2000, 36, 261–277. [Google Scholar] [CrossRef] [PubMed]
- Belbahri, L.; Boucher, C.; Candresse, T.; Nicole, M.; Ricci, P.; Keller, H. A local accumulation of the Ralstonia solanacearum PopA protein in transgenic tobacco renders a compatible plant-pathogen interaction incompatible. Plant. J. 2001, 28, 419–430. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Racapé, J.; Belbahri, L.; Engelhardt, S.; Lacombe, B.; Lee, J.; Lochman, J.; Marais, A.; Nicole, M.; Nurnberger, T.; Parlange, F.; et al. Ca2+-dependent lipid binding and membrane integration of PopA, a harpin-like elicitor of the hypersensitive response in tobacco. Mol. Microbiol. 2005, 58, 1406–1420. [Google Scholar] [CrossRef] [PubMed]
- Müller, V.; Grüber, G. ATP synthases: Structure, function and evolution of unique energy converters. Cell. Mol. Life Sci. 2003, 60, 474–494. [Google Scholar] [CrossRef]
- Rühle, T.; Leister, D. Assembly of F1F0-ATP synthases. Biochim. Biophy. Acta 2015, 1847, 849–860. [Google Scholar] [CrossRef] [Green Version]
- Ackerman, S.H.; Martin, J.; Tzagoloff, A. Characterization of ATP11 and detection of the encoded protein in mitochondria of Saccharomyces cerevisiae. J. Biol. Chem. 1992, 267, 7386–7394. [Google Scholar]
- Bowman, S.; Ackerman, S.H.; Griffiths, D.E.; Tzagoloff, A. Characterization of ATP12, a yeast nuclear gene required for the assembly of the mitochondrial F1-ATPase. J. Biol. Chem. 1991, 266, 7517–7523. [Google Scholar] [PubMed]
- Ashutosh; Kumar, P.; Dinesh Kumar, V.; Sharma, P.C.; Prakash, S.; Bhat, S.R. A novel orf108 co-transcribed with the atpA gene is associated with cytoplasmic male sterility in Brassica juncea carrying Moricandia arvensis cytoplasm. Plant. Cell Physiol. 2008, 49, 284–289. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wen, L.; Ruesch, K.L.; Ortega, V.M.; Kamps, T.L.; Gabay-Laughnan, S.; Chase, C.D. A nuclear restorer-of-fertility mutation disrupts accumulation of mitochondrial ATP synthase subunit alpha in developing pollen of S male-sterile maize. Genetics 2003, 165, 771–779. [Google Scholar] [PubMed]
- Choi, M.S.; Kim, W.; Lee, C.; Oh, C.S. Harpins, multifunctional proteins secreted by gram-negative plant-pathogenic bacteria. Mol. Plant. Microbe Interact. 2013, 26, 1115–1122. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, H.; Peng, J.; Bao, Z.; Meng, X.; Bonasera, J.M.; Chen, G.; Beer, S.V.; Dong, H. Downstream divergence of the ethylene signaling pathway for harpin-stimulated Arabidopsis growth and insect defense. Plant. Physiol. 2004, 136, 3628–3638. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ren, X.; Liu, F.; Bao, Z.; Zhang, C.; Wu, X.; Chen, L.; Liu, R.; Dong, H. Root growth of Arabidopsis thaliana is regulated by ethylene and abscisic acid signaling interaction in response to HrpNEa, a bacterial protein of harpin group. Plant. Mol. Biol. Report 2008, 26, 225–240. [Google Scholar] [CrossRef]
- Xie, Z.; Chen, Z. Harpin-induced hypersensitive cell death is associated with altered mitochondrial functions in tobacco cells. Mol. Plant. Microbe Interact. 2000, 13, 183–190. [Google Scholar] [CrossRef] [Green Version]
- Sripriya, P.; Vedantam, L.V.; Podile, A.R. Involvement of mitochondria and metacaspase elevation in harpinPss-induced cell death of Saccharomyces cerevisiae. J. Cell. Biochem. 2009, 107, 1150–1159. [Google Scholar] [CrossRef]
- Wei, Z.M.; Laby, R.J.; Zumoff, C.H.; Bauer, D.W.; He, S.Y.; Collmer, A.; Beer, S.V. Harpin, elicitor of the hypersensitive response produced by the plant pathogen Erwinia amylovora. Science 1992, 257, 85–88. [Google Scholar] [CrossRef]
- Oh, C.S.; Beer, S.V. AtHIPM, an ortholog of the apple HrpN-interacting protein, is a negative regulator of plant growth and mediates the growth-enhancing effect of HrpN in Arabidopsis. Plant. Physiol. 2007, 145, 426–436. [Google Scholar] [CrossRef] [Green Version]
- Campa, M.; Piazza, S.; Righetti, L.; Oh, C.S.; Conterno, L.; Borejsza-Wysocka, E.; Nagamangala, K.C.; Beer, S.V.; Aldwinckle, H.S.; Malnoy, M. HIPM is a susceptibility gene of Malus spp.: Reduced expression reduces susceptibility to Erwinia amylovora. Mol. Plant. Microbe Interact. 2019, 32, 167–175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miao, W.; Wang, X.; Song, C.; Wang, Y.; Ren, Y.; Wang, J. Transcriptome analysis of Hpa1Xoo transformed cotton revealed constitutive expression of genes in multiple signaling pathways related to disease resistance. J. Exp. Bot. 2010, 61, 4263–4275. [Google Scholar] [CrossRef] [PubMed]
- Bocsanczy, A.M.; Nissinen, R.M.; Oh, C.S.; Beer, S.V. HrpN of Erwinia amylovora functions in the translocation of DspA/E into plant cells. Mol. Plant. Pathol. 2008, 9, 425–434. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Chen, H.; Cai, T.; Deng, Y.; Zhuang, R.; Zhang, N.; Zeng, Y.; Zheng, Y.; Tang, R.; Pan, R.; et al. Overexpression of a novel peanut NBS-LRR gene AhRRS5 enhances disease resistance to Ralstonia solanacearum in tobacco. Plant. Biotechnol. J. 2017, 15, 39–55. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 1989. [Google Scholar]
- Wu, W.; Zhao, Z.; Luo, X.; Fan, X.; Zhuo, T.; Hu, X.; Liu, J.; Zou, H. Response regulator VemR regulates the transcription of flagellar rod gene flgG by interacting with σ54 factor RpoN2 in Xanthomonas citri ssp. citri. Mol. Plant. Pathol. 2019, 20, 372–381. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, H.; Zou, Y.; Shang, Y.; Lin, H.; Wang, Y.; Cai, R.; Tang, X.; Zhou, J.M. Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant. Physiol. 2008, 146, 368–376. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Schiff, M.; Dinesh-Kumar, S.P. Tobacco Rar1, EDS1 and NPR1/NIM1 like genes are required for N-mediated resistance to tobacco mosaic virus. Plant. J. 2008, 30, 415–429. [Google Scholar] [CrossRef]
- Sun, D.; Zhuo, T.; Hu, X.; Fan, X.; Zou, H. Identification of a Pseudomonas putida as biocontrol agent for tomato bacterial wilt disease. Biological Control. 2017, 114, 45–50. [Google Scholar] [CrossRef]
- Martinez-Garcia, J.F.; Monte, E.; Quail, P.H. A simple, rapid and quantitative method for preparing Arabidopsis protein extracts for immunoblot analysis. Plant. J. 1999, 20, 251–257. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [Green Version]






© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sun, T.; Wu, W.; Wu, H.; Rou, W.; Zhou, Y.; Zhuo, T.; Fan, X.; Hu, X.; Zou, H. Ralstonia solanacearum elicitor RipX Induces Defense Reaction by Suppressing the Mitochondrial atpA Gene in Host Plant. Int. J. Mol. Sci. 2020, 21, 2000. https://doi.org/10.3390/ijms21062000
Sun T, Wu W, Wu H, Rou W, Zhou Y, Zhuo T, Fan X, Hu X, Zou H. Ralstonia solanacearum elicitor RipX Induces Defense Reaction by Suppressing the Mitochondrial atpA Gene in Host Plant. International Journal of Molecular Sciences. 2020; 21(6):2000. https://doi.org/10.3390/ijms21062000
Chicago/Turabian StyleSun, Tingyan, Wei Wu, Haoxiang Wu, Wei Rou, Yinghui Zhou, Tao Zhuo, Xiaojing Fan, Xun Hu, and Huasong Zou. 2020. "Ralstonia solanacearum elicitor RipX Induces Defense Reaction by Suppressing the Mitochondrial atpA Gene in Host Plant" International Journal of Molecular Sciences 21, no. 6: 2000. https://doi.org/10.3390/ijms21062000