Molecular Characterization of Tc964, A Novel Antigenic Protein from Trypanosoma cruzi
Abstract
:1. Introduction
2. Results
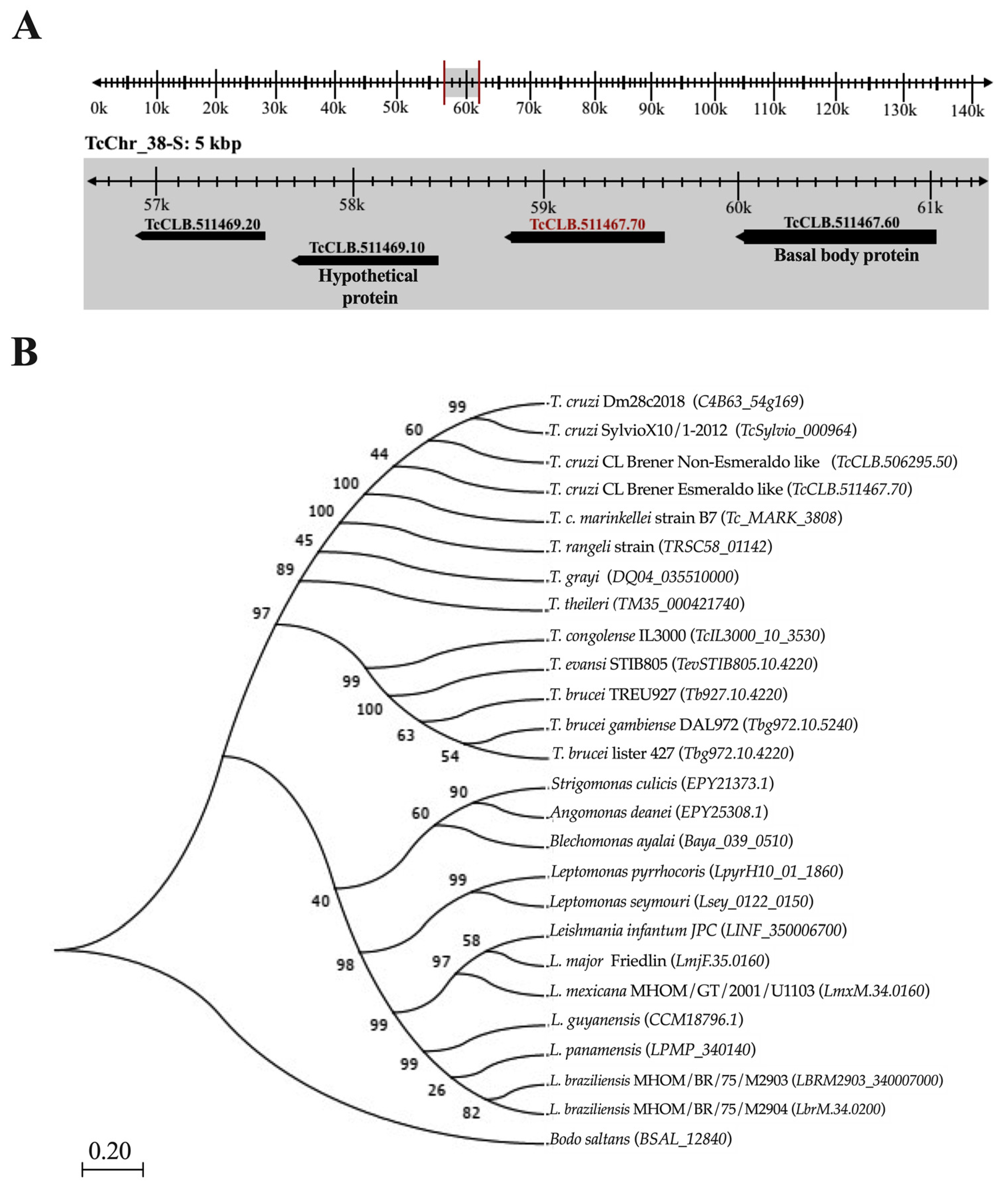
2.1. Tc964 Is Conserved in Different T. cruzi Discrete Typing Units (DTU) and Is Also Present in Other Kinetoplastids
2.2. Tc964 Protein May Be a Member of the GTPase Family
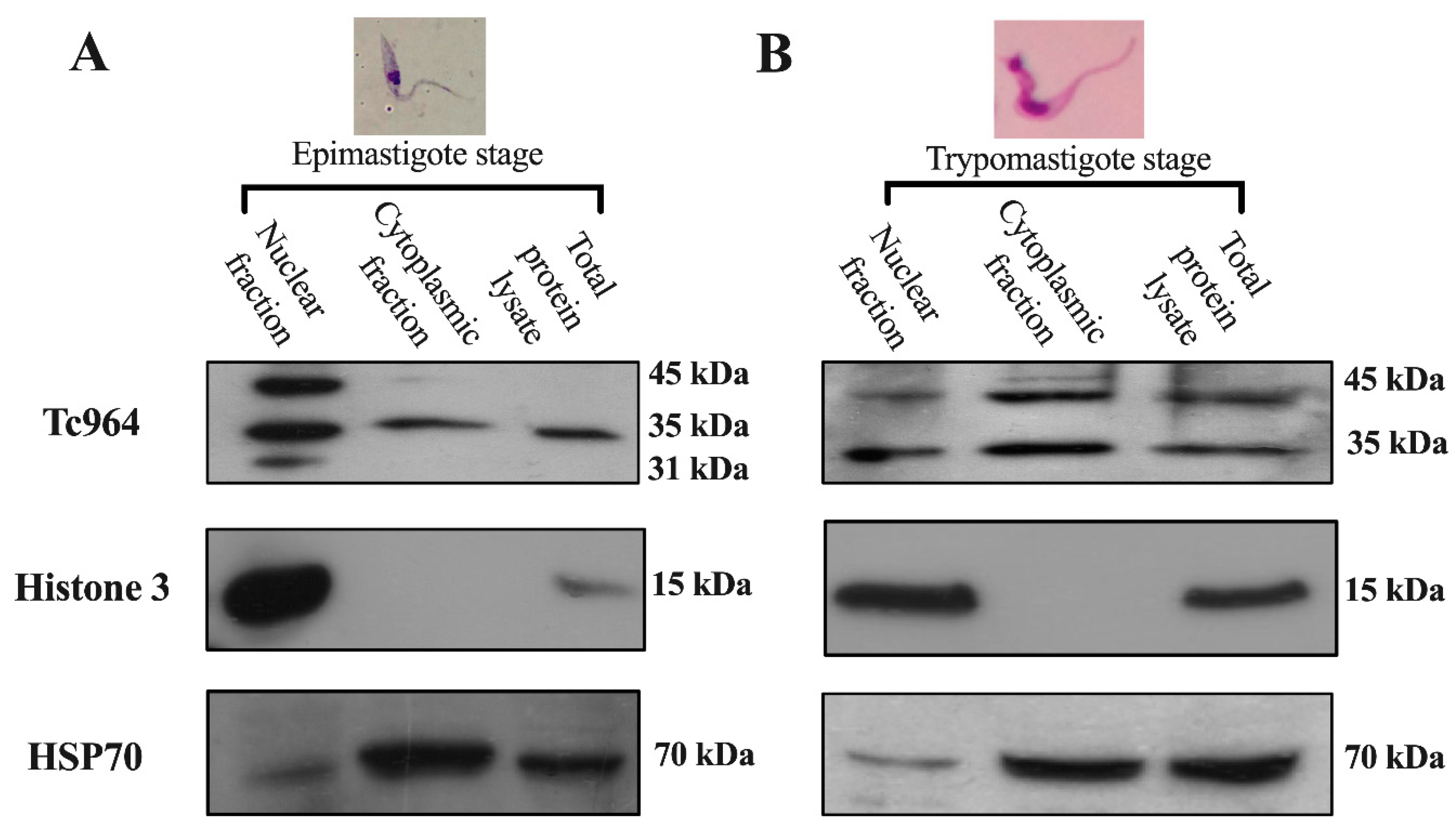
2.3. Tc964 is a Nucleocytoplasmic Protein
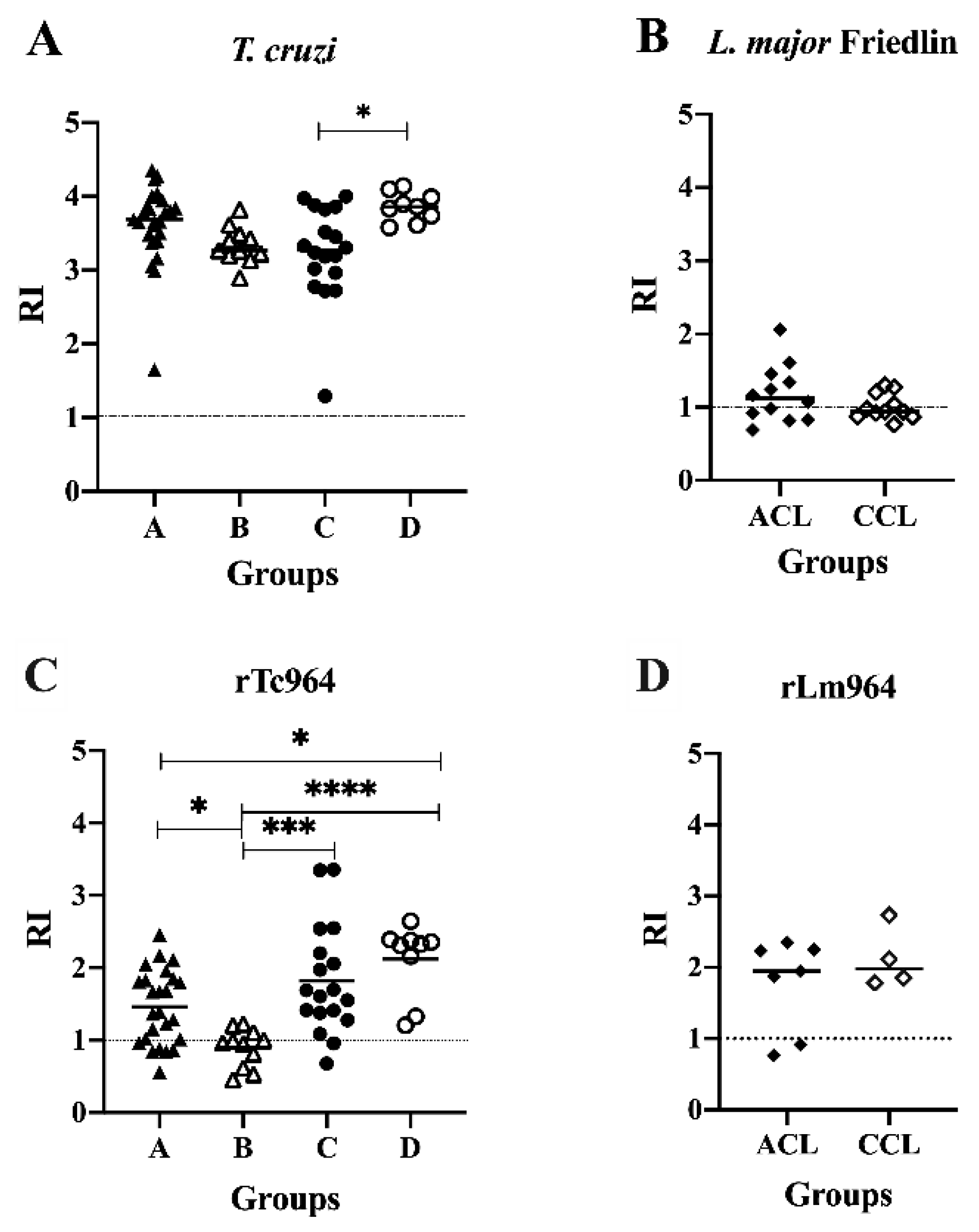
2.4. Tc964 is Recognized by Sera from Chronic Chagas Disease Patients
2.5. Tc964 Protein is Not Recognized by Sera from Leishmaniasis Patients
2.6. Two Peptides Derived from Tc964 Protein are Recognized by Sera from Chronic CD Patients
3. Discussion
4. Materials and Methods
4.1. Description of the TcCLB.511467.70 Gene in the Genome of T. cruzi
4.2. Phylogenetic Analysis of Protein Tc964
4.3. Fractionation and ELISA Test
4.3.1. Cultivation of Parasites
4.3.2. Subcellular Location of the Tc964 Protein of T. cruzi
4.3.3. Soluble Antigens from T. cruzi and L. major
4.4. Western Blot Assay
4.5. Structural Prediction of the Tc964 protein in T. cruzi and L. major
4.6. Cloning and Protein Expression
4.7. Prediction of Linear B-cell Epitopes and Peptide Synthesis
4.8. Sera Samples and Ethical Statement
4.9. ELISA Assay Protocol
4.10. Statistical and Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ACL | Active Cutaneous Leishmaniasis |
| CD | Chagas Disease |
| CCD | Chronic Chagas Disease |
| CCL | Curated Cutaneous Leishmaniasis |
| DTU | Discrete Typing Unit |
| ELISA | Enzyme-linked immunosorbent assay |
| kDa | Kilodaltons |
| PBS | phosphate buffered saline |
| RI | Reactivity index |
| rLm964 | Recombinant protein Lm964 |
| rTc964 | Recombinant protein Tc964 |
References
- Chagas, C. Nova tripanozomiaze humana. Mem. Inst. Oswaldo Cruz 1909, 1, 159–218. [Google Scholar] [CrossRef] [Green Version]
- Minchin, E. A new trypanosome parasitic in human beings. Nature 1910, 84, 142–144. [Google Scholar] [CrossRef]
- Perez-Molina, J.A.; Molina, I. Chagas disease. Lancet (Lond. Engl.) 2018, 391, 82–94. [Google Scholar] [CrossRef]
- Álvarez-Hernández, D.A.; Franyuti-Kelly, G.A.; Díaz López-Silva, R.; González Chávez, A.M.; González-Hermosillo, D.; Vásquez-López, R. Chagas disease: Current perspectives on a forgotten disease. Rev. Med. Hosp. Gen. (Mexico) 2016, 81, 154–164. [Google Scholar] [CrossRef]
- World Health Organization (WHO). Chagas Disease (American Trypanosomiasis). Available online: https://www.who.int/ (accessed on 17 April 2019).
- Organización Panamericana de la Salud. Guía Para el diagnóstico y el Tratamiento de la Enfermedad de Chagas. Washington, DC, USA: OPS. Available online: https://www.paho.org (accessed on 12 June 2019).
- Instituto Nacional de Salud. Informe de Evento de la Enfermedad de Chagas, Colombia, Periodo Epidemiológico I al VI de. Available online: https://www.ins.gov.co/ (accessed on 31 May 2019).
- Meymandi, S.; Hernandez, S.; Park, S.; Sanchez, D.R.; Forsyth, C. Treatment of Chagas disease in the United States. Curr. Treat Options Infect. Dis. 2018, 10, 373–388. [Google Scholar] [CrossRef] [Green Version]
- Montgomery, S.P.; Starr, M.C.; Cantey, P.T.; Edwards, M.S.; Meymandi, S.K. Neglected parasitic infections in the United States: Chagas disease. Am. J. Trop. Med. Hyg. 2014, 90, 814–818. [Google Scholar] [CrossRef] [Green Version]
- Balouz, V.; Agüero, F.; Buscaglia, C.A. Chagas disease diagnostic applications: Present knowledge and future steps. Adv. Parasitol. 2017, 97, 1–45. [Google Scholar] [CrossRef] [Green Version]
- Messenger, L.; Miles, M.; Bern, C. Between a bug and a hard place: Trypanosoma cruzi genetic diversity and the clinical outcomes of Chagas disease. Expert Rev. Anti-Infect. Ther. 2015, 13, 995–1029. [Google Scholar] [CrossRef] [Green Version]
- Daltro, R.T.; Leony, L.M.; Freitas, N.E.M.; Silva, Â.A.O.; Santos, E.F.; Del-Rei, R.P.; Brito, M.E.F.; BrandãoFilho, S.P.; Gomes, Y.M.; Silva, M.S.; et al. Cross-reactivity using chimeric Trypanosoma cruzi antigens: Diagnostic performance in settings co-endemic for Chagas disease and American cutaneous or visceral leishmaniasis. J. Clin. Microbiol. 2019, 57, e00762-19. [Google Scholar] [CrossRef] [Green Version]
- Dumonteil, E.; Herrera, C. Ten years of Chagas disease research: Looking back to achievements, looking ahead to challenges. PLoS Negl. Trop. Dis. 2017, 11, e0005422. [Google Scholar] [CrossRef]
- Santos, F.L.N.; De Souza, W.V.; Da Silva Barros, M.; Nakazawa, M.; Krieger, M.A.; de Miranda Gomes, Y. Chronic Chagas disease diagnosis: A comparative performance of commercial enzyme immunoassay tests. Am. J. Trop. Med. Hyg. 2016, 94, 1034–1039. [Google Scholar] [CrossRef] [PubMed]
- Thomas, M.C.; Fernández-Villegas, A.; Carrilero, B.; Marañón, C.; Saura, D.; Noya, O.; Segovia, M.; Alarcón de Noya, B.; Alonso, C.; López, M.C. Characterization of an immunodominant antigenic epitope from Trypanosoma cruzi as a biomarker of chronic Chagas’ disease pathology. Clin. Vaccine Immunol. 2012, 19, 167–173. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Florez, M.M.; de Oliveira, C.I.; Puerta, C.J.; Guzman, F.; Ayala, M.; Montoya, G.; Delgado, G. Synthetic peptides derived from ribosomal proteins of Leishmania spp. in mucocutaneous leishmaniasis: Diagnostic usefulness. Protein Pept. Lett. 2017, 24, 982–988. [Google Scholar] [CrossRef]
- Silveira, J.F.; Umezawa, E.S.; Luquetti, A.O. Chagas disease: Recombinant Trypanosoma cruzi antigens for serological diagnosis. Trends Parasitol. 2001, 17, 286–291. [Google Scholar] [CrossRef]
- Malchiodi, E.L.; Chiaramonte, M.G.; Taranto, N.J.; Zwirner, N.W.; Margni, R.A. Cross-reactivity studies and differential serodiagnosis of human infections caused by Trypanosoma cruzi and Leishmania spp; use of immunoblotting and ELISA with a purified antigen (Ag163B6). Clin. Exp. Immunol. 1994, 97, 417–423. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, E.; Ramírez, C.A.; Casas, J.C.; Ospina, M.I.; Requena, J.M.; Puerta, C.J. Characterization of the mRNA untranslated regions [UTR] of the Trypanosoma cruzi LYT1 isoforms derived by alternative trans-splicing. Univ. Sci. 2018, 23, 267–290. [Google Scholar] [CrossRef]
- Andrews, N.W.; Abrams, C.K.; Slatin, S.L.; Griffiths, G. A T. cruzi secreted protein immunologically related to the complement component C9: Evidence for membrane pore-forming activity at low pH. Cell 1990, 61, 1277–1287. [Google Scholar] [CrossRef]
- Benabdellah, K.; González, E.; González, A. Alternative trans splicing of the Trypanosoma cruzi LYT1 gene transcript results in compartmental and functional switch for the encoded protein. Mol. Microbiol. 2007, 65, 1559–1567. [Google Scholar] [CrossRef]
- Manning-Cela, R.; Cortés, A.; González-Rey, H.; Van Voorhis, W.; Swindle, J.; González, A. LYT1 protein is required for efficient in vitro infection by Trypanosoma cruzi. Infect. Immun. 2001, 69, 3916–3923. [Google Scholar] [CrossRef] [Green Version]
- Ruiz-Márvez, E.; Requena, J.M.; Puerta, C.J. Proteins associated to the UTRs of the LYT1 mRNAs. Status (unpublished; manuscript in preparation).
- Logan-Klumpler, F.J.; De Silva, N.; Boehme, U.; Rogers, M.B.; Velarde, G.; McQuillan, J.A.; Carver, T.; Aslett, M.; Olsen, C.; Subramanian, S.; et al. GeneDB–an annotation database for pathogens. Nucleic Acids Res. 2012, 40, D98–D108. [Google Scholar] [CrossRef]
- Aslett, M.; Aurrecoechea, C.; Berriman, M.; Brestelli, J.; Brunk, B.P.; Carrington, M.; Depledge, D.P.; Fischer, S.; Gajria, B.; Gao, X.; et al. TriTrypDB: A functional genomic resource for the Trypanosomatidae. Nucleic Acids Res. 2010, 38, D457–D462. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Benson, D.A.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Wheeler, D.L. GenBank. Nucleic Acids Res. 2005, 33, D34–D38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Sayed, N.M.; Myler, P.J.; Bartholomeu, D.C.; Nilsson, D.; Aggarwal, G.; Tran, A.N.; Ghedin, E.; Wortney, E.A.; Delcher, A.L.; Blandin, G.; et al. The genome sequence of Trypanosoma cruzi, etiologic agent of Chagas disease. Science 2005, 309, 409–415. [Google Scholar] [CrossRef] [Green Version]
- Berná, L.; Rodriguez, M.; Chiribao, M.L.; Parodi-Talice, A.; Pita, S.; Rijo, G.; Álvarez-Valin, F.; Robello, C. Expanding an expanded genome: Long-read sequencing of Trypanosoma cruzi. Microb. Genom. 2018, 4, e000177. [Google Scholar] [CrossRef] [PubMed]
- NCBI. Available online: https://www.ncbi.nlm.nih.gov/protein/PBJ69506 (accessed on 1 September 2019).
- Grisard, E.C.; Teixeira, S.M.; de Almeida, L.G.; Stoco, P.H.; Gerber, A.L.; Talavera-López, C.; Lima, O.C.; Andersson, B.; de Vasconcelos, A.T. Trypanosoma cruzi clone Dm28c draft genome sequence. Genome Announc. 2014, 30, e01114-13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franzén, O.; Arner, E.; Ferella, M.; Nilsson, D.; Respuela, P.; Carninci, P.; Hayashizaki, Y.; Åslund, L.; Andersson, B.; Daub, C.O. The short non-coding transcriptome of the protozoan parasite Trypanosoma cruzi. PLoS Negl. Trop. Dis. 2011, 5, e1283. [Google Scholar] [CrossRef] [Green Version]
- Franzén, O.; Talavera-López, C.; Ochaya, S.; Butler, C.E.; Messenger, L.A.; Lewis, M.D.; Llewellyn, M.S.; Marinkelle, C.J.; Tyler, K.M.; Miles, M.A.; et al. Comparative genomic analysis of human infective Trypanosoma cruzi lineages with the bat-restricted subspecies T. cruzi marinkellei. BMC Genom. 2012, 13, 531. [Google Scholar] [CrossRef]
- Maslov, D.A.; Opperdoes, F.R.; Kostygov, A.Y.; Hashimi, H.; Lukes, J.; Yurchenko, V. Recent advances in trypanosomatid research: Genome organization, expression, metabolism, taxonomy and evolution. Parasitology 2019, 146, 1–27. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. BLAST: Sequence searching using NCBI’s BLAST program. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Kim, D.E.; Chivian, D.; Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 2004, 32, W526–W531. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wild, K.; Bange, G.; Motiejunas, D.; Kribelbauer, J.; Hendricks, A.; Segnitz, B.; Wade, R.C.; Sinning, I. Structural basis for conserved regulation and adaptation of the signal recognition particle targeting complex. J. Mol. Biol. 2016, 428, 2880–2897. [Google Scholar] [CrossRef] [PubMed]
- Chandran, V.; Luisi, B.F. Recognition of enolase in the Escherichia coli RNA degradosome. J. Mol. Biol. 2006, 358, 8–15. [Google Scholar] [CrossRef]
- Das, P.; Babbar, P.; Malhotra, N.; Sharma, M.; Jachak, G.; Gonnade, R.G.; Shanmugam, D.; Harlos, K.; Yogavel, M.; Sharma, A.; et al. Specific stereoisomeric conformations determine the drug potency of cladosporin scaffold against malarial parasite. Int. J. Med. Chem. 2008, 61, 566–578. [Google Scholar] [CrossRef] [PubMed]
- Case, D.A.; Cheatham, T.E.; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R. The Amber biomolecular simulation programs. J. Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef] [Green Version]
- Wild, K.; Rosendal, K.R.; Sinning, I. A structural step into the SRP cycle. Mol. Microbiol. 2004, 53, 357–363. [Google Scholar] [CrossRef]
- Rosendal, K.R.; Wild, K.; Montoya, G.; Sinning, I. Crystal structure of the complete core of archaeal signal recognition particle and implications for interdomain communication. Proc. Natl. Acad. Sci. USA 2003, 25, 14701–14706. [Google Scholar] [CrossRef] [Green Version]
- The PyMOL Molecular Graphics System, Version 2.Schrodinger, L.L.C. Available online: https://pymol.org/2/#page-top (accessed on 13 September 2019).
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Soding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using clustal omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef]
- Xu, L.; Lubkov, V.; Taylor, L.J.; Bar-Sagi, D. Feedback regulation of Ras signaling by rabex-5-mediated ubiquitination. Curr. Biol. 2010, 20, 1372–1377. [Google Scholar] [CrossRef] [Green Version]
- Fu, X.; Liang, C.; Li, F.; Wang, L.; Wu, X.; Lu, A.; Xiao, G.; Zhang, G. The rules and functions of nucleocytoplasmic shuttling proteins. Int. J. Mol. Sci. 2018, 19, 1445. [Google Scholar] [CrossRef] [Green Version]
- Lobo, M.; Balouz, V.; Melli, L.; Carlevaro, G.; Cortina, M.E.; Cámara, M.; Cánepa, G.E.; Carmona, S.J.; Althech, J.; Campetella, O.; et al. Molecular and antigenic characterization of Trypanosoma cruzi TolT proteins. PLoS Negl. Trop. Dis. 2019, 13, e0007245. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X. Multistate GTPase Control Co-Translational Protein Targeting. Ph.D. Thesis, The California Institute of Technology, Pasadena, CA, USA, 2012. [Google Scholar] [CrossRef] [Green Version]
- Egea, P.F.; Tsuruta, H.; de Leon, G.P.; Napetschnig, J.; Walter, P.; Stroud, R.M. Structures of the signal recognition particle receptor from the archaeon Pyrococcus furiosus: Implications for the targeting step at the membrane. PLoS ONE 2008, 3, e3619. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Träger, C.; Rosenblad, M.A.; Ziehe, D.; Garcia-Petit, C.; Scharader, L.; Kock, K.; Richter, C.V.; Klinkert, B.; Narberhaus, F.; Herrmann, C.; et al. Evolution from the prokaryotic to the higher plant chloroplast signal recognition particle: The signal recognition RNA is conserved in plastids of a wide range of photosynthetic organisms. Plant Cell 2012, 24, 4819–4836. [Google Scholar] [CrossRef] [Green Version]
- Scott, M.S.; Calafell, S.J.; Thomas, D.Y.; Hallett, M.T. Refining protein subcellular localization. PLoS Comput. Biol. 2005, 1, 518–528. [Google Scholar] [CrossRef] [PubMed]
- Goody, R.S.; Muüller, M.P.; Wu, Y.W. Mechanisms of action of Rab proteins, key regulators of intracellular vesicular transport. Biol. Chem. 2017, 398, 565–575. [Google Scholar] [CrossRef] [PubMed]
- De la Vega, M.; Burrows, J.F.; Johnston, J.A. Ubiquitination. Small_GTPases 2011, 2, 192–201. [Google Scholar] [CrossRef] [Green Version]
- Nethe, M.; Hordijk, P.L. The role of ubiquitylation and degradation in Rho GTPase signalling. J. Cell Sci. 2010, 1, 4011–4018. [Google Scholar] [CrossRef] [Green Version]
- Gupta, I.; Aggarwal, S.; Singh, K.; Yadav, A.; Khan, S. Ubiquitin proteasome pathway proteins as potential drug targets in parasite Trypanosoma cruzi. Sci. Rep. 2018, 8, 1–12. [Google Scholar] [CrossRef]
- Ramírez, C.A.; Ruiz, E.; Rodríguez, E.; Burchmore, R.; Requena, J.M.; Puerta, C.J. Interactome of TcHy964 protein identified by its association with the 5′UTR of Trypanosoma cruzi LYT1 mRNA. In Proceedings of the Symposium on Omics Technologies for Research on Infectious Diseases; Centro Internacional de Entrenamiento e Investigaciones Médicas (CIDEIM): Cali, Colombia, 2017. [Google Scholar] [CrossRef]
- Flechas, I.D.; Cuellar, A.; Cucunubá, Z.M.; Rosas, F.; Velasco, V.; Steindel, M.; Thomas, M.C.; López, M.C.; González, J.M.; Puerta, C.J. Characterising the KMP-11 and HSP-70 recombinant antigens’ humoral immune response profile in chagasic patients. BMC Infect. Dis. 2009, 9, 186. [Google Scholar] [CrossRef] [Green Version]
- Passos, V.M.A.; Volpini, A.C.; Braga, E.M.; Lacerda, P.A.F.; Ouaissi, A.; Lima-Martins, M.V.C.; Krettli, A.U. Differential serodiagnosis of human infections caused by Trypanosoma cruzi and Leishmania spp. using ELISA with a recombinant antigen (rTc24). Mem. Inst. Oswaldo Cruz 1997, 92, 791–793. [Google Scholar] [CrossRef] [Green Version]
- Bottino, C.G.; Gomes, L.P.; Pereira, J.B.; Coura, J.R.; Provance, D.W.; De Simone, S.G. Chagas disease-specific antigens: Characterization of epitopes in CRA/FRA by synthetic peptide mapping and evaluation by ELISA-peptide assay. BMC Infect. Dis. 2013, 13, 568. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferreira, A.W.; Belem, Z.R.; Lemos, E.A.; Reed, S.G.; Campos-Neto, A. Enzyme-linked immunosorbent assay for serological diagnosis of Chagas’ disease employing a Trypanosoma cruzi recombinant antigen that consists of four different peptides. J. Clin. Microbiol. 2001, 39, 4390–4395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mucci, J.; Carmona, S.J.; Volcovich, R.; Altcheh, J.; Bracamonte, E.; Marco, J.D.; Nielsen, M.; Buscaglia, C.A.; Aguero, F. Next-generation ELISA diagnostic assay for Chagas disease based on the combination of short peptidic epitopes. PLoS Negl. Trop. Dis. 2017, 11, e0005972. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jackson, A.P.; Quail, M.A.; Berriman, M. Insights into the genome sequence of a free-living kinetoplastid: Bodo saltans (Kinetoplastida: Euglenozoa). BMC Genom. 2008, 9, 594. [Google Scholar] [CrossRef] [Green Version]
- Silva, L.; Nussenzweig, V. Sobre una cepa de Trypanosoma cruzi altamente virulenta para o camundongo branco. Fol. Clin. Biol. 1953, 20, 191–207. [Google Scholar]
- Ivens, A.C.; Peacock, C.S.; Worthey, E.A.; Murphy, L.; Aggarwal, G.; Berriman, M.; Sisk, E.; Rajandream, M.A.; Adlem, E.; Aert, R.; et al. The genome of the kinetoplastid parasite, Leishmania major. Science 2005, 309, 436–442. [Google Scholar] [CrossRef] [Green Version]
- Souza, A.P.; Soto, M.; Costa, J.M.; Boaventura, V.S.; de Oliveira, C.I.; Cristal, J.R.; Barral-Netto, M.; Barral, A. Towards a more precise serological diagnosis of human tegumentary leishmaniasis using Leishmania recombinant proteins. PLoS ONE 2013, 8, e66110. [Google Scholar] [CrossRef] [Green Version]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 97, 676–682. [Google Scholar] [CrossRef] [Green Version]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 2015, 10, 845–858. [Google Scholar] [CrossRef] [Green Version]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [Green Version]
- Benkert, P.; Biasini, M.; Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 2011, 3, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.U.; Zoete, V.; Michielin, O.; Guex, N. Defining and searching for structural motifs using deepview/swiss-pdbviewer. BMC Bioinform. 2012, 13, 173. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD-visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Andrew, S.; Titus, J.A.; Zumstein, L. Dialysis and concentration of protein solutions. Curr. Protoc. Immunol. 2001, 21, A3H1–A3H5. [Google Scholar]
- Yao, B.; Zhang, L.; Liang, S.; Zhang, C. SVMTriP: A method to predict antigenic epitopes using support vector machine to integrate tri-peptide similarity and propensity. PLoS ONE 2012, 9, e45152. [Google Scholar] [CrossRef] [Green Version]
- Rubinstein, N.D.; Mayrose, I.; Martz, E.; Pupko, T. Epitopia: A webserver for predicting B-cell epitopes. BMC Bioinform. 2009, 10, 287. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.; Liu, H.; Yang, J.; Chou, K.C. Prediction of linear B-cell epitopes using amino acid pair antigenicity scale. Amino Acids 2007, 33, 423–428. [Google Scholar] [CrossRef]
- Jespersen, M.C.; Peters, B.; Nielsen, M.; Marcatili, P. BepiPred-2.0: Improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res. 2017, 45, W24–W29. [Google Scholar] [CrossRef] [Green Version]
- Saha, S.; Raghava, G.P.S. Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins 2006, 65, 40–48. [Google Scholar] [CrossRef]
- Guzmán, F.; Wong, G.; Román, T.; Cárdenas, C.; Alvárez, C.; Schmitt, P.; Albericio, F.; Rojas, V. Identification of antimicrobial peptides from the microalgae Tetraselmis suecica (Kylin) Butcher and bactericidal activity improvement. Mar. Drugs 2019, 17, 453. [Google Scholar] [CrossRef] [Green Version]
- Yancy, C.W.; Jessup, M.; Bozkurt, B.; Butler, J.; Casey, D.E.; Colvin, M.; Drazner, M.H.; Filippatos, G.S.; Fonarow, G.C.; Givertz, M.M.; et al. 2017 ACC/AHA/HFSA focused update of the 2013 ACCF/AHA guideline for the management of heart failure. Circulation 2017, 136, 137–161. [Google Scholar] [CrossRef] [PubMed]
- Hunt, S.A.; Abraham, W.; Chin, M.; Feldman, A.; Francis, G.S.; Ganiats, T.G.; Jessup, M.; Konstam, M.; Mancini, D.; Michi, K.; et al. ACC/AHA 2005 guideline update for the diagnosis and management of chronic heart failure in the adult. Circulation 2005, 112, 154–235. [Google Scholar] [CrossRef] [Green Version]
- Lasso, P.; Mateus, J.; Pavía, P.; Rosas, F.; Roa, N.; Thomas, M.C.; López, M.C.; González, J.M.; Puerta, C.J.; Cuéllar, A. Inhibitory receptor expression on CD8+ T cells is linked to functional responses against Trypanosoma cruzi antigens in chronic chagasic patients. J. Immunol. 2015, 195, 3748–3758. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barral, A.; Honda, E.; Caldas, A.; Costa, J.; Vinhas, V.; Rowton, E.D.; Valenzuela, J.G.; Charlab, R.; Barral-Neto, M.; Ribeiro, J.M. Human immune response to sand fly salivary gland antigens: A useful epidemiological marker? Am. J. Trop. Med. Hyg. 2000, 62, 740–745. [Google Scholar] [CrossRef] [PubMed] [Green Version]



© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruiz-Márvez, E.; Ramírez, C.A.; Rodríguez, E.R.; Flórez, M.M.; Delgado, G.; Guzmán, F.; Gómez-Puertas, P.; Requena, J.M.; Puerta, C.J. Molecular Characterization of Tc964, A Novel Antigenic Protein from Trypanosoma cruzi. Int. J. Mol. Sci. 2020, 21, 2432. https://doi.org/10.3390/ijms21072432
Ruiz-Márvez E, Ramírez CA, Rodríguez ER, Flórez MM, Delgado G, Guzmán F, Gómez-Puertas P, Requena JM, Puerta CJ. Molecular Characterization of Tc964, A Novel Antigenic Protein from Trypanosoma cruzi. International Journal of Molecular Sciences. 2020; 21(7):2432. https://doi.org/10.3390/ijms21072432
Chicago/Turabian StyleRuiz-Márvez, Elizabeth, César Augusto Ramírez, Eliana Rocío Rodríguez, Magda Mellisa Flórez, Gabriela Delgado, Fanny Guzmán, Paulino Gómez-Puertas, José María Requena, and Concepción J. Puerta. 2020. "Molecular Characterization of Tc964, A Novel Antigenic Protein from Trypanosoma cruzi" International Journal of Molecular Sciences 21, no. 7: 2432. https://doi.org/10.3390/ijms21072432







